Gene Page: MAPKBP1
Summary ?
| GeneID | 23005 |
| Symbol | MAPKBP1 |
| Synonyms | JNKBP-1|JNKBP1 |
| Description | mitogen-activated protein kinase binding protein 1 |
| Reference | MIM:616786|HGNC:HGNC:29536|Ensembl:ENSG00000137802|HPRD:17465|Vega:OTTHUMG00000160227 |
| Gene type | protein-coding |
| Map location | 15q15.1 |
| Pascal p-value | 0.672 |
| Sherlock p-value | 0.297 |
| Fetal beta | 0.929 |
| eGene | Caudate basal ganglia Cerebellar Hemisphere Cerebellum Cortex Frontal Cortex BA9 Nucleus accumbens basal ganglia Myers' cis & trans Meta |
| Support | CompositeSet Darnell FMRP targets Ascano FMRP targets |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| PMID:cooccur | High-throughput literature-search | Systematic search in PubMed for genes co-occurring with SCZ keywords. A total of 3027 genes were included. | |
| Literature | High-throughput literature-search | Co-occurance with Schizophrenia keywords: schizophrenic | Click to show details |
Section I. Genetics and epigenetics annotation
 eQTL annotation
eQTL annotation
| SNP ID | Chromosome | Position | eGene | Gene Entrez ID | pvalue | qvalue | TSS distance | eQTL type |
|---|---|---|---|---|---|---|---|---|
| rs16829545 | chr2 | 151977407 | MAPKBP1 | 23005 | 1.478E-16 | trans | ||
| rs3845734 | chr2 | 171125572 | MAPKBP1 | 23005 | 0.01 | trans | ||
| rs7584986 | chr2 | 184111432 | MAPKBP1 | 23005 | 2.881E-5 | trans | ||
| rs2183142 | chr4 | 159232695 | MAPKBP1 | 23005 | 0.07 | trans | ||
| rs170776 | chr4 | 173276735 | MAPKBP1 | 23005 | 0.13 | trans | ||
| rs1396222 | chr4 | 173279496 | MAPKBP1 | 23005 | 0.13 | trans | ||
| rs335993 | chr4 | 173321789 | MAPKBP1 | 23005 | 0.18 | trans | ||
| rs335980 | chr4 | 173329784 | MAPKBP1 | 23005 | 0.11 | trans | ||
| rs335982 | chr4 | 173330945 | MAPKBP1 | 23005 | 0.15 | trans | ||
| rs336016 | chr4 | 173366576 | MAPKBP1 | 23005 | 0.17 | trans | ||
| rs337984 | chr4 | 173411662 | MAPKBP1 | 23005 | 0.09 | trans | ||
| rs17762315 | chr5 | 76807576 | MAPKBP1 | 23005 | 0.02 | trans | ||
| rs9461864 | chr6 | 33481468 | MAPKBP1 | 23005 | 0.01 | trans | ||
| rs7787830 | chr7 | 98797019 | MAPKBP1 | 23005 | 0.03 | trans | ||
| rs3118341 | chr9 | 25185518 | MAPKBP1 | 23005 | 0.16 | trans | ||
| rs9406868 | chr9 | 25223372 | MAPKBP1 | 23005 | 0.16 | trans | ||
| rs11139334 | chr9 | 84209393 | MAPKBP1 | 23005 | 0.17 | trans | ||
| rs16955618 | chr15 | 29937543 | MAPKBP1 | 23005 | 2.07E-19 | trans | ||
| rs11873184 | chr18 | 1584081 | MAPKBP1 | 23005 | 0.05 | trans | ||
| rs1041786 | chr21 | 22617710 | MAPKBP1 | 23005 | 0.17 | trans |
Section II. Transcriptome annotation
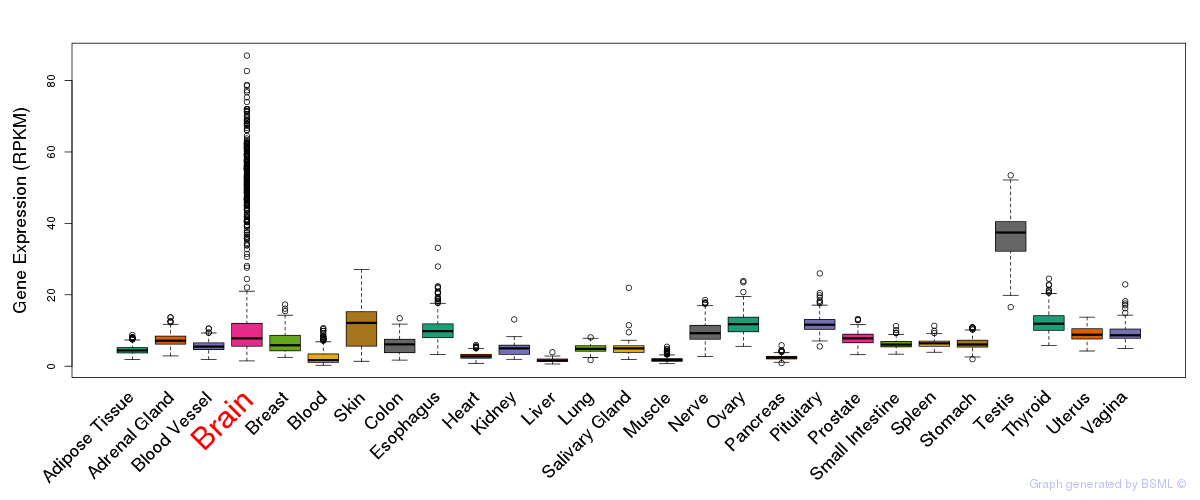
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| WNK1 | 0.92 | 0.91 |
| CLN8 | 0.88 | 0.89 |
| SPTLC2 | 0.87 | 0.88 |
| KIAA1147 | 0.85 | 0.88 |
| USP54 | 0.85 | 0.82 |
| DENND5A | 0.84 | 0.87 |
| DST | 0.84 | 0.83 |
| BBX | 0.84 | 0.81 |
| AFF1 | 0.84 | 0.82 |
| FAM55C | 0.83 | 0.86 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| ST20 | -0.50 | -0.57 |
| C1orf61 | -0.45 | -0.50 |
| IL32 | -0.44 | -0.52 |
| PFDN5 | -0.44 | -0.45 |
| TIMM8B | -0.43 | -0.45 |
| C8orf59 | -0.42 | -0.45 |
| C1orf54 | -0.42 | -0.50 |
| SYCP3 | -0.42 | -0.51 |
| AF347015.21 | -0.41 | -0.44 |
| SERF2 | -0.41 | -0.45 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| GRAESSMANN APOPTOSIS BY DOXORUBICIN UP | 1142 | 669 | All SZGR 2.0 genes in this pathway |
| GRAESSMANN RESPONSE TO MC AND DOXORUBICIN UP | 612 | 367 | All SZGR 2.0 genes in this pathway |
| FOSTER TOLERANT MACROPHAGE DN | 409 | 268 | All SZGR 2.0 genes in this pathway |
| KONDO COLON CANCER HCP WITH H3K27ME1 | 26 | 13 | All SZGR 2.0 genes in this pathway |
| SMID BREAST CANCER RELAPSE IN BRAIN DN | 85 | 58 | All SZGR 2.0 genes in this pathway |
| BOCHKIS FOXA2 TARGETS | 425 | 261 | All SZGR 2.0 genes in this pathway |
| GU PDEF TARGETS UP | 71 | 49 | All SZGR 2.0 genes in this pathway |
| MILI PSEUDOPODIA HAPTOTAXIS UP | 518 | 299 | All SZGR 2.0 genes in this pathway |
| YAGI AML WITH INV 16 TRANSLOCATION | 422 | 277 | All SZGR 2.0 genes in this pathway |
| PANGAS TUMOR SUPPRESSION BY SMAD1 AND SMAD5 DN | 157 | 106 | All SZGR 2.0 genes in this pathway |
| WAKABAYASHI ADIPOGENESIS PPARG RXRA BOUND 8D | 882 | 506 | All SZGR 2.0 genes in this pathway |
| WAKABAYASHI ADIPOGENESIS PPARG RXRA BOUND WITH H4K20ME1 MARK | 145 | 82 | All SZGR 2.0 genes in this pathway |
| FORTSCHEGGER PHF8 TARGETS DN | 784 | 464 | All SZGR 2.0 genes in this pathway |