Gene Page: GUF1
Summary ?
| GeneID | 60558 |
| Symbol | GUF1 |
| Synonyms | EF-4 |
| Description | GUF1 homolog, GTPase |
| Reference | HGNC:HGNC:25799|Ensembl:ENSG00000151806|HPRD:07821|Vega:OTTHUMG00000128608 |
| Gene type | protein-coding |
| Map location | 4p12 |
| Pascal p-value | 0.326 |
| Fetal beta | 0.468 |
| DMG | 1 (# studies) |
| eGene | Anterior cingulate cortex BA24 Caudate basal ganglia Cerebellar Hemisphere Cerebellum Cortex Frontal Cortex BA9 Hippocampus Hypothalamus Nucleus accumbens basal ganglia Putamen basal ganglia Myers' cis & trans Meta |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| DMG:Jaffe_2016 | Genome-wide DNA methylation analysis | This dataset includes 2,104 probes/CpGs associated with SZ patients (n=108) compared to 136 controls at Bonferroni-adjusted P < 0.05. | 1 |
Section I. Genetics and epigenetics annotation
 Differentially methylated gene
Differentially methylated gene
| Probe | Chromosome | Position | Nearest gene | P (dis) | Beta (dis) | FDR (dis) | Study |
|---|---|---|---|---|---|---|---|
| cg19516485 | 4 | 44680377 | GUF1 | 9.31E-10 | -0.011 | 1.11E-6 | DMG:Jaffe_2016 |
 eQTL annotation
eQTL annotation
| SNP ID | Chromosome | Position | eGene | Gene Entrez ID | pvalue | qvalue | TSS distance | eQTL type |
|---|---|---|---|---|---|---|---|---|
| rs17600797 | chr4 | 44626805 | GUF1 | 60558 | 5.893E-7 | cis | ||
| rs6850107 | chr4 | 44633865 | GUF1 | 60558 | 7.124E-6 | cis | ||
| rs10026088 | chr4 | 44744986 | GUF1 | 60558 | 0.07 | cis | ||
| rs11729477 | chr4 | 44770332 | GUF1 | 60558 | 9.095E-4 | cis | ||
| rs1580610 | chr4 | 44775014 | GUF1 | 60558 | 0.12 | cis | ||
| rs4145971 | chr4 | 44843213 | GUF1 | 60558 | 5.436E-7 | cis | ||
| rs10938369 | chr4 | 44860188 | GUF1 | 60558 | 0.16 | cis | ||
| rs6447385 | chr4 | 44880089 | GUF1 | 60558 | 0.15 | cis | ||
| rs17600797 | chr4 | 44626805 | GUF1 | 60558 | 1.151E-4 | trans | ||
| rs6850107 | chr4 | 44633865 | GUF1 | 60558 | 0 | trans | ||
| rs11729477 | chr4 | 44770332 | GUF1 | 60558 | 0.07 | trans | ||
| rs4145971 | chr4 | 44843213 | GUF1 | 60558 | 1.073E-4 | trans | ||
| rs16857236 | 4 | 44614810 | GUF1 | ENSG00000151806.9 | 1.405E-6 | 0 | -67648 | gtex_brain_ba24 |
| rs1439364 | 4 | 44615265 | GUF1 | ENSG00000151806.9 | 6.418E-7 | 0 | -67193 | gtex_brain_ba24 |
| rs11736925 | 4 | 44616283 | GUF1 | ENSG00000151806.9 | 6.483E-7 | 0 | -66175 | gtex_brain_ba24 |
| rs11734057 | 4 | 44616296 | GUF1 | ENSG00000151806.9 | 6.45E-7 | 0 | -66162 | gtex_brain_ba24 |
| rs17600797 | 4 | 44626806 | GUF1 | ENSG00000151806.9 | 8.083E-8 | 0 | -55652 | gtex_brain_ba24 |
| rs6831166 | 4 | 44630977 | GUF1 | ENSG00000151806.9 | 6.143E-9 | 0 | -51481 | gtex_brain_ba24 |
| rs6850107 | 4 | 44633866 | GUF1 | ENSG00000151806.9 | 2.592E-9 | 0 | -48592 | gtex_brain_ba24 |
| rs6813944 | 4 | 44635157 | GUF1 | ENSG00000151806.9 | 2.461E-9 | 0 | -47301 | gtex_brain_ba24 |
| rs10938361 | 4 | 44638931 | GUF1 | ENSG00000151806.9 | 4.813E-10 | 0 | -43527 | gtex_brain_ba24 |
| rs7679737 | 4 | 44639517 | GUF1 | ENSG00000151806.9 | 6.05E-10 | 0 | -42941 | gtex_brain_ba24 |
| rs4235132 | 4 | 44641392 | GUF1 | ENSG00000151806.9 | 6.01E-10 | 0 | -41066 | gtex_brain_ba24 |
| rs34018268 | 4 | 44644463 | GUF1 | ENSG00000151806.9 | 5.94E-10 | 0 | -37995 | gtex_brain_ba24 |
| rs12646633 | 4 | 44644687 | GUF1 | ENSG00000151806.9 | 5.94E-10 | 0 | -37771 | gtex_brain_ba24 |
| rs6852372 | 4 | 44651820 | GUF1 | ENSG00000151806.9 | 5.852E-10 | 0 | -30638 | gtex_brain_ba24 |
| rs1388310 | 4 | 44653414 | GUF1 | ENSG00000151806.9 | 3.714E-9 | 0 | -29044 | gtex_brain_ba24 |
| rs369684976 | 4 | 44657868 | GUF1 | ENSG00000151806.9 | 8.614E-10 | 0 | -24590 | gtex_brain_ba24 |
| rs12639722 | 4 | 44660248 | GUF1 | ENSG00000151806.9 | 5.852E-10 | 0 | -22210 | gtex_brain_ba24 |
| rs62305366 | 4 | 44664907 | GUF1 | ENSG00000151806.9 | 5.852E-10 | 0 | -17551 | gtex_brain_ba24 |
| rs35372782 | 4 | 44667338 | GUF1 | ENSG00000151806.9 | 5.852E-10 | 0 | -15120 | gtex_brain_ba24 |
| rs62305368 | 4 | 44673826 | GUF1 | ENSG00000151806.9 | 5.793E-10 | 0 | -8632 | gtex_brain_ba24 |
| rs200576312 | 4 | 44677677 | GUF1 | ENSG00000151806.9 | 5.703E-10 | 0 | -4781 | gtex_brain_ba24 |
| rs75324498 | 4 | 44677678 | GUF1 | ENSG00000151806.9 | 5.852E-10 | 0 | -4780 | gtex_brain_ba24 |
| rs10461070 | 4 | 44678737 | GUF1 | ENSG00000151806.9 | 5.852E-10 | 0 | -3721 | gtex_brain_ba24 |
| rs11734296 | 4 | 44679560 | GUF1 | ENSG00000151806.9 | 5.852E-10 | 0 | -2898 | gtex_brain_ba24 |
| rs62305372 | 4 | 44683541 | GUF1 | ENSG00000151806.9 | 5.852E-10 | 0 | 1083 | gtex_brain_ba24 |
| rs11931747 | 4 | 44687163 | GUF1 | ENSG00000151806.9 | 5.852E-10 | 0 | 4705 | gtex_brain_ba24 |
| rs12499960 | 4 | 44687401 | GUF1 | ENSG00000151806.9 | 3.122E-9 | 0 | 4943 | gtex_brain_ba24 |
| rs12639842 | 4 | 44691098 | GUF1 | ENSG00000151806.9 | 5.852E-10 | 0 | 8640 | gtex_brain_ba24 |
| rs71611910 | 4 | 44693499 | GUF1 | ENSG00000151806.9 | 5.852E-10 | 0 | 11041 | gtex_brain_ba24 |
| rs2306992 | 4 | 44694077 | GUF1 | ENSG00000151806.9 | 5.852E-10 | 0 | 11619 | gtex_brain_ba24 |
| rs201814154 | 4 | 44695520 | GUF1 | ENSG00000151806.9 | 5.763E-10 | 0 | 13062 | gtex_brain_ba24 |
| rs77297867 | 4 | 44695521 | GUF1 | ENSG00000151806.9 | 4.17E-10 | 0 | 13063 | gtex_brain_ba24 |
| rs13125773 | 4 | 44701256 | GUF1 | ENSG00000151806.9 | 9.136E-10 | 0 | 18798 | gtex_brain_ba24 |
| rs1565532 | 4 | 44702720 | GUF1 | ENSG00000151806.9 | 5.833E-10 | 0 | 20262 | gtex_brain_ba24 |
| rs16857402 | 4 | 44706453 | GUF1 | ENSG00000151806.9 | 5.769E-10 | 0 | 23995 | gtex_brain_ba24 |
| rs2709 | 4 | 44706919 | GUF1 | ENSG00000151806.9 | 5.852E-10 | 0 | 24461 | gtex_brain_ba24 |
| rs62305383 | 4 | 44708086 | GUF1 | ENSG00000151806.9 | 5.852E-10 | 0 | 25628 | gtex_brain_ba24 |
| rs71648801 | 4 | 44723646 | GUF1 | ENSG00000151806.9 | 4.082E-9 | 0 | 41188 | gtex_brain_ba24 |
| rs11945975 | 4 | 44724566 | GUF1 | ENSG00000151806.9 | 9.135E-10 | 0 | 42108 | gtex_brain_ba24 |
| rs11724396 | 4 | 44725334 | GUF1 | ENSG00000151806.9 | 5.852E-10 | 0 | 42876 | gtex_brain_ba24 |
| rs16857442 | 4 | 44739911 | GUF1 | ENSG00000151806.9 | 5.301E-7 | 0 | 57453 | gtex_brain_ba24 |
| rs1491300 | 4 | 44742727 | GUF1 | ENSG00000151806.9 | 5.317E-7 | 0 | 60269 | gtex_brain_ba24 |
| rs138839217 | 4 | 44747451 | GUF1 | ENSG00000151806.9 | 4.792E-7 | 0 | 64993 | gtex_brain_ba24 |
| rs112680663 | 4 | 44749297 | GUF1 | ENSG00000151806.9 | 5.334E-7 | 0 | 66839 | gtex_brain_ba24 |
| rs13112967 | 4 | 44750803 | GUF1 | ENSG00000151806.9 | 4.795E-7 | 0 | 68345 | gtex_brain_ba24 |
| rs11934131 | 4 | 44761571 | GUF1 | ENSG00000151806.9 | 5.334E-7 | 0 | 79113 | gtex_brain_ba24 |
| rs7675035 | 4 | 44764783 | GUF1 | ENSG00000151806.9 | 5.334E-7 | 0 | 82325 | gtex_brain_ba24 |
| rs12644039 | 4 | 44766644 | GUF1 | ENSG00000151806.9 | 3.939E-7 | 0 | 84186 | gtex_brain_ba24 |
| rs35152931 | 4 | 44766932 | GUF1 | ENSG00000151806.9 | 5.334E-7 | 0 | 84474 | gtex_brain_ba24 |
| rs11729477 | 4 | 44770333 | GUF1 | ENSG00000151806.9 | 5.334E-7 | 0 | 87875 | gtex_brain_ba24 |
| rs16857479 | 4 | 44770720 | GUF1 | ENSG00000151806.9 | 5.334E-7 | 0 | 88262 | gtex_brain_ba24 |
| rs11729736 | 4 | 44778190 | GUF1 | ENSG00000151806.9 | 5.212E-7 | 0 | 95732 | gtex_brain_ba24 |
| rs11724366 | 4 | 44785512 | GUF1 | ENSG00000151806.9 | 4.893E-7 | 0 | 103054 | gtex_brain_ba24 |
| rs6816989 | 4 | 44791911 | GUF1 | ENSG00000151806.9 | 5.643E-7 | 0 | 109453 | gtex_brain_ba24 |
| rs71984973 | 4 | 44793288 | GUF1 | ENSG00000151806.9 | 6.893E-7 | 0 | 110830 | gtex_brain_ba24 |
| rs11728530 | 4 | 44794111 | GUF1 | ENSG00000151806.9 | 5.526E-7 | 0 | 111653 | gtex_brain_ba24 |
| rs36047545 | 4 | 44795072 | GUF1 | ENSG00000151806.9 | 5.544E-7 | 0 | 112614 | gtex_brain_ba24 |
| rs7683933 | 4 | 44798080 | GUF1 | ENSG00000151806.9 | 4.923E-7 | 0 | 115622 | gtex_brain_ba24 |
| rs12639624 | 4 | 44801988 | GUF1 | ENSG00000151806.9 | 5.334E-7 | 0 | 119530 | gtex_brain_ba24 |
| rs57068467 | 4 | 44806404 | GUF1 | ENSG00000151806.9 | 5.334E-7 | 0 | 123946 | gtex_brain_ba24 |
| rs12639682 | 4 | 44811032 | GUF1 | ENSG00000151806.9 | 5.338E-7 | 0 | 128574 | gtex_brain_ba24 |
| rs36022022 | 4 | 44818450 | GUF1 | ENSG00000151806.9 | 5.347E-7 | 0 | 135992 | gtex_brain_ba24 |
| rs11730341 | 4 | 44823800 | GUF1 | ENSG00000151806.9 | 5.352E-7 | 0 | 141342 | gtex_brain_ba24 |
| rs34521789 | 4 | 44824918 | GUF1 | ENSG00000151806.9 | 5.352E-7 | 0 | 142460 | gtex_brain_ba24 |
| rs62409354 | 4 | 44825505 | GUF1 | ENSG00000151806.9 | 5.352E-7 | 0 | 143047 | gtex_brain_ba24 |
| rs35765022 | 4 | 44827368 | GUF1 | ENSG00000151806.9 | 4.024E-7 | 0 | 144910 | gtex_brain_ba24 |
| rs11734975 | 4 | 44827537 | GUF1 | ENSG00000151806.9 | 5.844E-7 | 0 | 145079 | gtex_brain_ba24 |
| rs71816407 | 4 | 44832654 | GUF1 | ENSG00000151806.9 | 6.01E-7 | 0 | 150196 | gtex_brain_ba24 |
| rs12642145 | 4 | 44834738 | GUF1 | ENSG00000151806.9 | 5.348E-7 | 0 | 152280 | gtex_brain_ba24 |
| rs143320673 | 4 | 44835036 | GUF1 | ENSG00000151806.9 | 5.137E-7 | 0 | 152578 | gtex_brain_ba24 |
| rs924176 | 4 | 44842838 | GUF1 | ENSG00000151806.9 | 5.37E-7 | 0 | 160380 | gtex_brain_ba24 |
| rs4145971 | 4 | 44843214 | GUF1 | ENSG00000151806.9 | 5.37E-7 | 0 | 160756 | gtex_brain_ba24 |
| rs6813425 | 4 | 44661520 | GUF1 | ENSG00000151806.9 | 2.343E-6 | 0.04 | -20938 | gtex_brain_putamen_basal |
| rs11732880 | 4 | 44675514 | GUF1 | ENSG00000151806.9 | 1.927E-6 | 0.04 | -6944 | gtex_brain_putamen_basal |
| rs397788506 | 4 | 44704790 | GUF1 | ENSG00000151806.9 | 2.648E-6 | 0.04 | 22332 | gtex_brain_putamen_basal |
| rs3046153 | 4 | 44727783 | GUF1 | ENSG00000151806.9 | 1.927E-6 | 0.04 | 45325 | gtex_brain_putamen_basal |
Section II. Transcriptome annotation
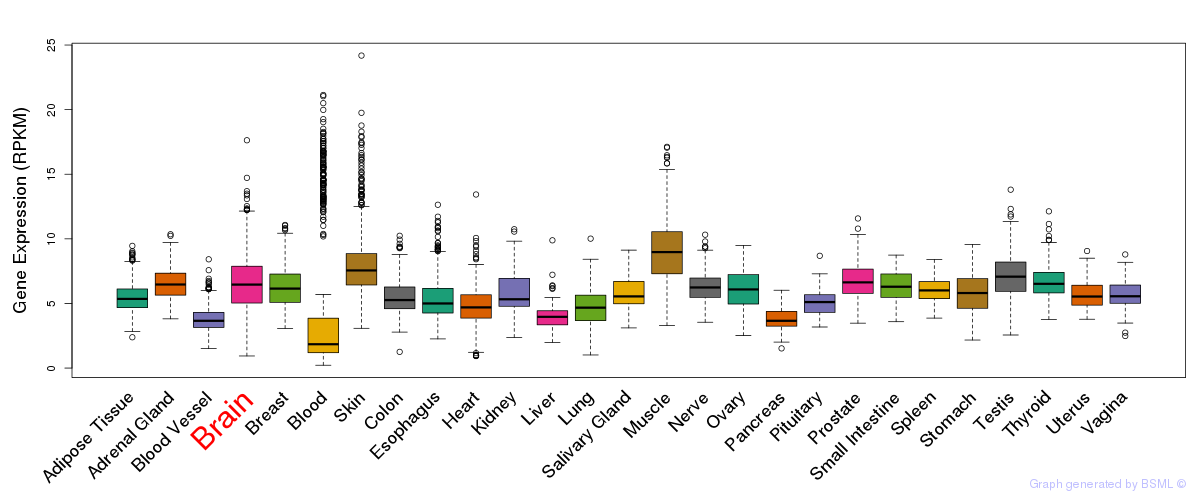
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
No co-expressed genes in brain regions
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| RODRIGUES THYROID CARCINOMA POORLY DIFFERENTIATED UP | 633 | 376 | All SZGR 2.0 genes in this pathway |
| DAIRKEE TERT TARGETS DN | 124 | 79 | All SZGR 2.0 genes in this pathway |
| BYSTRYKH HEMATOPOIESIS STEM CELL QTL TRANS | 882 | 572 | All SZGR 2.0 genes in this pathway |
| IVANOVA HEMATOPOIESIS STEM CELL SHORT TERM | 32 | 15 | All SZGR 2.0 genes in this pathway |