Gene Page: ZC3H3
Summary ?
| GeneID | 23144 |
| Symbol | ZC3H3 |
| Synonyms | ZC3HDC3 |
| Description | zinc finger CCCH-type containing 3 |
| Reference | HGNC:HGNC:28972|Ensembl:ENSG00000014164|HPRD:15697|Vega:OTTHUMG00000165127 |
| Gene type | protein-coding |
| Map location | 8q24.3 |
| Pascal p-value | 0.08 |
| DMG | 1 (# studies) |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| DMG:Wockner_2014 | Genome-wide DNA methylation analysis | This dataset includes 4641 differentially methylated probes corresponding to 2929 unique genes between schizophrenia patients (n=24) and controls (n=24). | 2 |
Section I. Genetics and epigenetics annotation
 Differentially methylated gene
Differentially methylated gene
| Probe | Chromosome | Position | Nearest gene | P (dis) | Beta (dis) | FDR (dis) | Study |
|---|---|---|---|---|---|---|---|
| cg25834632 | 8 | 144590015 | ZC3H3 | 4.21E-6 | 0.912 | 0.01 | DMG:Wockner_2014 |
| cg03612190 | 8 | 144565290 | ZC3H3 | 5.577E-4 | 0.468 | 0.049 | DMG:Wockner_2014 |
Section II. Transcriptome annotation
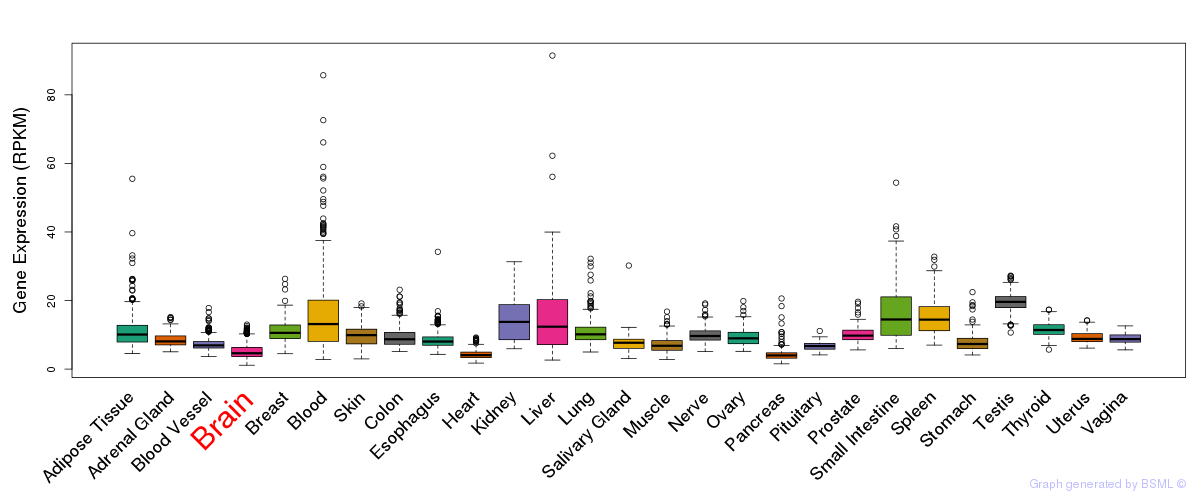
General gene expression (GTEx)
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| INSR | 0.95 | 0.96 |
| MFHAS1 | 0.94 | 0.96 |
| BACH2 | 0.94 | 0.94 |
| MYST4 | 0.94 | 0.95 |
| ASXL3 | 0.94 | 0.94 |
| CELSR3 | 0.93 | 0.94 |
| IGF1R | 0.93 | 0.94 |
| FAM117B | 0.93 | 0.95 |
| KIAA0240 | 0.93 | 0.98 |
| FAT4 | 0.93 | 0.92 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| SERPINB6 | -0.67 | -0.77 |
| HSD17B14 | -0.63 | -0.81 |
| AIFM3 | -0.62 | -0.76 |
| C5orf53 | -0.62 | -0.74 |
| TSC22D4 | -0.62 | -0.78 |
| HEPN1 | -0.61 | -0.72 |
| S100B | -0.61 | -0.84 |
| AF347015.31 | -0.61 | -0.87 |
| HLA-F | -0.61 | -0.72 |
| RAMP1 | -0.61 | -0.78 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| DEURIG T CELL PROLYMPHOCYTIC LEUKEMIA UP | 368 | 234 | All SZGR 2.0 genes in this pathway |
| DOANE RESPONSE TO ANDROGEN UP | 184 | 125 | All SZGR 2.0 genes in this pathway |
| GRAESSMANN APOPTOSIS BY SERUM DEPRIVATION DN | 234 | 147 | All SZGR 2.0 genes in this pathway |
| GRAESSMANN RESPONSE TO MC AND SERUM DEPRIVATION DN | 84 | 54 | All SZGR 2.0 genes in this pathway |
| LIAO METASTASIS | 539 | 324 | All SZGR 2.0 genes in this pathway |
| NIKOLSKY BREAST CANCER 8Q23 Q24 AMPLICON | 157 | 87 | All SZGR 2.0 genes in this pathway |
| PENG LEUCINE DEPRIVATION DN | 187 | 122 | All SZGR 2.0 genes in this pathway |
| SHEPARD BMYB MORPHOLINO UP | 205 | 126 | All SZGR 2.0 genes in this pathway |
| PENG GLUCOSE DEPRIVATION DN | 169 | 112 | All SZGR 2.0 genes in this pathway |
| AGUIRRE PANCREATIC CANCER COPY NUMBER UP | 298 | 174 | All SZGR 2.0 genes in this pathway |
| YAGI AML WITH T 8 21 TRANSLOCATION | 368 | 247 | All SZGR 2.0 genes in this pathway |
| MARTENS TRETINOIN RESPONSE UP | 857 | 456 | All SZGR 2.0 genes in this pathway |
| DELACROIX RAR BOUND ES | 462 | 273 | All SZGR 2.0 genes in this pathway |