Gene Page: NPLOC4
Summary ?
| GeneID | 55666 |
| Symbol | NPLOC4 |
| Synonyms | NPL4 |
| Description | NPL4 homolog, ubiquitin recognition factor |
| Reference | MIM:606590|HGNC:HGNC:18261|Ensembl:ENSG00000182446|HPRD:06975|Vega:OTTHUMG00000177990 |
| Gene type | protein-coding |
| Map location | 17qter |
| Pascal p-value | 0.057 |
| Sherlock p-value | 0.02 |
| Fetal beta | -0.845 |
| eGene | Cortex |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS |
Section I. Genetics and epigenetics annotation
Section II. Transcriptome annotation
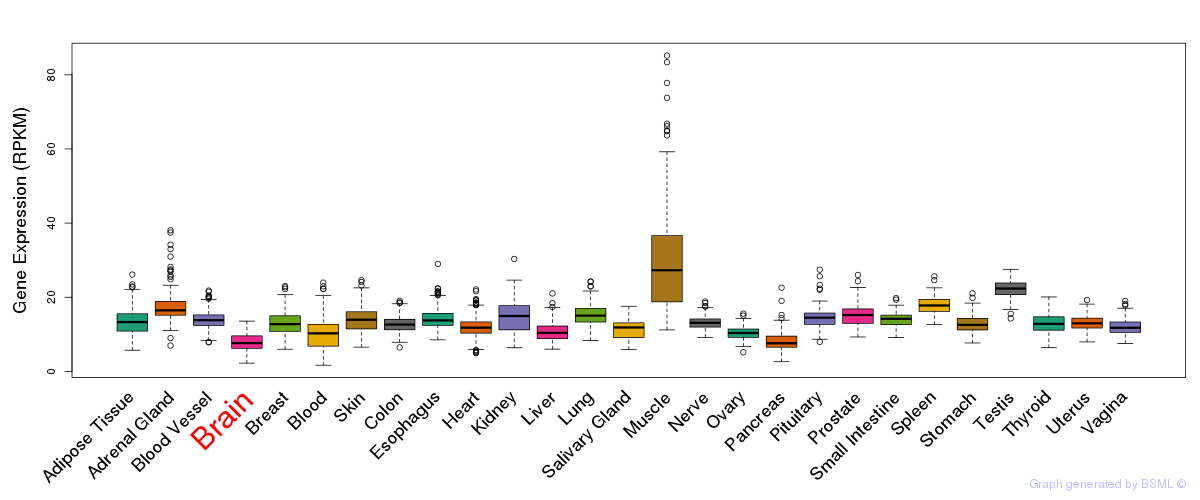
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| ZNF821 | 0.93 | 0.92 |
| U2AF1 | 0.93 | 0.90 |
| NAGK | 0.92 | 0.94 |
| TRA2A | 0.92 | 0.90 |
| RPAIN | 0.92 | 0.91 |
| PDAP1 | 0.91 | 0.92 |
| ZSCAN21 | 0.91 | 0.88 |
| BAT1 | 0.91 | 0.92 |
| ZNF311 | 0.91 | 0.87 |
| EWSR1 | 0.91 | 0.91 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| HLA-F | -0.75 | -0.77 |
| AF347015.27 | -0.74 | -0.89 |
| AF347015.31 | -0.74 | -0.87 |
| AIFM3 | -0.73 | -0.76 |
| AF347015.33 | -0.73 | -0.88 |
| ALDOC | -0.72 | -0.73 |
| MT-CO2 | -0.72 | -0.86 |
| TSC22D4 | -0.72 | -0.78 |
| MT-CYB | -0.72 | -0.87 |
| C5orf53 | -0.71 | -0.71 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| NAKAMURA TUMOR ZONE PERIPHERAL VS CENTRAL UP | 285 | 181 | All SZGR 2.0 genes in this pathway |
| HEIDENBLAD AMPLICON 8Q24 UP | 40 | 23 | All SZGR 2.0 genes in this pathway |
| BENPORATH OCT4 TARGETS | 290 | 172 | All SZGR 2.0 genes in this pathway |
| NIKOLSKY BREAST CANCER 17Q21 Q25 AMPLICON | 335 | 181 | All SZGR 2.0 genes in this pathway |
| DURCHDEWALD SKIN CARCINOGENESIS DN | 264 | 168 | All SZGR 2.0 genes in this pathway |
| MONNIER POSTRADIATION TUMOR ESCAPE UP | 393 | 244 | All SZGR 2.0 genes in this pathway |