Gene Page: GPR180
Summary ?
| GeneID | 160897 |
| Symbol | GPR180 |
| Synonyms | ITR |
| Description | G protein-coupled receptor 180 |
| Reference | MIM:607787|HGNC:HGNC:28899|Ensembl:ENSG00000152749|HPRD:16258|Vega:OTTHUMG00000017207 |
| Gene type | protein-coding |
| Map location | 13q32.1 |
| Pascal p-value | 0.602 |
| Fetal beta | -0.276 |
| DMG | 1 (# studies) |
| eGene | Hippocampus Putamen basal ganglia Meta |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| DMG:Jaffe_2016 | Genome-wide DNA methylation analysis | This dataset includes 2,104 probes/CpGs associated with SZ patients (n=108) compared to 136 controls at Bonferroni-adjusted P < 0.05. | 1 |
Section I. Genetics and epigenetics annotation
 Differentially methylated gene
Differentially methylated gene
| Probe | Chromosome | Position | Nearest gene | P (dis) | Beta (dis) | FDR (dis) | Study |
|---|---|---|---|---|---|---|---|
| cg25740925 | 13 | 95254690 | GPR180 | 2.62E-8 | -0.015 | 8.36E-6 | DMG:Jaffe_2016 |
 eQTL annotation
eQTL annotation
| SNP ID | Chromosome | Position | eGene | Gene Entrez ID | pvalue | qvalue | TSS distance | eQTL type |
|---|---|---|---|---|---|---|---|---|
| rs7322704 | 13 | 95217146 | GPR180 | ENSG00000152749.7 | 1.227E-6 | 0 | -37011 | gtex_brain_putamen_basal |
| rs1408999 | 13 | 95220009 | GPR180 | ENSG00000152749.7 | 8.702E-9 | 0 | -34148 | gtex_brain_putamen_basal |
| rs7331814 | 13 | 95222544 | GPR180 | ENSG00000152749.7 | 3.27E-9 | 0 | -31613 | gtex_brain_putamen_basal |
| rs7339106 | 13 | 95223903 | GPR180 | ENSG00000152749.7 | 4.643E-9 | 0 | -30254 | gtex_brain_putamen_basal |
| rs3742107 | 13 | 95227340 | GPR180 | ENSG00000152749.7 | 4.643E-9 | 0 | -26817 | gtex_brain_putamen_basal |
| rs7321239 | 13 | 95229288 | GPR180 | ENSG00000152749.7 | 4.414E-9 | 0 | -24869 | gtex_brain_putamen_basal |
| rs2275648 | 13 | 95231981 | GPR180 | ENSG00000152749.7 | 4.643E-9 | 0 | -22176 | gtex_brain_putamen_basal |
| rs9556402 | 13 | 95233607 | GPR180 | ENSG00000152749.7 | 5.111E-7 | 0 | -20550 | gtex_brain_putamen_basal |
| rs9590037 | 13 | 95234223 | GPR180 | ENSG00000152749.7 | 5.111E-7 | 0 | -19934 | gtex_brain_putamen_basal |
| rs4773801 | 13 | 95235175 | GPR180 | ENSG00000152749.7 | 4.643E-9 | 0 | -18982 | gtex_brain_putamen_basal |
| rs7330873 | 13 | 95235890 | GPR180 | ENSG00000152749.7 | 4.643E-9 | 0 | -18267 | gtex_brain_putamen_basal |
| rs9524544 | 13 | 95236963 | GPR180 | ENSG00000152749.7 | 4.643E-9 | 0 | -17194 | gtex_brain_putamen_basal |
| rs6492716 | 13 | 95237460 | GPR180 | ENSG00000152749.7 | 5.108E-7 | 0 | -16697 | gtex_brain_putamen_basal |
| rs6492717 | 13 | 95237526 | GPR180 | ENSG00000152749.7 | 1.654E-7 | 0 | -16631 | gtex_brain_putamen_basal |
| rs6492718 | 13 | 95237793 | GPR180 | ENSG00000152749.7 | 4.643E-9 | 0 | -16364 | gtex_brain_putamen_basal |
| rs200958171 | 13 | 95238464 | GPR180 | ENSG00000152749.7 | 4.706E-8 | 0 | -15693 | gtex_brain_putamen_basal |
| rs34350725 | 13 | 95238470 | GPR180 | ENSG00000152749.7 | 4.643E-9 | 0 | -15687 | gtex_brain_putamen_basal |
| rs9524545 | 13 | 95238884 | GPR180 | ENSG00000152749.7 | 4.273E-8 | 0 | -15273 | gtex_brain_putamen_basal |
| rs9524547 | 13 | 95239068 | GPR180 | ENSG00000152749.7 | 8.93E-9 | 0 | -15089 | gtex_brain_putamen_basal |
| rs397723832 | 13 | 95239427 | GPR180 | ENSG00000152749.7 | 1.355E-8 | 0 | -14730 | gtex_brain_putamen_basal |
| rs9516434 | 13 | 95240054 | GPR180 | ENSG00000152749.7 | 2.268E-7 | 0 | -14103 | gtex_brain_putamen_basal |
| rs7998025 | 13 | 95240291 | GPR180 | ENSG00000152749.7 | 4.273E-8 | 0 | -13866 | gtex_brain_putamen_basal |
| rs7997333 | 13 | 95240561 | GPR180 | ENSG00000152749.7 | 1.038E-7 | 0 | -13596 | gtex_brain_putamen_basal |
| rs34405130 | 13 | 95241307 | GPR180 | ENSG00000152749.7 | 4.273E-8 | 0 | -12850 | gtex_brain_putamen_basal |
| rs4473052 | 13 | 95241568 | GPR180 | ENSG00000152749.7 | 1E-7 | 0 | -12589 | gtex_brain_putamen_basal |
| rs4608209 | 13 | 95241634 | GPR180 | ENSG00000152749.7 | 2.801E-8 | 0 | -12523 | gtex_brain_putamen_basal |
| rs66560756 | 13 | 95242282 | GPR180 | ENSG00000152749.7 | 4.775E-9 | 0 | -11875 | gtex_brain_putamen_basal |
| rs9524548 | 13 | 95242352 | GPR180 | ENSG00000152749.7 | 4.273E-8 | 0 | -11805 | gtex_brain_putamen_basal |
| rs9524549 | 13 | 95242386 | GPR180 | ENSG00000152749.7 | 4.273E-8 | 0 | -11771 | gtex_brain_putamen_basal |
| rs9301979 | 13 | 95243270 | GPR180 | ENSG00000152749.7 | 4.715E-9 | 0 | -10887 | gtex_brain_putamen_basal |
| rs9524550 | 13 | 95244087 | GPR180 | ENSG00000152749.7 | 4.643E-9 | 0 | -10070 | gtex_brain_putamen_basal |
| rs1112370 | 13 | 95246440 | GPR180 | ENSG00000152749.7 | 4.643E-9 | 0 | -7717 | gtex_brain_putamen_basal |
| rs9524551 | 13 | 95246883 | GPR180 | ENSG00000152749.7 | 4.273E-8 | 0 | -7274 | gtex_brain_putamen_basal |
| rs7997035 | 13 | 95247626 | GPR180 | ENSG00000152749.7 | 4.273E-8 | 0 | -6531 | gtex_brain_putamen_basal |
| rs9516437 | 13 | 95248905 | GPR180 | ENSG00000152749.7 | 1.913E-8 | 0 | -5252 | gtex_brain_putamen_basal |
| rs9516438 | 13 | 95249017 | GPR180 | ENSG00000152749.7 | 5.957E-7 | 0 | -5140 | gtex_brain_putamen_basal |
| rs9301981 | 13 | 95250102 | GPR180 | ENSG00000152749.7 | 2.26E-8 | 0 | -4055 | gtex_brain_putamen_basal |
| rs9561641 | 13 | 95250528 | GPR180 | ENSG00000152749.7 | 1.018E-6 | 0 | -3629 | gtex_brain_putamen_basal |
| rs57292562 | 13 | 95250609 | GPR180 | ENSG00000152749.7 | 3.852E-7 | 0 | -3548 | gtex_brain_putamen_basal |
| rs9516440 | 13 | 95250671 | GPR180 | ENSG00000152749.7 | 2.322E-8 | 0 | -3486 | gtex_brain_putamen_basal |
| rs71207556 | 13 | 95250777 | GPR180 | ENSG00000152749.7 | 2.832E-8 | 0 | -3380 | gtex_brain_putamen_basal |
| rs9516441 | 13 | 95250841 | GPR180 | ENSG00000152749.7 | 2.4E-8 | 0 | -3316 | gtex_brain_putamen_basal |
| rs9516443 | 13 | 95251103 | GPR180 | ENSG00000152749.7 | 2.195E-8 | 0 | -3054 | gtex_brain_putamen_basal |
| rs9524553 | 13 | 95251508 | GPR180 | ENSG00000152749.7 | 2.364E-8 | 0 | -2649 | gtex_brain_putamen_basal |
| rs9524554 | 13 | 95251540 | GPR180 | ENSG00000152749.7 | 2.364E-8 | 0 | -2617 | gtex_brain_putamen_basal |
| rs9561642 | 13 | 95251796 | GPR180 | ENSG00000152749.7 | 2.38E-8 | 0 | -2361 | gtex_brain_putamen_basal |
| rs9301983 | 13 | 95252147 | GPR180 | ENSG00000152749.7 | 2.396E-8 | 0 | -2010 | gtex_brain_putamen_basal |
| rs3897473 | 13 | 95253920 | GPR180 | ENSG00000152749.7 | 2.519E-8 | 0 | -237 | gtex_brain_putamen_basal |
| rs3897474 | 13 | 95254031 | GPR180 | ENSG00000152749.7 | 2.491E-8 | 0 | -126 | gtex_brain_putamen_basal |
| rs9516445 | 13 | 95256891 | GPR180 | ENSG00000152749.7 | 2.721E-8 | 0 | 2734 | gtex_brain_putamen_basal |
| rs35253817 | 13 | 95258242 | GPR180 | ENSG00000152749.7 | 2.824E-8 | 0 | 4085 | gtex_brain_putamen_basal |
| rs9524555 | 13 | 95259536 | GPR180 | ENSG00000152749.7 | 2.899E-8 | 0 | 5379 | gtex_brain_putamen_basal |
| rs9524556 | 13 | 95260421 | GPR180 | ENSG00000152749.7 | 2.724E-8 | 0 | 6264 | gtex_brain_putamen_basal |
| rs9524557 | 13 | 95261775 | GPR180 | ENSG00000152749.7 | 2.822E-8 | 0 | 7618 | gtex_brain_putamen_basal |
| rs7981788 | 13 | 95263924 | GPR180 | ENSG00000152749.7 | 2.898E-8 | 0 | 9767 | gtex_brain_putamen_basal |
| rs9524559 | 13 | 95264604 | GPR180 | ENSG00000152749.7 | 2.906E-8 | 0 | 10447 | gtex_brain_putamen_basal |
| rs9556405 | 13 | 95266244 | GPR180 | ENSG00000152749.7 | 1.671E-8 | 0 | 12087 | gtex_brain_putamen_basal |
| rs9524560 | 13 | 95268942 | GPR180 | ENSG00000152749.7 | 2.72E-7 | 0 | 14785 | gtex_brain_putamen_basal |
| rs9524561 | 13 | 95269009 | GPR180 | ENSG00000152749.7 | 2.713E-7 | 0 | 14852 | gtex_brain_putamen_basal |
| rs9524562 | 13 | 95269233 | GPR180 | ENSG00000152749.7 | 2.721E-7 | 0 | 15076 | gtex_brain_putamen_basal |
| rs9524563 | 13 | 95269541 | GPR180 | ENSG00000152749.7 | 2.722E-7 | 0 | 15384 | gtex_brain_putamen_basal |
| rs9516448 | 13 | 95270392 | GPR180 | ENSG00000152749.7 | 4.793E-7 | 0 | 16235 | gtex_brain_putamen_basal |
| rs9561645 | 13 | 95272210 | GPR180 | ENSG00000152749.7 | 4.829E-7 | 0 | 18053 | gtex_brain_putamen_basal |
| rs1572935 | 13 | 95272552 | GPR180 | ENSG00000152749.7 | 4.843E-7 | 0 | 18395 | gtex_brain_putamen_basal |
| rs9524566 | 13 | 95276735 | GPR180 | ENSG00000152749.7 | 4.958E-7 | 0 | 22578 | gtex_brain_putamen_basal |
| rs9524568 | 13 | 95279505 | GPR180 | ENSG00000152749.7 | 5.285E-7 | 0 | 25348 | gtex_brain_putamen_basal |
| rs2298057 | 13 | 95280923 | GPR180 | ENSG00000152749.7 | 5.236E-7 | 0 | 26766 | gtex_brain_putamen_basal |
| rs2298055 | 13 | 95281296 | GPR180 | ENSG00000152749.7 | 5.267E-7 | 0 | 27139 | gtex_brain_putamen_basal |
| rs4771890 | 13 | 95283295 | GPR180 | ENSG00000152749.7 | 4.194E-7 | 0 | 29138 | gtex_brain_putamen_basal |
Section II. Transcriptome annotation
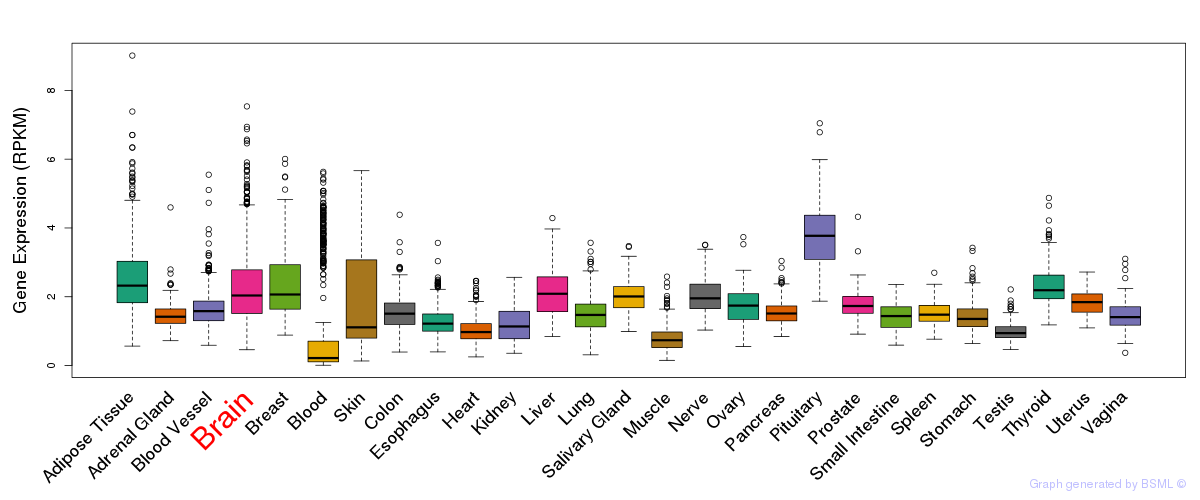
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
No co-expressed genes in brain regions
Section III. Gene Ontology annotation
| Cellular component | GO term | Evidence | Neuro keywords | PubMed ID |
|---|---|---|---|---|
| GO:0016020 | membrane | IEA | - | |
| GO:0016021 | integral to membrane | IEA | - |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| ZHOU INFLAMMATORY RESPONSE LIVE UP | 485 | 293 | All SZGR 2.0 genes in this pathway |
| ZHOU INFLAMMATORY RESPONSE FIMA DN | 284 | 156 | All SZGR 2.0 genes in this pathway |
| RODRIGUES THYROID CARCINOMA POORLY DIFFERENTIATED UP | 633 | 376 | All SZGR 2.0 genes in this pathway |
| NUYTTEN EZH2 TARGETS UP | 1037 | 673 | All SZGR 2.0 genes in this pathway |
| GEORGES TARGETS OF MIR192 AND MIR215 | 893 | 528 | All SZGR 2.0 genes in this pathway |
| VANTVEER BREAST CANCER METASTASIS DN | 121 | 65 | All SZGR 2.0 genes in this pathway |
| IVANOVA HEMATOPOIESIS INTERMEDIATE PROGENITOR | 149 | 84 | All SZGR 2.0 genes in this pathway |
| XU GH1 EXOGENOUS TARGETS DN | 120 | 69 | All SZGR 2.0 genes in this pathway |
| XU GH1 AUTOCRINE TARGETS DN | 142 | 94 | All SZGR 2.0 genes in this pathway |
| VANTVEER BREAST CANCER ESR1 DN | 240 | 153 | All SZGR 2.0 genes in this pathway |
| VANTVEER BREAST CANCER POOR PROGNOSIS | 55 | 30 | All SZGR 2.0 genes in this pathway |
| CHANDRAN METASTASIS UP | 221 | 135 | All SZGR 2.0 genes in this pathway |
| MIYAGAWA TARGETS OF EWSR1 ETS FUSIONS DN | 229 | 135 | All SZGR 2.0 genes in this pathway |
| GOBERT OLIGODENDROCYTE DIFFERENTIATION DN | 1080 | 713 | All SZGR 2.0 genes in this pathway |
| FORTSCHEGGER PHF8 TARGETS UP | 279 | 155 | All SZGR 2.0 genes in this pathway |
Section VI. microRNA annotation
| miRNA family | Target position | miRNA ID | miRNA seq | ||
|---|---|---|---|---|---|
| UTR start | UTR end | Match method | |||
| miR-199 | 978 | 984 | 1A | hsa-miR-199a | CCCAGUGUUCAGACUACCUGUUC |
| hsa-miR-199b | CCCAGUGUUUAGACUAUCUGUUC | ||||
| hsa-miR-199a | CCCAGUGUUCAGACUACCUGUUC | ||||
| hsa-miR-199b | CCCAGUGUUUAGACUAUCUGUUC | ||||
| miR-203.1 | 872 | 878 | 1A | hsa-miR-203 | UGAAAUGUUUAGGACCACUAG |
| miR-25/32/92/363/367 | 43 | 50 | 1A,m8 | hsa-miR-25brain | CAUUGCACUUGUCUCGGUCUGA |
| hsa-miR-32 | UAUUGCACAUUACUAAGUUGC | ||||
| hsa-miR-92 | UAUUGCACUUGUCCCGGCCUG | ||||
| hsa-miR-367 | AAUUGCACUUUAGCAAUGGUGA | ||||
| hsa-miR-92bSZ | UAUUGCACUCGUCCCGGCCUC | ||||
| miR-362 | 923 | 930 | 1A,m8 | hsa-miR-362 | AAUCCUUGGAACCUAGGUGUGAGU |
| miR-376 | 950 | 957 | 1A,m8 | hsa-miR-376a | AUCAUAGAGGAAAAUCCACGU |
| hsa-miR-376b | AUCAUAGAGGAAAAUCCAUGUU | ||||
- SZ: miRNAs which differentially expressed in brain cortex of schizophrenia patients comparing with control samples using microarray. Click here to see the list of SZ related miRNAs.
- Brain: miRNAs which are expressed in brain based on miRNA microarray expression studies. Click here to see the list of brain related miRNAs.