Gene Page: STMN4
Summary ?
| GeneID | 81551 |
| Symbol | STMN4 |
| Synonyms | RB3 |
| Description | stathmin 4 |
| Reference | HGNC:HGNC:16078|Ensembl:ENSG00000015592|HPRD:18120|Vega:OTTHUMG00000099461 |
| Gene type | protein-coding |
| Map location | 8p21.2 |
| Pascal p-value | 0.043 |
| Sherlock p-value | 0.622 |
| Fetal beta | 0.592 |
| eGene | Caudate basal ganglia Cortex Hippocampus Nucleus accumbens basal ganglia Putamen basal ganglia Meta |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:GWAScat | Genome-wide Association Studies | This data set includes 560 SNPs associated with schizophrenia. A total of 486 genes were mapped to these SNPs within 50kb. | |
| CV:GWASdb | Genome-wide Association Studies | GWASdb records for schizophrenia | |
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| PMID:cooccur | High-throughput literature-search | Systematic search in PubMed for genes co-occurring with SCZ keywords. A total of 3027 genes were included. | |
| GSMA_I | Genome scan meta-analysis | Psr: 0.031 | |
| GSMA_IIE | Genome scan meta-analysis (European-ancestry samples) | Psr: 0.00057 | |
| GSMA_IIA | Genome scan meta-analysis (All samples) | Psr: 0.03086 | |
| Literature | High-throughput literature-search | Co-occurance with Schizophrenia keywords: schizophrenia,schizophrenias | Click to show details |
Section I. Genetics and epigenetics annotation
 eQTL annotation
eQTL annotation
| SNP ID | Chromosome | Position | eGene | Gene Entrez ID | pvalue | qvalue | TSS distance | eQTL type |
|---|---|---|---|---|---|---|---|---|
| rs1562330 | 8 | 27098436 | STMN4 | ENSG00000015592.12 | 2.57902E-15 | 0 | 17501 | gtex_brain_putamen_basal |
| rs17366947 | 8 | 27103024 | STMN4 | ENSG00000015592.12 | 2.57902E-15 | 0 | 12913 | gtex_brain_putamen_basal |
| rs10481349 | 8 | 27104727 | STMN4 | ENSG00000015592.12 | 2.51219E-15 | 0 | 11210 | gtex_brain_putamen_basal |
| rs28407392 | 8 | 27117727 | STMN4 | ENSG00000015592.12 | 1.25788E-11 | 0 | -1790 | gtex_brain_putamen_basal |
| rs62502378 | 8 | 27119012 | STMN4 | ENSG00000015592.12 | 1.79266E-6 | 0 | -3075 | gtex_brain_putamen_basal |
| rs1345134 | 8 | 27119538 | STMN4 | ENSG00000015592.12 | 2.52833E-6 | 0 | -3601 | gtex_brain_putamen_basal |
Section II. Transcriptome annotation
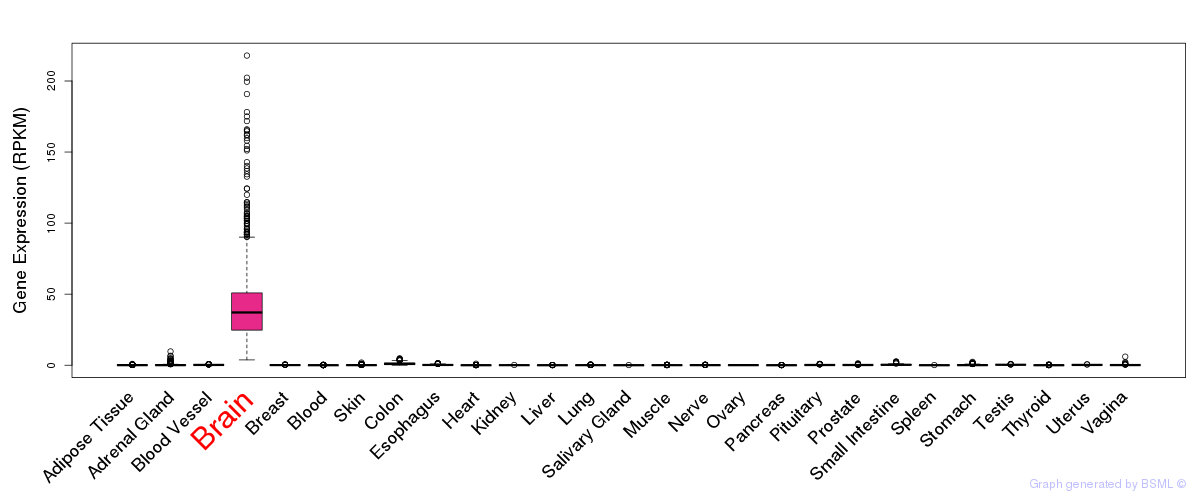
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
No co-expressed genes in brain regions
Section III. Gene Ontology annotation
| Biological process | GO term | Evidence | Neuro keywords | PubMed ID |
|---|---|---|---|---|
| GO:0007242 | intracellular signaling cascade | IEA | - |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| LEE NEURAL CREST STEM CELL UP | 146 | 99 | All SZGR 2.0 genes in this pathway |
| ZHOU INFLAMMATORY RESPONSE FIMA UP | 544 | 308 | All SZGR 2.0 genes in this pathway |
| ZHOU INFLAMMATORY RESPONSE LPS UP | 431 | 237 | All SZGR 2.0 genes in this pathway |
| RIGGI EWING SARCOMA PROGENITOR UP | 430 | 288 | All SZGR 2.0 genes in this pathway |
| MCMURRAY TP53 HRAS COOPERATION RESPONSE DN | 67 | 46 | All SZGR 2.0 genes in this pathway |
| VERHAAK GLIOBLASTOMA PRONEURAL | 177 | 132 | All SZGR 2.0 genes in this pathway |
| PLASARI TGFB1 TARGETS 10HR UP | 199 | 143 | All SZGR 2.0 genes in this pathway |
Section VI. microRNA annotation
| miRNA family | Target position | miRNA ID | miRNA seq | ||
|---|---|---|---|---|---|
| UTR start | UTR end | Match method | |||
| miR-135 | 173 | 179 | m8 | hsa-miR-135a | UAUGGCUUUUUAUUCCUAUGUGA |
| hsa-miR-135b | UAUGGCUUUUCAUUCCUAUGUG | ||||
| miR-150 | 210 | 216 | 1A | hsa-miR-150 | UCUCCCAACCCUUGUACCAGUG |
| miR-433-3p | 522 | 528 | m8 | hsa-miR-433brain | AUCAUGAUGGGCUCCUCGGUGU |
- SZ: miRNAs which differentially expressed in brain cortex of schizophrenia patients comparing with control samples using microarray. Click here to see the list of SZ related miRNAs.
- Brain: miRNAs which are expressed in brain based on miRNA microarray expression studies. Click here to see the list of brain related miRNAs.