Gene Page: URGCP
Summary ?
| GeneID | 55665 |
| Symbol | URGCP |
| Synonyms | URG4 |
| Description | upregulator of cell proliferation |
| Reference | MIM:610337|HGNC:HGNC:30890|Ensembl:ENSG00000106608|HPRD:11663|Vega:OTTHUMG00000155245 |
| Gene type | protein-coding |
| Map location | 7p13 |
| Pascal p-value | 0.814 |
| Sherlock p-value | 0.489 |
| Fetal beta | 0.514 |
| eGene | Caudate basal ganglia Cortex |
| Support | Ascano FMRP targets |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:GWASdb | Genome-wide Association Studies | GWASdb records for schizophrenia | |
| CV:PGCnp | Genome-wide Association Study | GWAS |
Section I. Genetics and epigenetics annotation
Section II. Transcriptome annotation
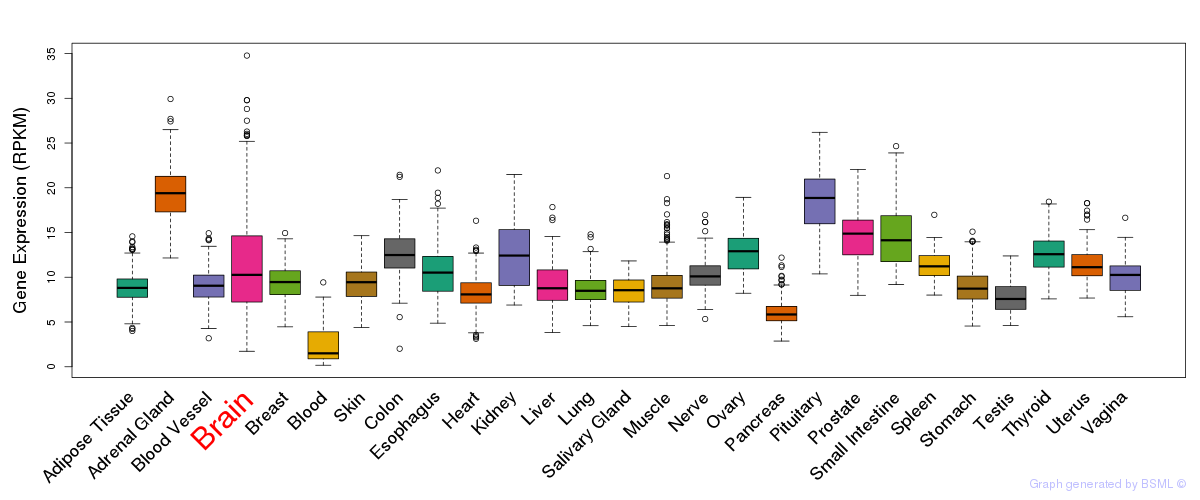
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| ZNF836 | 0.95 | 0.94 |
| ZBTB8A | 0.94 | 0.92 |
| ZFP37 | 0.94 | 0.92 |
| ZNF3 | 0.94 | 0.88 |
| TMEM57 | 0.94 | 0.94 |
| EPC1 | 0.93 | 0.90 |
| GMEB1 | 0.93 | 0.91 |
| ZNF551 | 0.93 | 0.88 |
| ZNF564 | 0.93 | 0.92 |
| ZFP30 | 0.93 | 0.92 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| HLA-F | -0.76 | -0.81 |
| AIFM3 | -0.75 | -0.82 |
| ALDOC | -0.74 | -0.77 |
| FBXO2 | -0.73 | -0.71 |
| TSC22D4 | -0.73 | -0.83 |
| PTH1R | -0.73 | -0.77 |
| AF347015.27 | -0.73 | -0.90 |
| AF347015.31 | -0.73 | -0.90 |
| C5orf53 | -0.73 | -0.72 |
| CLU | -0.72 | -0.75 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| IVANOVA HEMATOPOIESIS STEM CELL LONG TERM | 302 | 191 | All SZGR 2.0 genes in this pathway |
| CHENG RESPONSE TO NICKEL ACETATE | 45 | 29 | All SZGR 2.0 genes in this pathway |
| DOUGLAS BMI1 TARGETS UP | 566 | 371 | All SZGR 2.0 genes in this pathway |
| MONNIER POSTRADIATION TUMOR ESCAPE UP | 393 | 244 | All SZGR 2.0 genes in this pathway |
| ACEVEDO LIVER CANCER DN | 540 | 340 | All SZGR 2.0 genes in this pathway |
| FEVR CTNNB1 TARGETS UP | 682 | 433 | All SZGR 2.0 genes in this pathway |
| KRIEG KDM3A TARGETS NOT HYPOXIA | 208 | 107 | All SZGR 2.0 genes in this pathway |
| ZWANG CLASS 1 TRANSIENTLY INDUCED BY EGF | 516 | 308 | All SZGR 2.0 genes in this pathway |