Gene Page: PLAA
Summary ?
| GeneID | 9373 |
| Symbol | PLAA |
| Synonyms | DOA1|PLA2P|PLAP |
| Description | phospholipase A2 activating protein |
| Reference | MIM:603873|HGNC:HGNC:9043|Ensembl:ENSG00000137055|HPRD:04850|Vega:OTTHUMG00000019708 |
| Gene type | protein-coding |
| Map location | 9p21 |
| Pascal p-value | 0.002 |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:GWAScat | Genome-wide Association Studies | This data set includes 560 SNPs associated with schizophrenia. A total of 486 genes were mapped to these SNPs within 50kb. | |
| CV:GWASdb | Genome-wide Association Studies | GWASdb records for schizophrenia | |
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| PMID:cooccur | High-throughput literature-search | Systematic search in PubMed for genes co-occurring with SCZ keywords. A total of 3027 genes were included. | |
| Literature | High-throughput literature-search | Co-occurance with Schizophrenia keywords: schizophrenia,schizophrenias | Click to show details |
Section I. Genetics and epigenetics annotation
Section II. Transcriptome annotation
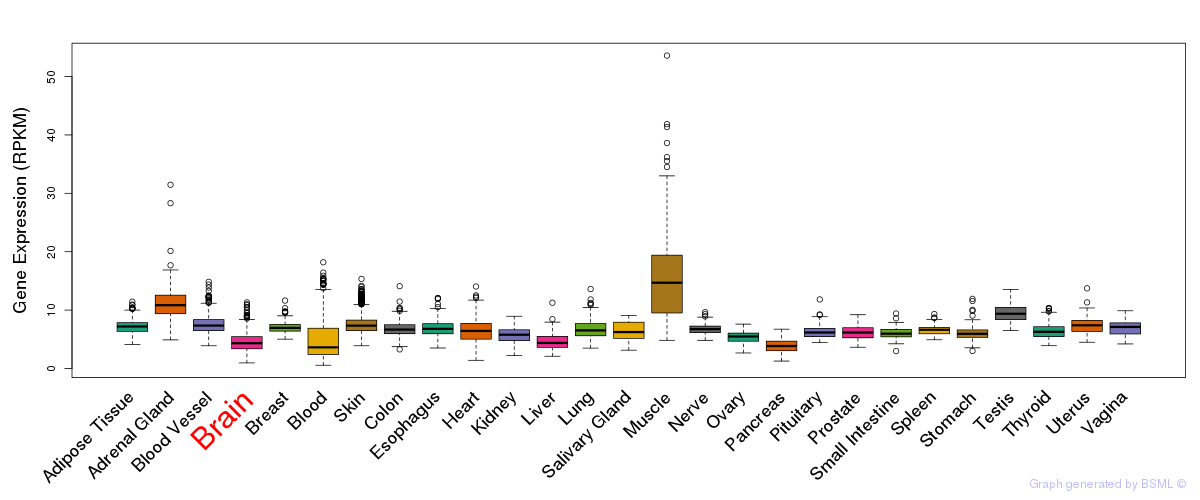
General gene expression (GTEx)
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| TMC6 | 0.78 | 0.84 |
| CHADL | 0.77 | 0.84 |
| PPAP2C | 0.74 | 0.78 |
| RTKN | 0.73 | 0.81 |
| TFEB | 0.73 | 0.80 |
| GSN | 0.73 | 0.78 |
| SIRT2 | 0.73 | 0.82 |
| MAG | 0.73 | 0.78 |
| RHOG | 0.73 | 0.77 |
| VWA1 | 0.72 | 0.80 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| SLC24A5 | -0.51 | -0.61 |
| SNRPB2 | -0.51 | -0.63 |
| SNORA32 | -0.50 | -0.64 |
| ZBED5 | -0.50 | -0.61 |
| ZNF300 | -0.50 | -0.53 |
| CCNG2 | -0.50 | -0.57 |
| DSCC1 | -0.49 | -0.57 |
| SNX7 | -0.49 | -0.59 |
| ZNF682 | -0.49 | -0.54 |
| AIM2 | -0.49 | -0.62 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| GARY CD5 TARGETS DN | 431 | 263 | All SZGR 2.0 genes in this pathway |
| DIAZ CHRONIC MEYLOGENOUS LEUKEMIA UP | 1382 | 904 | All SZGR 2.0 genes in this pathway |
| GAL LEUKEMIC STEM CELL DN | 244 | 153 | All SZGR 2.0 genes in this pathway |
| KINSEY TARGETS OF EWSR1 FLII FUSION UP | 1278 | 748 | All SZGR 2.0 genes in this pathway |
| BENPORATH OCT4 TARGETS | 290 | 172 | All SZGR 2.0 genes in this pathway |
| KENNY CTNNB1 TARGETS DN | 52 | 34 | All SZGR 2.0 genes in this pathway |
| SHEPARD BMYB MORPHOLINO UP | 205 | 126 | All SZGR 2.0 genes in this pathway |
| IVANOVA HEMATOPOIESIS INTERMEDIATE PROGENITOR | 149 | 84 | All SZGR 2.0 genes in this pathway |
| MARSON BOUND BY E2F4 UNSTIMULATED | 728 | 415 | All SZGR 2.0 genes in this pathway |
| WANG TUMOR INVASIVENESS DN | 210 | 128 | All SZGR 2.0 genes in this pathway |
| YOSHIMURA MAPK8 TARGETS DN | 366 | 257 | All SZGR 2.0 genes in this pathway |
| LEE BMP2 TARGETS DN | 882 | 538 | All SZGR 2.0 genes in this pathway |