Gene Page: TIMM50
Summary ?
| GeneID | 92609 |
| Symbol | TIMM50 |
| Synonyms | TIM50|TIM50L |
| Description | translocase of inner mitochondrial membrane 50 |
| Reference | MIM:607381|HGNC:HGNC:23656|Ensembl:ENSG00000105197|HPRD:12118| |
| Gene type | protein-coding |
| Map location | 19q13.2 |
| Pascal p-value | 0.017 |
| Fetal beta | 0.681 |
| DMG | 1 (# studies) |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| DMG:Jaffe_2016 | Genome-wide DNA methylation analysis | This dataset includes 2,104 probes/CpGs associated with SZ patients (n=108) compared to 136 controls at Bonferroni-adjusted P < 0.05. | 2 |
Section I. Genetics and epigenetics annotation
 Differentially methylated gene
Differentially methylated gene
| Probe | Chromosome | Position | Nearest gene | P (dis) | Beta (dis) | FDR (dis) | Study |
|---|---|---|---|---|---|---|---|
| cg11865883 | 19 | 39971137 | TIMM50 | 6.53E-9 | -0.015 | 3.38E-6 | DMG:Jaffe_2016 |
| cg00945719 | 19 | 39971230 | TIMM50 | 5.06E-8 | -0.013 | 1.35E-5 | DMG:Jaffe_2016 |
Section II. Transcriptome annotation
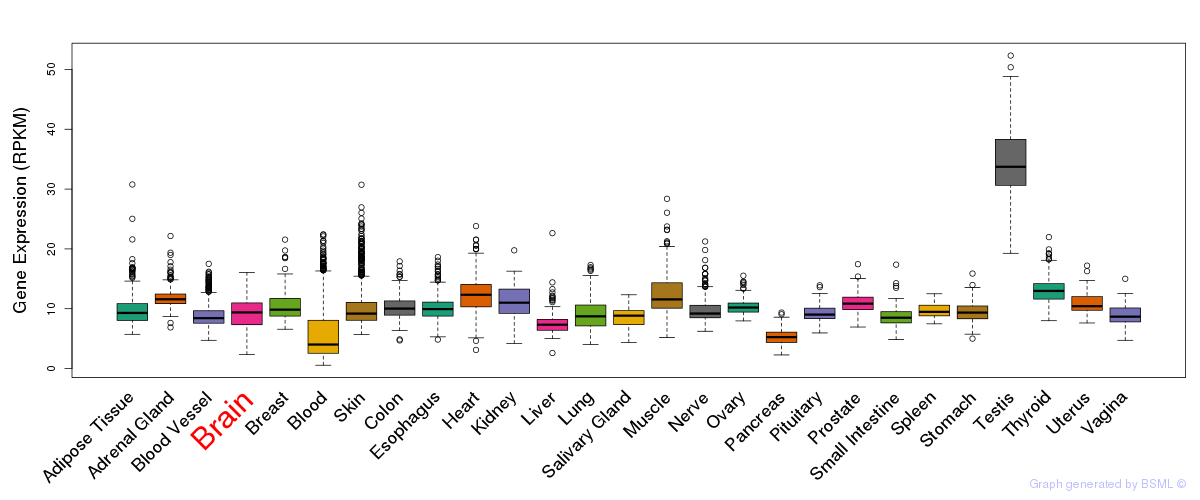
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
No co-expressed genes in brain regions
Section IV. Protein-protein interaction annotation
| Interactors | Aliases B | Official full name B | Experimental | Source | PubMed ID |
|---|---|---|---|---|---|
| AKTIP | FT1 | FTS | AKT interacting protein | Affinity Capture-MS | BioGRID | 17353931 |
| ARAF | A-RAF | ARAF1 | PKS2 | RAFA1 | v-raf murine sarcoma 3611 viral oncogene homolog | A-RAF interacts with TIMM50. | BIND | 12620389 |
| ARAF | A-RAF | ARAF1 | PKS2 | RAFA1 | v-raf murine sarcoma 3611 viral oncogene homolog | ARAF (A-Raf) interacts with TIMM50 (hC22). | BIND | 12620389 |
| BRAF | B-RAF1 | BRAF1 | FLJ95109 | MGC126806 | MGC138284 | RAFB1 | v-raf murine sarcoma viral oncogene homolog B1 | B-RAF interacts with TIMM50. | BIND | 12620389 |
| MAGED1 | DLXIN-1 | NRAGE | melanoma antigen family D, 1 | Affinity Capture-MS | BioGRID | 17353931 |
| MAP3K3 | MAPKKK3 | MEKK3 | mitogen-activated protein kinase kinase kinase 3 | - | HPRD | 14743216 |
| MAP3K7IP2 | FLJ21885 | KIAA0733 | TAB2 | mitogen-activated protein kinase kinase kinase 7 interacting protein 2 | - | HPRD | 14743216 |
| NFKB2 | LYT-10 | LYT10 | nuclear factor of kappa light polypeptide gene enhancer in B-cells 2 (p49/p100) | - | HPRD | 14743216 |
| NFKBIA | IKBA | MAD-3 | NFKBI | nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, alpha | - | HPRD | 14743216 |
| NFKBIB | IKBB | TRIP9 | nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, beta | - | HPRD | 14743216 |
| PAFAH1B2 | - | platelet-activating factor acetylhydrolase, isoform Ib, beta subunit 30kDa | Two-hybrid | BioGRID | 16169070 |
| PELO | CGI-17 | PRO1770 | pelota homolog (Drosophila) | Affinity Capture-MS | BioGRID | 17353931 |
| PHB2 | BAP | BCAP37 | Bap37 | MGC117268 | PNAS-141 | REA | p22 | prohibitin 2 | Affinity Capture-MS | BioGRID | 17353931 |
| RAF1 | CRAF | NS5 | Raf-1 | c-Raf | v-raf-1 murine leukemia viral oncogene homolog 1 | C-RAF interacts with TIMM50. | BIND | 12620389 |
| RAF1 | CRAF | NS5 | Raf-1 | c-Raf | v-raf-1 murine leukemia viral oncogene homolog 1 | RAF1 (C-Raf) interacts with TIMM50 (hC22). | BIND | 12620389 |
| TIMM23 | MGC22767 | PRO1197 | TIM23 | translocase of inner mitochondrial membrane 23 homolog (yeast) | - | HPRD,BioGRID | 12437925 |15044455 |
| TNFRSF10B | CD262 | DR5 | KILLER | KILLER/DR5 | TRAIL-R2 | TRAILR2 | TRICK2 | TRICK2A | TRICK2B | TRICKB | ZTNFR9 | tumor necrosis factor receptor superfamily, member 10b | Two-hybrid | BioGRID | 15044455 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| REACTOME MITOCHONDRIAL PROTEIN IMPORT | 58 | 24 | All SZGR 2.0 genes in this pathway |
| REACTOME METABOLISM OF PROTEINS | 518 | 242 | All SZGR 2.0 genes in this pathway |
| CASORELLI ACUTE PROMYELOCYTIC LEUKEMIA DN | 663 | 425 | All SZGR 2.0 genes in this pathway |
| HORIUCHI WTAP TARGETS DN | 310 | 188 | All SZGR 2.0 genes in this pathway |
| ZHOU INFLAMMATORY RESPONSE FIMA UP | 544 | 308 | All SZGR 2.0 genes in this pathway |
| ZHOU INFLAMMATORY RESPONSE LPS UP | 431 | 237 | All SZGR 2.0 genes in this pathway |
| COLDREN GEFITINIB RESISTANCE UP | 85 | 57 | All SZGR 2.0 genes in this pathway |
| WEI MYCN TARGETS WITH E BOX | 795 | 478 | All SZGR 2.0 genes in this pathway |
| MOREAUX B LYMPHOCYTE MATURATION BY TACI DN | 73 | 45 | All SZGR 2.0 genes in this pathway |
| MOREAUX MULTIPLE MYELOMA BY TACI DN | 172 | 107 | All SZGR 2.0 genes in this pathway |
| KRIGE RESPONSE TO TOSEDOSTAT 6HR UP | 953 | 554 | All SZGR 2.0 genes in this pathway |
| KRIGE RESPONSE TO TOSEDOSTAT 24HR DN | 1011 | 592 | All SZGR 2.0 genes in this pathway |
| MARSON BOUND BY E2F4 UNSTIMULATED | 728 | 415 | All SZGR 2.0 genes in this pathway |
| STEIN ESRRA TARGETS UP | 388 | 234 | All SZGR 2.0 genes in this pathway |
| KUUSELO PANCREATIC CANCER 19Q13 AMPLIFICATION | 35 | 22 | All SZGR 2.0 genes in this pathway |
| STEIN ESRRA TARGETS | 535 | 325 | All SZGR 2.0 genes in this pathway |