Gene Page: SLC22A6
Summary ?
| GeneID | 9356 |
| Symbol | SLC22A6 |
| Synonyms | HOAT1|OAT1|PAHT|ROAT1 |
| Description | solute carrier family 22 member 6 |
| Reference | MIM:607582|HGNC:HGNC:10970|Ensembl:ENSG00000197901|HPRD:12125|Vega:OTTHUMG00000167767 |
| Gene type | protein-coding |
| Map location | 11q12.3 |
| Pascal p-value | 0.241 |
| Fetal beta | -0.324 |
| eGene | Myers' cis & trans Meta |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| PMID:cooccur | High-throughput literature-search | Systematic search in PubMed for genes co-occurring with SCZ keywords. A total of 3027 genes were included. | |
| Literature | High-throughput literature-search | Co-occurance with Schizophrenia keywords: schizophrenia,schizophrenias | Click to show details |
Section I. Genetics and epigenetics annotation
 eQTL annotation
eQTL annotation
| SNP ID | Chromosome | Position | eGene | Gene Entrez ID | pvalue | qvalue | TSS distance | eQTL type |
|---|---|---|---|---|---|---|---|---|
| rs7541471 | chr1 | 39034942 | SLC22A6 | 9356 | 8.918E-4 | trans | ||
| rs16825217 | chr1 | 39039168 | SLC22A6 | 9356 | 0.02 | trans | ||
| rs10493495 | chr1 | 72722583 | SLC22A6 | 9356 | 0.1 | trans | ||
| rs7541200 | chr1 | 181327476 | SLC22A6 | 9356 | 0.02 | trans | ||
| rs9309267 | chr2 | 55850332 | SLC22A6 | 9356 | 0.16 | trans | ||
| rs6739788 | chr2 | 55862354 | SLC22A6 | 9356 | 0.16 | trans | ||
| rs17015803 | chr2 | 120191430 | SLC22A6 | 9356 | 0 | trans | ||
| rs17022377 | chr4 | 95783024 | SLC22A6 | 9356 | 0.12 | trans | ||
| rs313946 | chr4 | 113975904 | SLC22A6 | 9356 | 0.01 | trans | ||
| rs16880508 | chr5 | 8396460 | SLC22A6 | 9356 | 0.18 | trans | ||
| rs7734944 | chr5 | 39646529 | SLC22A6 | 9356 | 0.11 | trans | ||
| rs295936 | chr5 | 58865856 | SLC22A6 | 9356 | 0.07 | trans | ||
| rs11250964 | chr10 | 2005552 | SLC22A6 | 9356 | 0.13 | trans | ||
| rs3750769 | chr10 | 71859865 | SLC22A6 | 9356 | 0.09 | trans | ||
| rs11016704 | chr10 | 128989145 | SLC22A6 | 9356 | 0.04 | trans | ||
| rs481843 | chr11 | 116525866 | SLC22A6 | 9356 | 0.2 | trans | ||
| rs17023015 | chr12 | 95053554 | SLC22A6 | 9356 | 0.16 | trans | ||
| rs9529974 | chr13 | 72821935 | SLC22A6 | 9356 | 0.08 | trans | ||
| rs7403047 | chr15 | 38822791 | SLC22A6 | 9356 | 0.14 | trans | ||
| rs4787772 | chr16 | 25855167 | SLC22A6 | 9356 | 0.01 | trans | ||
| rs1824248 | chr18 | 31073821 | SLC22A6 | 9356 | 0.08 | trans | ||
| rs12455504 | chr18 | 42122043 | SLC22A6 | 9356 | 0.05 | trans | ||
| snp_a-1913345 | 0 | SLC22A6 | 9356 | 0.04 | trans | |||
| rs16979011 | chr21 | 28154453 | SLC22A6 | 9356 | 0.07 | trans | ||
| rs2830502 | chr21 | 28157753 | SLC22A6 | 9356 | 0.07 | trans | ||
| rs17255719 | chrX | 12144766 | SLC22A6 | 9356 | 0.01 | trans | ||
| rs4893621 | chrX | 51993653 | SLC22A6 | 9356 | 0.15 | trans | ||
| rs12395234 | chrX | 52617968 | SLC22A6 | 9356 | 0.11 | trans | ||
| rs6568279 | 0 | SLC22A6 | 9356 | 0.13 | trans | |||
| rs12556892 | chrX | 93417735 | SLC22A6 | 9356 | 0.07 | trans | ||
| rs808754 | chrX | 93438033 | SLC22A6 | 9356 | 0.07 | trans | ||
| rs6529399 | chrX | 128763582 | SLC22A6 | 9356 | 0.01 | trans | ||
| rs3761581 | chrX | 128789720 | SLC22A6 | 9356 | 0.06 | trans | ||
| rs242163 | chrX | 132311662 | SLC22A6 | 9356 | 0.04 | trans | ||
| rs17252704 | chrX | 148612450 | SLC22A6 | 9356 | 0.05 | trans |
Section II. Transcriptome annotation
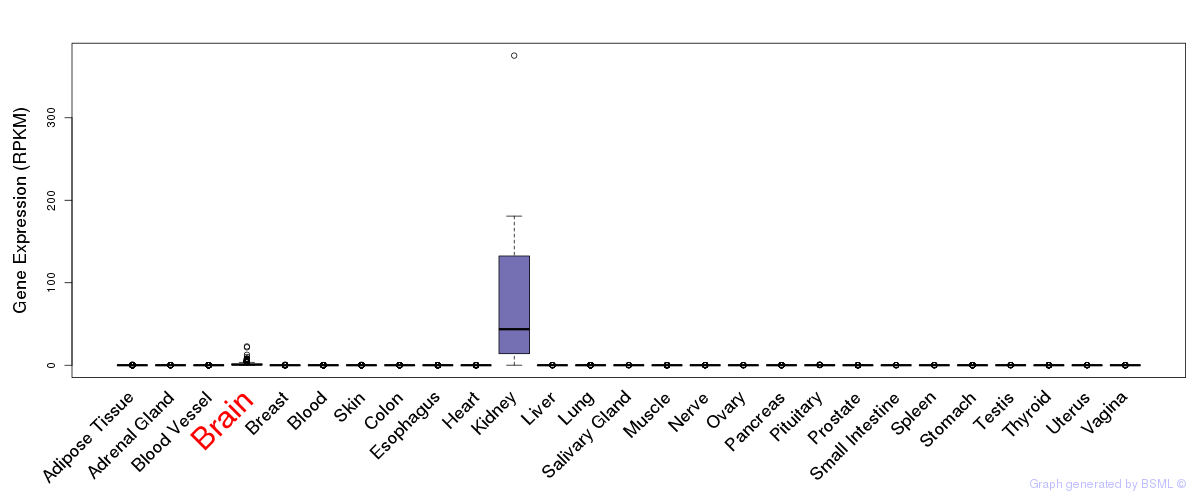
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| CHD9 | 0.91 | 0.89 |
| AP001011.3 | 0.91 | 0.87 |
| ZNF292 | 0.91 | 0.89 |
| ZNF518B | 0.91 | 0.89 |
| UBR5 | 0.90 | 0.86 |
| BTBD7 | 0.90 | 0.85 |
| MGA | 0.90 | 0.88 |
| C20orf74 | 0.90 | 0.84 |
| ARID2 | 0.90 | 0.87 |
| XRN1 | 0.90 | 0.86 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| AIFM3 | -0.68 | -0.77 |
| HLA-F | -0.68 | -0.77 |
| AF347015.31 | -0.67 | -0.83 |
| LHPP | -0.67 | -0.69 |
| TSC22D4 | -0.67 | -0.78 |
| ALDOC | -0.66 | -0.74 |
| HEPN1 | -0.65 | -0.75 |
| TNFSF12 | -0.65 | -0.72 |
| PTH1R | -0.65 | -0.74 |
| MT-CO2 | -0.65 | -0.83 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| REACTOME TRANSMEMBRANE TRANSPORT OF SMALL MOLECULES | 413 | 270 | All SZGR 2.0 genes in this pathway |
| REACTOME SLC MEDIATED TRANSMEMBRANE TRANSPORT | 241 | 157 | All SZGR 2.0 genes in this pathway |
| REACTOME TRANSPORT OF GLUCOSE AND OTHER SUGARS BILE SALTS AND ORGANIC ACIDS METAL IONS AND AMINE COMPOUNDS | 89 | 54 | All SZGR 2.0 genes in this pathway |
| REACTOME ORGANIC CATION ANION ZWITTERION TRANSPORT | 13 | 9 | All SZGR 2.0 genes in this pathway |
| JAZAG TGFB1 SIGNALING VIA SMAD4 DN | 66 | 38 | All SZGR 2.0 genes in this pathway |
| ROPERO HDAC2 TARGETS | 114 | 71 | All SZGR 2.0 genes in this pathway |
| FLECHNER BIOPSY KIDNEY TRANSPLANT REJECTED VS OK DN | 546 | 351 | All SZGR 2.0 genes in this pathway |
| BAELDE DIABETIC NEPHROPATHY UP | 86 | 48 | All SZGR 2.0 genes in this pathway |
| BILANGES SERUM SENSITIVE VIA TSC1 | 23 | 12 | All SZGR 2.0 genes in this pathway |