Gene Page: KRT85
Summary ?
| GeneID | 3891 |
| Symbol | KRT85 |
| Synonyms | ECTD4|HB5|Hb-5|K85|KRTHB5|hHb5 |
| Description | keratin 85 |
| Reference | MIM:602767|HGNC:HGNC:6462|Ensembl:ENSG00000135443|HPRD:04139|Vega:OTTHUMG00000169633 |
| Gene type | protein-coding |
| Map location | 12q13 |
| Pascal p-value | 0.029 |
| Fetal beta | -0.114 |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:GWASdb | Genome-wide Association Studies | GWASdb records for schizophrenia | |
| CV:PGCnp | Genome-wide Association Study | GWAS |
Section I. Genetics and epigenetics annotation
Section II. Transcriptome annotation
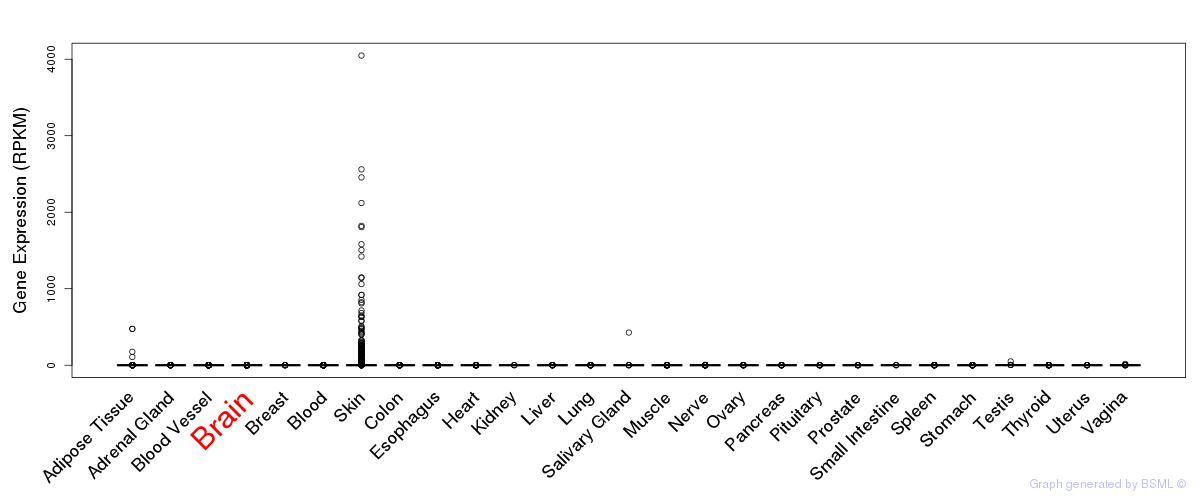
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| SIDT2 | 0.87 | 0.88 |
| DENND5A | 0.86 | 0.85 |
| TMEM189 | 0.86 | 0.84 |
| TMEM214 | 0.86 | 0.86 |
| IPO13 | 0.86 | 0.80 |
| TECPR2 | 0.84 | 0.87 |
| ABCG1 | 0.84 | 0.82 |
| TBC1D9B | 0.84 | 0.84 |
| CPOX | 0.83 | 0.80 |
| UNC5B | 0.83 | 0.82 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| SYCP3 | -0.50 | -0.46 |
| AL050337.1 | -0.48 | -0.39 |
| FAM159B | -0.48 | -0.55 |
| AF347015.21 | -0.48 | -0.33 |
| CLEC2B | -0.47 | -0.32 |
| C1orf54 | -0.46 | -0.34 |
| ST20 | -0.46 | -0.47 |
| SPDYA | -0.46 | -0.42 |
| C1orf61 | -0.45 | -0.33 |
| GNG11 | -0.44 | -0.35 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| SHEN SMARCA2 TARGETS DN | 357 | 212 | All SZGR 2.0 genes in this pathway |
| SATO SILENCED BY METHYLATION IN PANCREATIC CANCER 1 | 419 | 273 | All SZGR 2.0 genes in this pathway |
| HOWLIN CITED1 TARGETS 1 UP | 35 | 25 | All SZGR 2.0 genes in this pathway |
| MARTINEZ RB1 TARGETS UP | 673 | 430 | All SZGR 2.0 genes in this pathway |
| MARTINEZ TP53 TARGETS DN | 593 | 372 | All SZGR 2.0 genes in this pathway |
| MARTINEZ RB1 AND TP53 TARGETS DN | 591 | 366 | All SZGR 2.0 genes in this pathway |
| BRUECKNER TARGETS OF MIRLET7A3 UP | 111 | 69 | All SZGR 2.0 genes in this pathway |
| SWEET LUNG CANCER KRAS DN | 435 | 289 | All SZGR 2.0 genes in this pathway |
| MEISSNER NPC ICP WITH H3 UNMETHYLATED | 24 | 15 | All SZGR 2.0 genes in this pathway |
| MIKKELSEN MEF HCP WITH H3 UNMETHYLATED | 228 | 119 | All SZGR 2.0 genes in this pathway |
| MARTENS TRETINOIN RESPONSE UP | 857 | 456 | All SZGR 2.0 genes in this pathway |
| BRUINS UVC RESPONSE VIA TP53 GROUP A | 898 | 516 | All SZGR 2.0 genes in this pathway |