Gene Page: MORN1
Summary ?
| GeneID | 79906 |
| Symbol | MORN1 |
| Synonyms | - |
| Description | MORN repeat containing 1 |
| Reference | HGNC:HGNC:25852|Ensembl:ENSG00000116151|HPRD:07843|Vega:OTTHUMG00000001402 |
| Gene type | protein-coding |
| Map location | 1p36.33-p36.32 |
| Pascal p-value | 0.039 |
| Sherlock p-value | 0.248 |
| Fetal beta | -0.423 |
| DMG | 1 (# studies) |
| eGene | Caudate basal ganglia Cortex Frontal Cortex BA9 Myers' cis & trans Meta |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| DMG:Wockner_2014 | Genome-wide DNA methylation analysis | This dataset includes 4641 differentially methylated probes corresponding to 2929 unique genes between schizophrenia patients (n=24) and controls (n=24). | 3 |
Section I. Genetics and epigenetics annotation
 Differentially methylated gene
Differentially methylated gene
| Probe | Chromosome | Position | Nearest gene | P (dis) | Beta (dis) | FDR (dis) | Study |
|---|---|---|---|---|---|---|---|
| cg27054514 | 1 | 2252995 | MORN1 | 7.72E-5 | 0.359 | 0.025 | DMG:Wockner_2014 |
| cg09438972 | 1 | 2316354 | MORN1 | 4.423E-4 | 0.565 | 0.045 | DMG:Wockner_2014 |
| cg04286030 | 1 | 2313373 | MORN1 | 4.659E-4 | -0.422 | 0.046 | DMG:Wockner_2014 |
 eQTL annotation
eQTL annotation
| SNP ID | Chromosome | Position | eGene | Gene Entrez ID | pvalue | qvalue | TSS distance | eQTL type |
|---|---|---|---|---|---|---|---|---|
| rs17392743 | chr1 | 54764240 | MORN1 | 79906 | 0.13 | trans | ||
| rs17572651 | chr1 | 218943612 | MORN1 | 79906 | 0 | trans | ||
| rs16829545 | chr2 | 151977407 | MORN1 | 79906 | 8.075E-11 | trans | ||
| rs3845734 | chr2 | 171125572 | MORN1 | 79906 | 0.01 | trans | ||
| rs7584986 | chr2 | 184111432 | MORN1 | 79906 | 0 | trans | ||
| rs7635430 | chr3 | 180152035 | MORN1 | 79906 | 0.07 | trans | ||
| rs17762315 | chr5 | 76807576 | MORN1 | 79906 | 0.01 | trans | ||
| rs11185824 | chr10 | 91393711 | MORN1 | 79906 | 0.17 | trans | ||
| rs16955618 | chr15 | 29937543 | MORN1 | 79906 | 7.056E-18 | trans | ||
| rs17711752 | chr16 | 34436184 | MORN1 | 79906 | 0.19 | trans | ||
| rs1041786 | chr21 | 22617710 | MORN1 | 79906 | 0.02 | trans |
Section II. Transcriptome annotation
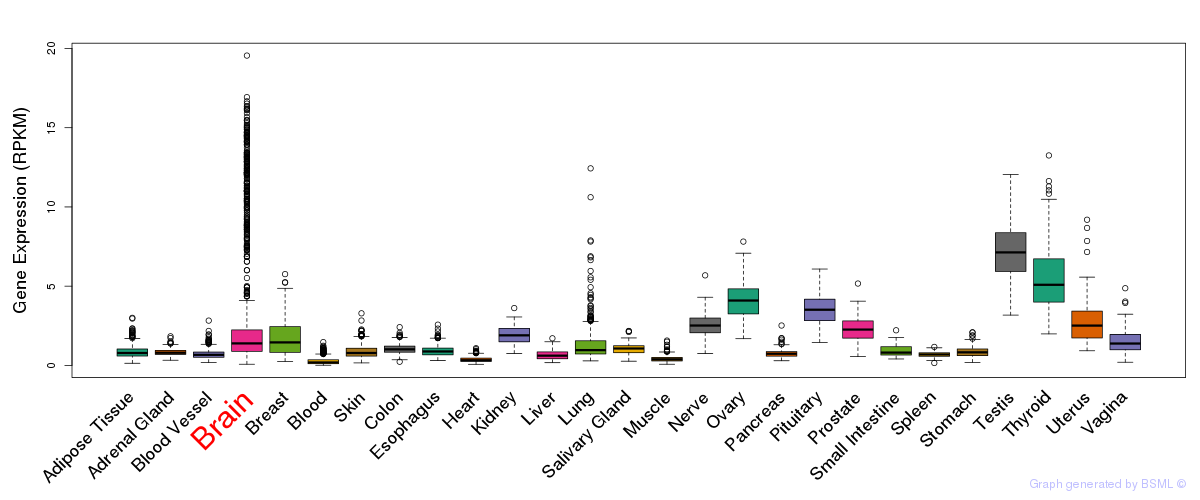
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
No co-expressed genes in brain regions
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| ZHOU INFLAMMATORY RESPONSE FIMA UP | 544 | 308 | All SZGR 2.0 genes in this pathway |
| ZHOU INFLAMMATORY RESPONSE LPS UP | 431 | 237 | All SZGR 2.0 genes in this pathway |
| JAERVINEN AMPLIFIED IN LARYNGEAL CANCER | 40 | 24 | All SZGR 2.0 genes in this pathway |
| LOPEZ MBD TARGETS | 957 | 597 | All SZGR 2.0 genes in this pathway |
| ROPERO HDAC2 TARGETS | 114 | 71 | All SZGR 2.0 genes in this pathway |
| LEE LIVER CANCER SURVIVAL UP | 185 | 112 | All SZGR 2.0 genes in this pathway |
| MARTENS TRETINOIN RESPONSE DN | 841 | 431 | All SZGR 2.0 genes in this pathway |
| BRUINS UVC RESPONSE VIA TP53 GROUP D | 280 | 158 | All SZGR 2.0 genes in this pathway |