rs12326991
Homo sapiens
T>C / T>G
None
SNV (Single Nucleotide Variation)
C=0295 (8847/29902,GnomAD)
C=0268 (7805/29118,TOPMED)
C=0281 (1407/5008,1000G)
C=0401 (1547/3854,ALSPAC)
C=0381 (1413/3708,TWINSUK)
C=0268 (7805/29118,TOPMED)
C=0281 (1407/5008,1000G)
C=0401 (1547/3854,ALSPAC)
C=0381 (1413/3708,TWINSUK)
chr18:78418452 (GRCh38.p7) (18q23)
AD
GWASdb2
Genomic Coordinates
| Sequence Name | Change(s) |
|---|---|
| GRCh38.p7 chr 18 | NC_000018.10:g.78418452T>C |
| GRCh38.p7 chr 18 | NC_000018.10:g.78418452T>G |
| GRCh37.p13 chr 18 | NC_000018.9:g.76178452T>C |
| GRCh37.p13 chr 18 | NC_000018.9:g.76178452T>G |
Population Frequency
| Study | Population | Group | Sample # | Ref Allele | Alt Allele |
|---|---|---|---|---|---|
| 1000Genomes | African | Sub | 1322 | T=0.908 | C=0.092 |
| 1000Genomes | American | Sub | 694 | T=0.710 | C=0.290 |
| 1000Genomes | East Asian | Sub | 1008 | T=0.638 | C=0.362 |
| 1000Genomes | Europe | Sub | 1006 | T=0.628 | C=0.372 |
| 1000Genomes | Global | Study-wide | 5008 | T=0.719 | C=0.281 |
| 1000Genomes | South Asian | Sub | 978 | T=0.650 | C=0.350 |
| The Avon Longitudinal Study of Parents and Children | PARENT AND CHILD COHORT | Study-wide | 3854 | T=0.599 | C=0.401 |
| The Genome Aggregation Database | African | Sub | 8714 | T=0.870 | G=0.000 |
| The Genome Aggregation Database | American | Sub | 836 | T=0.740 | G=0.00, |
| The Genome Aggregation Database | East Asian | Sub | 1612 | T=0.612 | G=0.000 |
| The Genome Aggregation Database | Europe | Sub | 18438 | T=0.633 | G=0.000 |
| The Genome Aggregation Database | Global | Study-wide | 29902 | T=0.704 | G=0.000 |
| The Genome Aggregation Database | Other | Sub | 302 | T=0.650 | G=0.00, |
| Trans-Omics for Precision Medicine | Global | Study-wide | 29118 | T=0.732 | C=0.268 |
| UK 10K study - Twins | TWIN COHORT | Study-wide | 3708 | T=0.619 | C=0.381 |
| PMID | Title | Author | Journal |
|---|---|---|---|
| 20201924 | Genome-wide association study of alcohol dependence implicates a region on chromosome 11. | Edenberg HJ | Alcohol Clin Exp Res |
P-Value
| SNP ID | p-value | Traits | Study |
|---|---|---|---|
| rs12326991 | 0.000762 | alcohol dependence | 20201924 |
eQTL of rs12326991 in Brain tissues (GTEx Analysis Release V7)
| Position (v37) | eGene | GeneID | Variant | p-value | TSS | Tissue | |
|---|---|---|---|---|---|---|---|
| There is no eQTL annotation for this SNP | |||||||
meQTL of rs12326991 in Fetal Brain
| Probe ID | Position | Gene | beta | p-value | |||
|---|---|---|---|---|---|---|---|
| There is no meQTL annotation for this SNP | |||||||
Genomic View
Chromatin Interaction
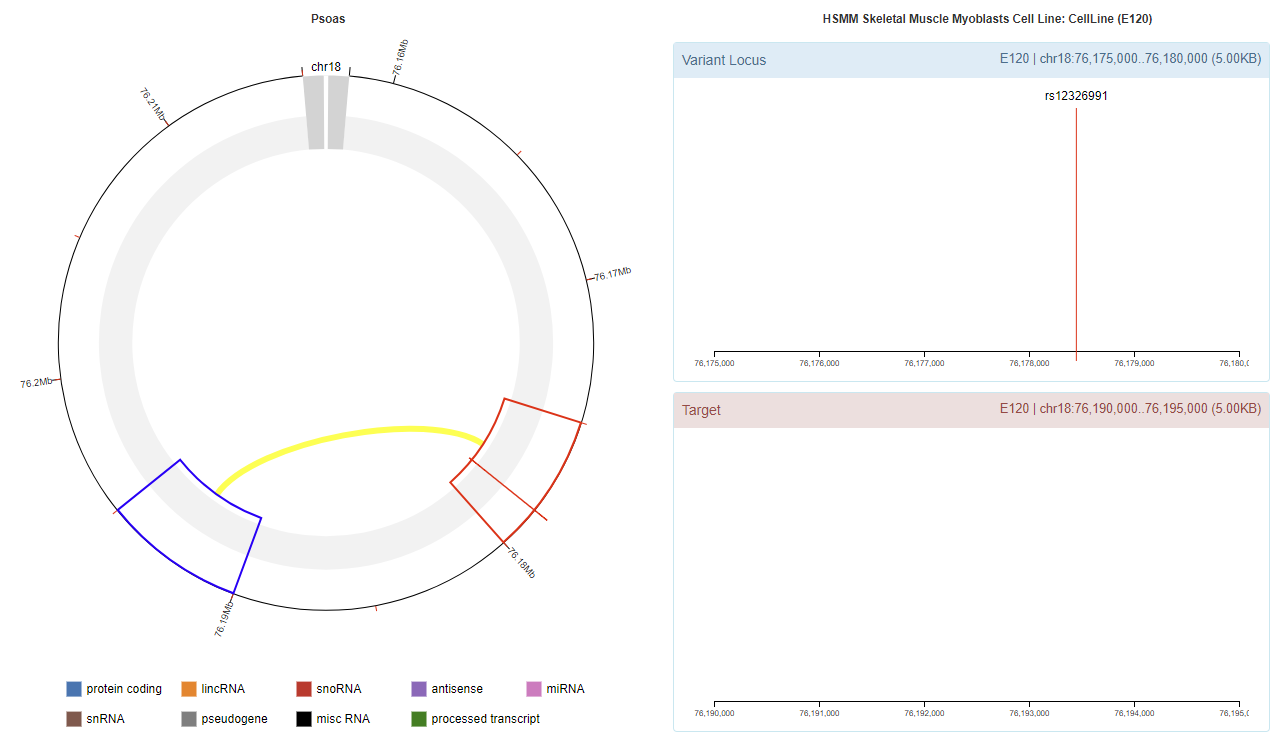
Enhancer Annotation (GRCh37.p13)
| Chromosome | Start | End | Region | Distance ( -/+ : Up/Downstream ) |
|---|---|---|---|---|
| chr18 | 76856486 | 76856582 | E067 | 33886 |
| chr18 | 76856598 | 76856664 | E067 | 33998 |
| chr18 | 76837302 | 76837489 | E068 | 14702 |
| chr18 | 76837919 | 76837979 | E068 | 15319 |
| chr18 | 76838025 | 76838075 | E068 | 15425 |
| chr18 | 76838145 | 76838195 | E068 | 15545 |
| chr18 | 76853280 | 76853370 | E068 | 30680 |
| chr18 | 76853692 | 76853742 | E068 | 31092 |
| chr18 | 76854132 | 76854287 | E068 | 31532 |
| chr18 | 76867854 | 76867936 | E068 | 45254 |
| chr18 | 76868077 | 76868169 | E068 | 45477 |
| chr18 | 76868499 | 76868549 | E068 | 45899 |
| chr18 | 76868612 | 76868663 | E068 | 46012 |
| chr18 | 76854132 | 76854287 | E069 | 31532 |
| chr18 | 76867854 | 76867936 | E069 | 45254 |
| chr18 | 76868077 | 76868169 | E069 | 45477 |
| chr18 | 76868499 | 76868549 | E069 | 45899 |
| chr18 | 76868612 | 76868663 | E069 | 46012 |
| chr18 | 76868716 | 76868811 | E069 | 46116 |
| chr18 | 76805970 | 76806024 | E070 | -16576 |
| chr18 | 76806308 | 76806399 | E070 | -16201 |
| chr18 | 76831079 | 76831144 | E070 | 8479 |
| chr18 | 76831162 | 76831364 | E070 | 8562 |
| chr18 | 76831387 | 76831536 | E070 | 8787 |
| chr18 | 76837302 | 76837489 | E071 | 14702 |
| chr18 | 76837919 | 76837979 | E071 | 15319 |
| chr18 | 76838025 | 76838075 | E071 | 15425 |
| chr18 | 76838145 | 76838195 | E071 | 15545 |
| chr18 | 76845090 | 76845260 | E071 | 22490 |
| chr18 | 76845432 | 76845526 | E071 | 22832 |
| chr18 | 76845678 | 76845832 | E071 | 23078 |
| chr18 | 76852949 | 76853130 | E071 | 30349 |
| chr18 | 76853280 | 76853370 | E071 | 30680 |
| chr18 | 76853692 | 76853742 | E071 | 31092 |
| chr18 | 76854132 | 76854287 | E071 | 31532 |
| chr18 | 76868499 | 76868549 | E071 | 45899 |
| chr18 | 76868612 | 76868663 | E071 | 46012 |
| chr18 | 76868716 | 76868811 | E071 | 46116 |
| chr18 | 76831079 | 76831144 | E072 | 8479 |
| chr18 | 76831162 | 76831364 | E072 | 8562 |
| chr18 | 76831387 | 76831536 | E072 | 8787 |
| chr18 | 76867854 | 76867936 | E072 | 45254 |
| chr18 | 76868077 | 76868169 | E072 | 45477 |
| chr18 | 76868499 | 76868549 | E072 | 45899 |
| chr18 | 76868612 | 76868663 | E072 | 46012 |
| chr18 | 76868716 | 76868811 | E072 | 46116 |
| chr18 | 76831079 | 76831144 | E073 | 8479 |
| chr18 | 76831162 | 76831364 | E073 | 8562 |
| chr18 | 76831387 | 76831536 | E073 | 8787 |
| chr18 | 76831549 | 76831816 | E073 | 8949 |
| chr18 | 76831079 | 76831144 | E074 | 8479 |
| chr18 | 76831162 | 76831364 | E074 | 8562 |
| chr18 | 76852949 | 76853130 | E074 | 30349 |
| chr18 | 76853280 | 76853370 | E074 | 30680 |
| chr18 | 76853692 | 76853742 | E074 | 31092 |
| chr18 | 76854132 | 76854287 | E074 | 31532 |
| chr18 | 76854463 | 76854513 | E074 | 31863 |
| chr18 | 76854517 | 76854582 | E074 | 31917 |
| chr18 | 76854770 | 76854820 | E074 | 32170 |
| chr18 | 76854871 | 76854996 | E074 | 32271 |
| chr18 | 76855173 | 76855223 | E074 | 32573 |
| chr18 | 76855270 | 76855473 | E074 | 32670 |
| chr18 | 76855506 | 76855722 | E074 | 32906 |
| chr18 | 76855992 | 76856137 | E074 | 33392 |
| chr18 | 76856486 | 76856582 | E074 | 33886 |
| chr18 | 76856598 | 76856664 | E074 | 33998 |
| chr18 | 76867854 | 76867936 | E074 | 45254 |
| chr18 | 76868077 | 76868169 | E074 | 45477 |
| chr18 | 76868499 | 76868549 | E074 | 45899 |
| chr18 | 76868612 | 76868663 | E074 | 46012 |
| chr18 | 76868716 | 76868811 | E074 | 46116 |
| chr18 | 76831079 | 76831144 | E081 | 8479 |
| chr18 | 76831162 | 76831364 | E081 | 8562 |
Promoter Annotation (GRCh37.p13)
| Chromosome | Start | End | Region | Distance(-/+:Up/Downstream) |
|---|---|---|---|---|
| chr18 | 76828110 | 76830565 | E067 | 5510 |
| chr18 | 76828110 | 76830565 | E068 | 5510 |
| chr18 | 76828110 | 76830565 | E069 | 5510 |
| chr18 | 76828110 | 76830565 | E070 | 5510 |
| chr18 | 76828110 | 76830565 | E071 | 5510 |
| chr18 | 76828110 | 76830565 | E072 | 5510 |
| chr18 | 76828110 | 76830565 | E073 | 5510 |
| chr18 | 76828110 | 76830565 | E074 | 5510 |
| chr18 | 76828110 | 76830565 | E082 | 5510 |