Gene Page: SNURF
Summary ?
| GeneID | 8926 |
| Symbol | SNURF |
| Synonyms | - |
| Description | SNRPN upstream reading frame |
| Reference | HGNC:HGNC:11171|HPRD:18083| |
| Gene type | protein-coding |
| Map location | 15q12 |
| Pascal p-value | 0.239 |
| Sherlock p-value | 0.496 |
| Fetal beta | -0.92 |
| eGene | Caudate basal ganglia Myers' cis & trans |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CNV:YES | Copy number variation studies | Manual curation | |
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| PMID:cooccur | High-throughput literature-search | Systematic search in PubMed for genes co-occurring with SCZ keywords. A total of 3027 genes were included. | |
| Literature | High-throughput literature-search | Co-occurance with Schizophrenia keywords: schizophrenia,schizophrenias | Click to show details |
Section I. Genetics and epigenetics annotation
 eQTL annotation
eQTL annotation
| SNP ID | Chromosome | Position | eGene | Gene Entrez ID | pvalue | qvalue | TSS distance | eQTL type |
|---|---|---|---|---|---|---|---|---|
| rs2105242 | chr1 | 163568328 | SNURF | 8926 | 0.08 | trans |
Section II. Transcriptome annotation
General gene expression (GTEx)
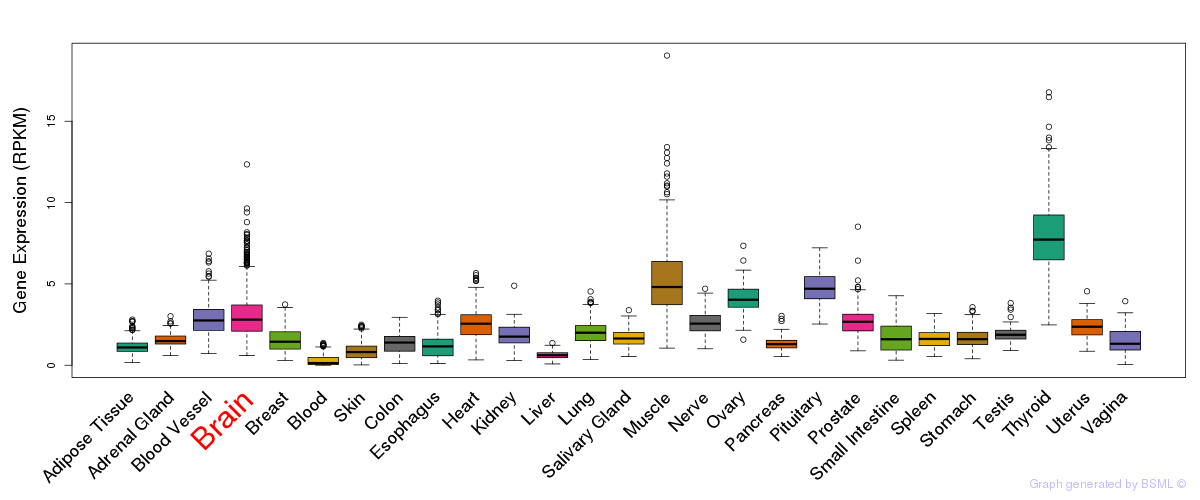
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| NRGN | 0.66 | 0.62 |
| ITPKA | 0.60 | 0.60 |
| CAMK1 | 0.59 | 0.53 |
| NECAB3 | 0.59 | 0.47 |
| CCDC85B | 0.57 | 0.60 |
| OCEL1 | 0.57 | 0.56 |
| KREMEN2 | 0.57 | 0.61 |
| CACNA1F | 0.57 | 0.60 |
| GRASP | 0.56 | 0.62 |
| FAM131A | 0.56 | 0.51 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| ZNF766 | -0.42 | -0.48 |
| ZNF345 | -0.42 | -0.48 |
| TUBB2B | -0.42 | -0.45 |
| ZNF3 | -0.41 | -0.45 |
| UPF3B | -0.41 | -0.54 |
| THOC2 | -0.41 | -0.51 |
| TIGD1 | -0.41 | -0.45 |
| ZNF311 | -0.41 | -0.43 |
| CCDC123 | -0.41 | -0.43 |
| AC004017.1 | -0.41 | -0.43 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| PID AR PATHWAY | 61 | 46 | All SZGR 2.0 genes in this pathway |
| AFFAR YY1 TARGETS UP | 214 | 133 | All SZGR 2.0 genes in this pathway |
| LEIN LOCALIZED TO PROXIMAL DENDRITES | 37 | 26 | All SZGR 2.0 genes in this pathway |
| MUELLER PLURINET | 299 | 189 | All SZGR 2.0 genes in this pathway |
| COLINA TARGETS OF 4EBP1 AND 4EBP2 | 356 | 214 | All SZGR 2.0 genes in this pathway |
| TERAO AOX4 TARGETS SKIN DN | 27 | 15 | All SZGR 2.0 genes in this pathway |
| BRIDEAU IMPRINTED GENES | 63 | 47 | All SZGR 2.0 genes in this pathway |