Gene Page: REG1B
Summary ?
| GeneID | 5968 |
| Symbol | REG1B |
| Synonyms | PSPS2|REGH|REGI-BETA|REGL |
| Description | regenerating family member 1 beta |
| Reference | MIM:167771|HGNC:HGNC:9952|Ensembl:ENSG00000172023|HPRD:01339|Vega:OTTHUMG00000130019 |
| Gene type | protein-coding |
| Map location | 2p12 |
| Pascal p-value | 0.006 |
| Fetal beta | -0.169 |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:GWASdb | Genome-wide Association Studies | GWASdb records for schizophrenia | |
| CV:PGCnp | Genome-wide Association Study | GWAS |
Section I. Genetics and epigenetics annotation
Section II. Transcriptome annotation
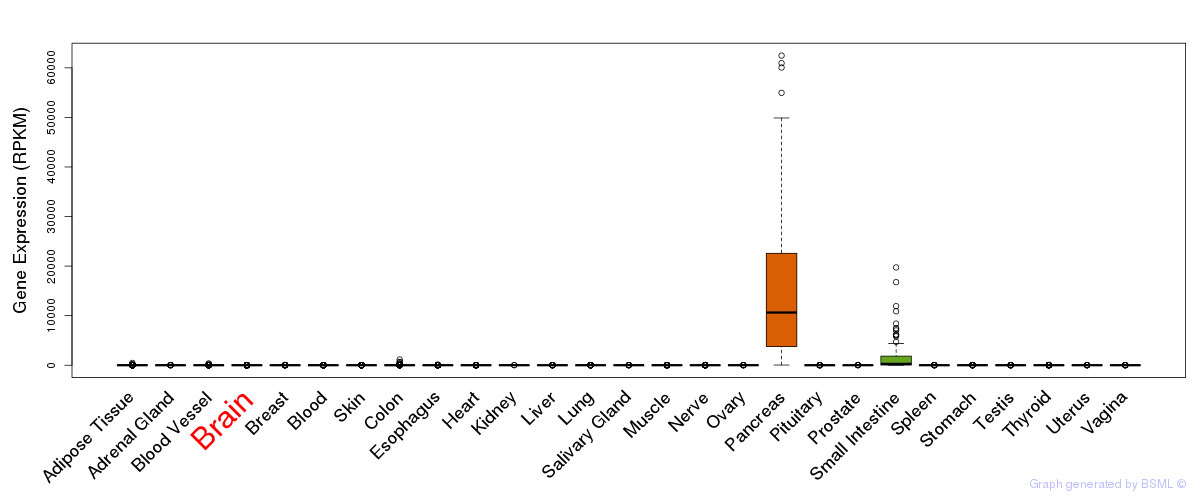
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| IQWD1 | 0.85 | 0.85 |
| PPP3R1 | 0.83 | 0.88 |
| GLRB | 0.83 | 0.88 |
| SCG5 | 0.83 | 0.84 |
| SUCLA2 | 0.83 | 0.85 |
| TSNAX | 0.83 | 0.83 |
| PPP3CB | 0.83 | 0.83 |
| SMAP2 | 0.83 | 0.83 |
| NCKAP1 | 0.82 | 0.80 |
| PCMT1 | 0.82 | 0.80 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| AF347015.18 | -0.55 | -0.39 |
| EFEMP2 | -0.52 | -0.55 |
| AF347015.21 | -0.51 | -0.28 |
| AF347015.26 | -0.51 | -0.31 |
| AL139819.3 | -0.51 | -0.43 |
| RAB34 | -0.50 | -0.54 |
| NFKB2 | -0.50 | -0.49 |
| SLC2A4RG | -0.50 | -0.50 |
| AP002478.3 | -0.50 | -0.45 |
| AF347015.2 | -0.50 | -0.29 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| WATANABE RECTAL CANCER RADIOTHERAPY RESPONSIVE UP | 108 | 67 | All SZGR 2.0 genes in this pathway |
| ZHOU INFLAMMATORY RESPONSE LIVE DN | 384 | 220 | All SZGR 2.0 genes in this pathway |
| SABATES COLORECTAL ADENOMA UP | 141 | 75 | All SZGR 2.0 genes in this pathway |
| GAUSSMANN MLL AF4 FUSION TARGETS G UP | 238 | 135 | All SZGR 2.0 genes in this pathway |
| RASHI RESPONSE TO IONIZING RADIATION 1 | 45 | 27 | All SZGR 2.0 genes in this pathway |
| IGLESIAS E2F TARGETS DN | 16 | 11 | All SZGR 2.0 genes in this pathway |
| IVANOVA HEMATOPOIESIS STEM CELL AND PROGENITOR | 681 | 420 | All SZGR 2.0 genes in this pathway |
| FEVR CTNNB1 TARGETS UP | 682 | 433 | All SZGR 2.0 genes in this pathway |
| NABA ECM AFFILIATED | 171 | 89 | All SZGR 2.0 genes in this pathway |
| NABA MATRISOME ASSOCIATED | 753 | 411 | All SZGR 2.0 genes in this pathway |
| NABA MATRISOME | 1028 | 559 | All SZGR 2.0 genes in this pathway |