Gene Page: ACOT6
Summary ?
| GeneID | 641372 |
| Symbol | ACOT6 |
| Synonyms | C14orf42|c14_5530 |
| Description | acyl-CoA thioesterase 6 |
| Reference | MIM:614267|HGNC:HGNC:33159|Ensembl:ENSG00000205669|Vega:OTTHUMG00000171609 |
| Gene type | protein-coding |
| Map location | 14q24.3 |
| Pascal p-value | 0.226 |
| eGene | Anterior cingulate cortex BA24 Caudate basal ganglia Cortex Frontal Cortex BA9 Nucleus accumbens basal ganglia Putamen basal ganglia |
| Support | CompositeSet |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| DNM:Xu_2012 | Whole Exome Sequencing analysis | De novo mutations of 4 genes were identified by exome sequencing of 795 samples in this study |
Section I. Genetics and epigenetics annotation
 DNM table
DNM table
| Gene | Chromosome | Position | Ref | Alt | Transcript | AA change | Mutation type | Sift | CG46 | Trait | Study |
|---|---|---|---|---|---|---|---|---|---|---|---|
| ACOT6 | chr14 | 74086428 | A | G | NM_001037162 | p.170Y>C | missense | Schizophrenia | DNM:Xu_2012 |
 eQTL annotation
eQTL annotation
| SNP ID | Chromosome | Position | eGene | Gene Entrez ID | pvalue | qvalue | TSS distance | eQTL type |
|---|---|---|---|---|---|---|---|---|
| rs10150828 | 14 | 74077485 | ACOT6 | ENSG00000205669.2 | 1.423E-6 | 0.01 | -164 | gtex_brain_ba24 |
| rs12587206 | 14 | 74079486 | ACOT6 | ENSG00000205669.2 | 1.164E-6 | 0.01 | 1837 | gtex_brain_ba24 |
| rs11620812 | 14 | 74081365 | ACOT6 | ENSG00000205669.2 | 1.057E-6 | 0.01 | 3716 | gtex_brain_ba24 |
| rs10136341 | 14 | 74083456 | ACOT6 | ENSG00000205669.2 | 1.08E-6 | 0.01 | 5807 | gtex_brain_ba24 |
| rs5809620 | 14 | 74084619 | ACOT6 | ENSG00000205669.2 | 4.887E-7 | 0.01 | 6970 | gtex_brain_ba24 |
| rs58429599 | 14 | 74085327 | ACOT6 | ENSG00000205669.2 | 1.08E-6 | 0.01 | 7678 | gtex_brain_ba24 |
| rs7156422 | 14 | 74087563 | ACOT6 | ENSG00000205669.2 | 1.084E-6 | 0.01 | 9914 | gtex_brain_ba24 |
| rs12436753 | 14 | 74087919 | ACOT6 | ENSG00000205669.2 | 1.112E-6 | 0.01 | 10270 | gtex_brain_ba24 |
| rs7202 | 14 | 73742281 | ACOT6 | ENSG00000205669.2 | 6.042E-7 | 0 | -335368 | gtex_brain_putamen_basal |
| rs10137126 | 14 | 74025873 | ACOT6 | ENSG00000205669.2 | 4.067E-7 | 0 | -51776 | gtex_brain_putamen_basal |
| rs201125749 | 14 | 74043640 | ACOT6 | ENSG00000205669.2 | 5.843E-7 | 0 | -34009 | gtex_brain_putamen_basal |
| rs56013842 | 14 | 74043641 | ACOT6 | ENSG00000205669.2 | 3.541E-7 | 0 | -34008 | gtex_brain_putamen_basal |
| rs11620997 | 14 | 74044260 | ACOT6 | ENSG00000205669.2 | 3.281E-7 | 0 | -33389 | gtex_brain_putamen_basal |
| rs10144816 | 14 | 74045907 | ACOT6 | ENSG00000205669.2 | 3.872E-7 | 0 | -31742 | gtex_brain_putamen_basal |
| rs28410246 | 14 | 74046219 | ACOT6 | ENSG00000205669.2 | 1.598E-7 | 0 | -31430 | gtex_brain_putamen_basal |
| rs11159027 | 14 | 74047516 | ACOT6 | ENSG00000205669.2 | 2.85E-7 | 0 | -30133 | gtex_brain_putamen_basal |
| rs10143135 | 14 | 74049986 | ACOT6 | ENSG00000205669.2 | 3.375E-7 | 0 | -27663 | gtex_brain_putamen_basal |
| rs3742819 | 14 | 74058832 | ACOT6 | ENSG00000205669.2 | 8.79E-7 | 0 | -18817 | gtex_brain_putamen_basal |
| rs2009883 | 14 | 74061457 | ACOT6 | ENSG00000205669.2 | 4.163E-8 | 0 | -16192 | gtex_brain_putamen_basal |
| rs11624125 | 14 | 74064707 | ACOT6 | ENSG00000205669.2 | 4.799E-8 | 0 | -12942 | gtex_brain_putamen_basal |
| rs10138579 | 14 | 74068301 | ACOT6 | ENSG00000205669.2 | 4.108E-8 | 0 | -9348 | gtex_brain_putamen_basal |
| rs199680410 | 14 | 74069787 | ACOT6 | ENSG00000205669.2 | 1.661E-9 | 0 | -7862 | gtex_brain_putamen_basal |
| rs12590899 | 14 | 74070844 | ACOT6 | ENSG00000205669.2 | 4.108E-8 | 0 | -6805 | gtex_brain_putamen_basal |
| rs11159029 | 14 | 74071251 | ACOT6 | ENSG00000205669.2 | 6.376E-9 | 0 | -6398 | gtex_brain_putamen_basal |
| rs10137762 | 14 | 74073461 | ACOT6 | ENSG00000205669.2 | 4.109E-8 | 0 | -4188 | gtex_brain_putamen_basal |
| rs7156378 | 14 | 74073516 | ACOT6 | ENSG00000205669.2 | 4.109E-8 | 0 | -4133 | gtex_brain_putamen_basal |
| rs12323653 | 14 | 74074391 | ACOT6 | ENSG00000205669.2 | 4.072E-8 | 0 | -3258 | gtex_brain_putamen_basal |
| rs10141401 | 14 | 74074804 | ACOT6 | ENSG00000205669.2 | 2.777E-9 | 0 | -2845 | gtex_brain_putamen_basal |
| rs12590145 | 14 | 74076794 | ACOT6 | ENSG00000205669.2 | 2.479E-8 | 0 | -855 | gtex_brain_putamen_basal |
| rs10150828 | 14 | 74077485 | ACOT6 | ENSG00000205669.2 | 1.348E-8 | 0 | -164 | gtex_brain_putamen_basal |
| rs12587206 | 14 | 74079486 | ACOT6 | ENSG00000205669.2 | 3.606E-9 | 0 | 1837 | gtex_brain_putamen_basal |
| rs11620812 | 14 | 74081365 | ACOT6 | ENSG00000205669.2 | 3.728E-9 | 0 | 3716 | gtex_brain_putamen_basal |
| rs10136341 | 14 | 74083456 | ACOT6 | ENSG00000205669.2 | 3.775E-9 | 0 | 5807 | gtex_brain_putamen_basal |
| rs12891009 | 14 | 74083846 | ACOT6 | ENSG00000205669.2 | 2.479E-8 | 0 | 6197 | gtex_brain_putamen_basal |
| rs5809620 | 14 | 74084619 | ACOT6 | ENSG00000205669.2 | 4.108E-9 | 0 | 6970 | gtex_brain_putamen_basal |
| rs58429599 | 14 | 74085327 | ACOT6 | ENSG00000205669.2 | 3.775E-9 | 0 | 7678 | gtex_brain_putamen_basal |
| rs11626046 | 14 | 74085707 | ACOT6 | ENSG00000205669.2 | 4.072E-8 | 0 | 8058 | gtex_brain_putamen_basal |
| rs7156422 | 14 | 74087563 | ACOT6 | ENSG00000205669.2 | 3.741E-9 | 0 | 9914 | gtex_brain_putamen_basal |
| rs12436753 | 14 | 74087919 | ACOT6 | ENSG00000205669.2 | 3.705E-9 | 0 | 10270 | gtex_brain_putamen_basal |
| rs12878358 | 14 | 74088234 | ACOT6 | ENSG00000205669.2 | 2.293E-8 | 0 | 10585 | gtex_brain_putamen_basal |
| rs10134911 | 14 | 74095256 | ACOT6 | ENSG00000205669.2 | 1.533E-7 | 0 | 17607 | gtex_brain_putamen_basal |
| rs4899474 | 14 | 74096603 | ACOT6 | ENSG00000205669.2 | 6.633E-7 | 0 | 18954 | gtex_brain_putamen_basal |
Section II. Transcriptome annotation
General gene expression (GTEx)
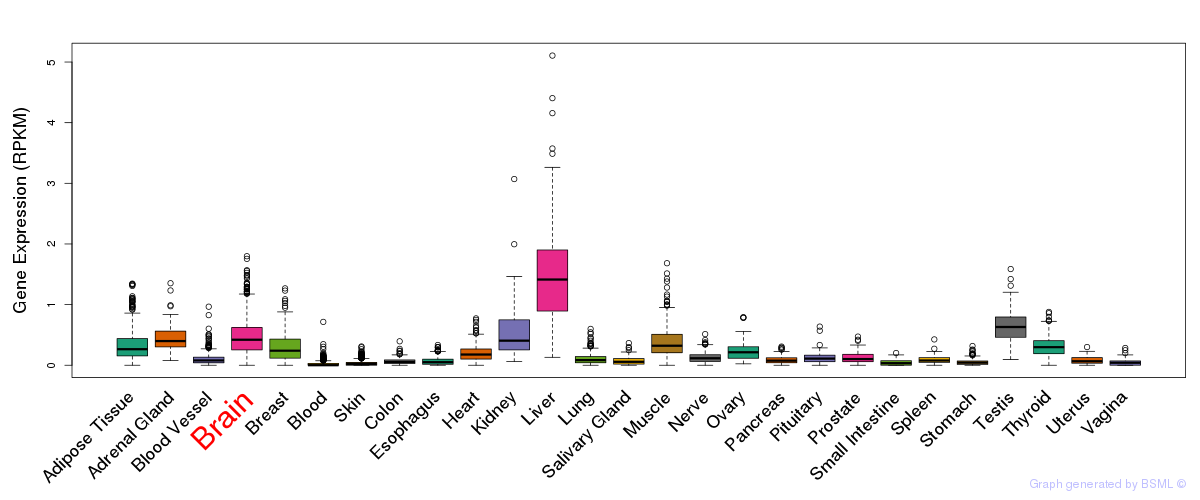
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
No co-expressed genes in brain regions
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| ZHOU INFLAMMATORY RESPONSE LIVE UP | 485 | 293 | All SZGR 2.0 genes in this pathway |
| ZHOU INFLAMMATORY RESPONSE LPS UP | 431 | 237 | All SZGR 2.0 genes in this pathway |
| CHEBOTAEV GR TARGETS UP | 77 | 62 | All SZGR 2.0 genes in this pathway |
| GRAESSMANN APOPTOSIS BY SERUM DEPRIVATION UP | 552 | 347 | All SZGR 2.0 genes in this pathway |
| GRAESSMANN APOPTOSIS BY DOXORUBICIN UP | 1142 | 669 | All SZGR 2.0 genes in this pathway |
| GRAESSMANN RESPONSE TO MC AND DOXORUBICIN UP | 612 | 367 | All SZGR 2.0 genes in this pathway |
| BERNARD PPAPDC1B TARGETS DN | 58 | 39 | All SZGR 2.0 genes in this pathway |
| MEISSNER NPC HCP WITH H3 UNMETHYLATED | 536 | 296 | All SZGR 2.0 genes in this pathway |
| CHYLA CBFA2T3 TARGETS DN | 242 | 146 | All SZGR 2.0 genes in this pathway |
| ZWANG TRANSIENTLY UP BY 2ND EGF PULSE ONLY | 1725 | 838 | All SZGR 2.0 genes in this pathway |