Gene Page: LINC00339
Summary ?
| GeneID | 29092 |
| Symbol | LINC00339 |
| Synonyms | HSPC157|NCRNA00339 |
| Description | long intergenic non-protein coding RNA 339 |
| Reference | HGNC:HGNC:25011|Ensembl:ENSG00000218510| |
| Gene type | ncRNA |
| Map location | 1p36.12 |
| Pascal p-value | 0.02 |
| Sherlock p-value | 1.106E-5 |
| Fetal beta | 0.367 |
| eGene | Anterior cingulate cortex BA24 Caudate basal ganglia Cerebellar Hemisphere Cerebellum Cortex Frontal Cortex BA9 Hippocampus Hypothalamus Nucleus accumbens basal ganglia Putamen basal ganglia |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:GWASdb | Genome-wide Association Studies | GWASdb records for schizophrenia | |
| CV:PGCnp | Genome-wide Association Study | GWAS |
Section I. Genetics and epigenetics annotation
 eQTL annotation
eQTL annotation
| SNP ID | Chromosome | Position | eGene | Gene Entrez ID | pvalue | qvalue | TSS distance | eQTL type |
|---|---|---|---|---|---|---|---|---|
| rs12097709 | 1 | 22347138 | LINC00339 | ENSG00000218510.3 | 3.537E-7 | 0 | -4543 | gtex_brain_ba24 |
| rs12097775 | 1 | 22347252 | LINC00339 | ENSG00000218510.3 | 2.486E-8 | 0 | -4429 | gtex_brain_ba24 |
| rs2865198 | 1 | 22348104 | LINC00339 | ENSG00000218510.3 | 2.235E-8 | 0 | -3577 | gtex_brain_ba24 |
| rs11586488 | 1 | 22348556 | LINC00339 | ENSG00000218510.3 | 2.206E-8 | 0 | -3125 | gtex_brain_ba24 |
| rs10917120 | 1 | 22350165 | LINC00339 | ENSG00000218510.3 | 3.987E-9 | 0 | -1516 | gtex_brain_ba24 |
| rs12061255 | 1 | 22350547 | LINC00339 | ENSG00000218510.3 | 3.81E-9 | 0 | -1134 | gtex_brain_ba24 |
| rs3754505 | 1 | 22351326 | LINC00339 | ENSG00000218510.3 | 3.584E-9 | 0 | -355 | gtex_brain_ba24 |
| rs11801382 | 1 | 22352108 | LINC00339 | ENSG00000218510.3 | 3.45E-9 | 0 | 427 | gtex_brain_ba24 |
| rs74813454 | 1 | 22352395 | LINC00339 | ENSG00000218510.3 | 6.437E-10 | 0 | 714 | gtex_brain_ba24 |
| rs141271718 | 1 | 22353345 | LINC00339 | ENSG00000218510.3 | 2.056E-7 | 0 | 1664 | gtex_brain_ba24 |
| rs3768571 | 1 | 22354327 | LINC00339 | ENSG00000218510.3 | 5.589E-8 | 0 | 2646 | gtex_brain_ba24 |
| rs61778044 | 1 | 22355631 | LINC00339 | ENSG00000218510.3 | 5.696E-8 | 0 | 3950 | gtex_brain_ba24 |
| rs56408422 | 1 | 22360138 | LINC00339 | ENSG00000218510.3 | 5.473E-8 | 0 | 8457 | gtex_brain_ba24 |
| rs12748456 | 1 | 22370157 | LINC00339 | ENSG00000218510.3 | 2.711E-7 | 0 | 18476 | gtex_brain_ba24 |
| rs6661161 | 1 | 22373671 | LINC00339 | ENSG00000218510.3 | 2.701E-7 | 0 | 21990 | gtex_brain_ba24 |
| rs71638823 | 1 | 22374032 | LINC00339 | ENSG00000218510.3 | 1.5E-6 | 0 | 22351 | gtex_brain_ba24 |
| rs11579182 | 1 | 22374693 | LINC00339 | ENSG00000218510.3 | 1.699E-6 | 0 | 23012 | gtex_brain_ba24 |
| rs11580039 | 1 | 22375190 | LINC00339 | ENSG00000218510.3 | 2.015E-6 | 0 | 23509 | gtex_brain_ba24 |
| rs3754501 | 1 | 22377543 | LINC00339 | ENSG00000218510.3 | 2.043E-6 | 0 | 25862 | gtex_brain_ba24 |
| rs3754500 | 1 | 22377718 | LINC00339 | ENSG00000218510.3 | 2.015E-6 | 0 | 26037 | gtex_brain_ba24 |
| rs3754497 | 1 | 22378670 | LINC00339 | ENSG00000218510.3 | 1.999E-6 | 0 | 26989 | gtex_brain_ba24 |
| rs34446016 | 1 | 22458637 | LINC00339 | ENSG00000218510.3 | 1.846E-6 | 0 | 106956 | gtex_brain_ba24 |
| rs55809030 | 1 | 22462703 | LINC00339 | ENSG00000218510.3 | 1.407E-6 | 0 | 111022 | gtex_brain_ba24 |
| rs12097709 | 1 | 22347138 | LINC00339 | ENSG00000218510.3 | 6.683E-12 | 0 | -4543 | gtex_brain_putamen_basal |
| rs12097775 | 1 | 22347252 | LINC00339 | ENSG00000218510.3 | 7.572E-12 | 0 | -4429 | gtex_brain_putamen_basal |
| rs2865198 | 1 | 22348104 | LINC00339 | ENSG00000218510.3 | 1.059E-12 | 0 | -3577 | gtex_brain_putamen_basal |
| rs11586488 | 1 | 22348556 | LINC00339 | ENSG00000218510.3 | 1.806E-12 | 0 | -3125 | gtex_brain_putamen_basal |
| rs3117047 | 1 | 22348587 | LINC00339 | ENSG00000218510.3 | 1.477E-6 | 0 | -3094 | gtex_brain_putamen_basal |
| rs11588264 | 1 | 22349150 | LINC00339 | ENSG00000218510.3 | 4.51E-7 | 0 | -2531 | gtex_brain_putamen_basal |
| rs10799730 | 1 | 22349615 | LINC00339 | ENSG00000218510.3 | 8.505E-7 | 0 | -2066 | gtex_brain_putamen_basal |
| rs10917120 | 1 | 22350165 | LINC00339 | ENSG00000218510.3 | 8.04E-14 | 0 | -1516 | gtex_brain_putamen_basal |
| rs12061255 | 1 | 22350547 | LINC00339 | ENSG00000218510.3 | 7.229E-14 | 0 | -1134 | gtex_brain_putamen_basal |
| rs2473307 | 1 | 22350834 | LINC00339 | ENSG00000218510.3 | 1.975E-6 | 0 | -847 | gtex_brain_putamen_basal |
| rs3754505 | 1 | 22351326 | LINC00339 | ENSG00000218510.3 | 4.381E-14 | 0 | -355 | gtex_brain_putamen_basal |
| rs61778042 | 1 | 22351947 | LINC00339 | ENSG00000218510.3 | 5.082E-12 | 0 | 266 | gtex_brain_putamen_basal |
| rs2255282 | 1 | 22352040 | LINC00339 | ENSG00000218510.3 | 2.432E-7 | 0 | 359 | gtex_brain_putamen_basal |
| rs11801382 | 1 | 22352108 | LINC00339 | ENSG00000218510.3 | 2.618E-14 | 0 | 427 | gtex_brain_putamen_basal |
| rs74813454 | 1 | 22352395 | LINC00339 | ENSG00000218510.3 | 2.211E-12 | 0 | 714 | gtex_brain_putamen_basal |
| rs141271718 | 1 | 22353345 | LINC00339 | ENSG00000218510.3 | 4.374E-7 | 0 | 1664 | gtex_brain_putamen_basal |
| rs2501291 | 1 | 22353414 | LINC00339 | ENSG00000218510.3 | 2.252E-6 | 0 | 1733 | gtex_brain_putamen_basal |
| rs2473302 | 1 | 22353824 | LINC00339 | ENSG00000218510.3 | 2.24E-6 | 0 | 2143 | gtex_brain_putamen_basal |
| rs3835733 | 1 | 22353835 | LINC00339 | ENSG00000218510.3 | 2.198E-6 | 0 | 2154 | gtex_brain_putamen_basal |
| rs2473301 | 1 | 22353874 | LINC00339 | ENSG00000218510.3 | 2.236E-6 | 0 | 2193 | gtex_brain_putamen_basal |
| rs2501290 | 1 | 22353986 | LINC00339 | ENSG00000218510.3 | 1.419E-8 | 0 | 2305 | gtex_brain_putamen_basal |
| rs2501289 | 1 | 22353989 | LINC00339 | ENSG00000218510.3 | 1.419E-8 | 0 | 2308 | gtex_brain_putamen_basal |
| rs2473300 | 1 | 22354246 | LINC00339 | ENSG00000218510.3 | 2.208E-6 | 0 | 2565 | gtex_brain_putamen_basal |
| rs2473298 | 1 | 22354277 | LINC00339 | ENSG00000218510.3 | 1.777E-6 | 0 | 2596 | gtex_brain_putamen_basal |
| rs3768571 | 1 | 22354327 | LINC00339 | ENSG00000218510.3 | 4.725E-11 | 0 | 2646 | gtex_brain_putamen_basal |
| rs2473297 | 1 | 22354537 | LINC00339 | ENSG00000218510.3 | 2.157E-6 | 0 | 2856 | gtex_brain_putamen_basal |
| rs2473296 | 1 | 22354587 | LINC00339 | ENSG00000218510.3 | 1.619E-6 | 0 | 2906 | gtex_brain_putamen_basal |
| rs2473295 | 1 | 22354866 | LINC00339 | ENSG00000218510.3 | 2.165E-6 | 0 | 3185 | gtex_brain_putamen_basal |
| rs61778044 | 1 | 22355631 | LINC00339 | ENSG00000218510.3 | 4.23E-11 | 0 | 3950 | gtex_brain_putamen_basal |
| rs2473294 | 1 | 22356098 | LINC00339 | ENSG00000218510.3 | 2.146E-6 | 0 | 4417 | gtex_brain_putamen_basal |
| rs2473293 | 1 | 22356138 | LINC00339 | ENSG00000218510.3 | 2.165E-6 | 0 | 4457 | gtex_brain_putamen_basal |
| rs2473292 | 1 | 22356155 | LINC00339 | ENSG00000218510.3 | 2.165E-6 | 0 | 4474 | gtex_brain_putamen_basal |
| rs1883421 | 1 | 22356640 | LINC00339 | ENSG00000218510.3 | 2.165E-6 | 0 | 4959 | gtex_brain_putamen_basal |
| rs1063116 | 1 | 22356809 | LINC00339 | ENSG00000218510.3 | 2.165E-6 | 0 | 5128 | gtex_brain_putamen_basal |
| rs1063117 | 1 | 22356888 | LINC00339 | ENSG00000218510.3 | 2.197E-6 | 0 | 5207 | gtex_brain_putamen_basal |
| rs1063118 | 1 | 22356905 | LINC00339 | ENSG00000218510.3 | 2.165E-6 | 0 | 5224 | gtex_brain_putamen_basal |
| rs760923 | 1 | 22357217 | LINC00339 | ENSG00000218510.3 | 2.165E-6 | 0 | 5536 | gtex_brain_putamen_basal |
| rs2473291 | 1 | 22357549 | LINC00339 | ENSG00000218510.3 | 2.165E-6 | 0 | 5868 | gtex_brain_putamen_basal |
| rs56408422 | 1 | 22360138 | LINC00339 | ENSG00000218510.3 | 3.803E-10 | 0 | 8457 | gtex_brain_putamen_basal |
| rs12748456 | 1 | 22370157 | LINC00339 | ENSG00000218510.3 | 1.65E-7 | 0 | 18476 | gtex_brain_putamen_basal |
Section II. Transcriptome annotation
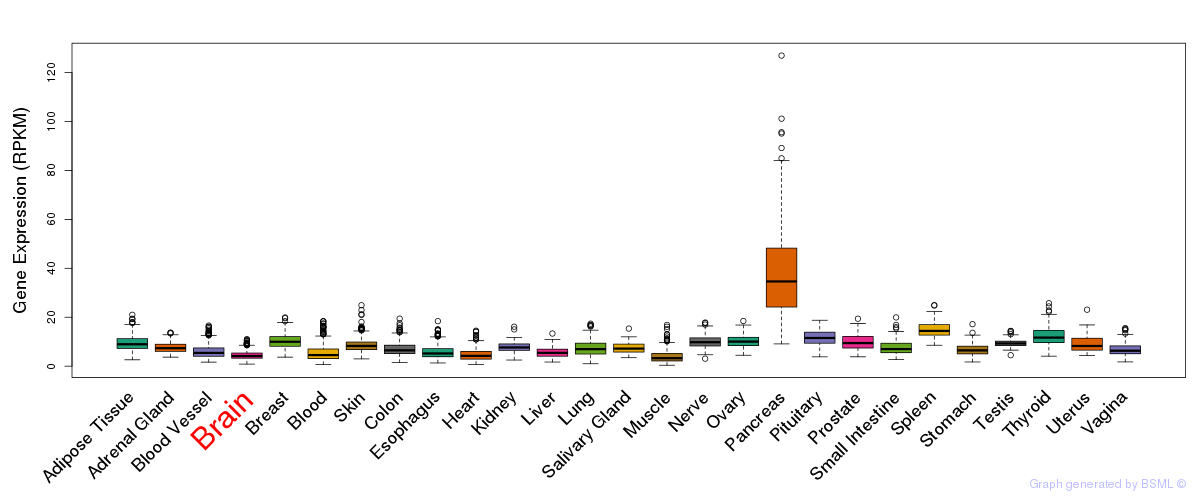
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
No co-expressed genes in brain regions
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| THUM SYSTOLIC HEART FAILURE UP | 423 | 283 | All SZGR 2.0 genes in this pathway |
| KINSEY TARGETS OF EWSR1 FLII FUSION UP | 1278 | 748 | All SZGR 2.0 genes in this pathway |
| KIM MYCN AMPLIFICATION TARGETS DN | 103 | 59 | All SZGR 2.0 genes in this pathway |
| GOERING BLOOD HDL CHOLESTEROL QTL CIS | 13 | 7 | All SZGR 2.0 genes in this pathway |
| BENPORATH CYCLING GENES | 648 | 385 | All SZGR 2.0 genes in this pathway |
| CHANDRAN METASTASIS DN | 306 | 191 | All SZGR 2.0 genes in this pathway |
| WHITFIELD CELL CYCLE S | 162 | 86 | All SZGR 2.0 genes in this pathway |