Gene Page: AP4E1
Summary ?
| GeneID | 23431 |
| Symbol | AP4E1 |
| Synonyms | CPSQ4|SPG51 |
| Description | adaptor related protein complex 4 epsilon 1 subunit |
| Reference | MIM:607244|HGNC:HGNC:573|Ensembl:ENSG00000081014|HPRD:06258|Vega:OTTHUMG00000172458 |
| Gene type | protein-coding |
| Map location | 15q21.2 |
| Pascal p-value | 0.014 |
| DMG | 1 (# studies) |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| DMG:Jaffe_2016 | Genome-wide DNA methylation analysis | This dataset includes 2,104 probes/CpGs associated with SZ patients (n=108) compared to 136 controls at Bonferroni-adjusted P < 0.05. | 1 |
Section I. Genetics and epigenetics annotation
 Differentially methylated gene
Differentially methylated gene
| Probe | Chromosome | Position | Nearest gene | P (dis) | Beta (dis) | FDR (dis) | Study |
|---|---|---|---|---|---|---|---|
| cg27183811 | 15 | 51200847 | AP4E1 | 5.89E-10 | -0.009 | 9.07E-7 | DMG:Jaffe_2016 |
Section II. Transcriptome annotation
General gene expression (GTEx)
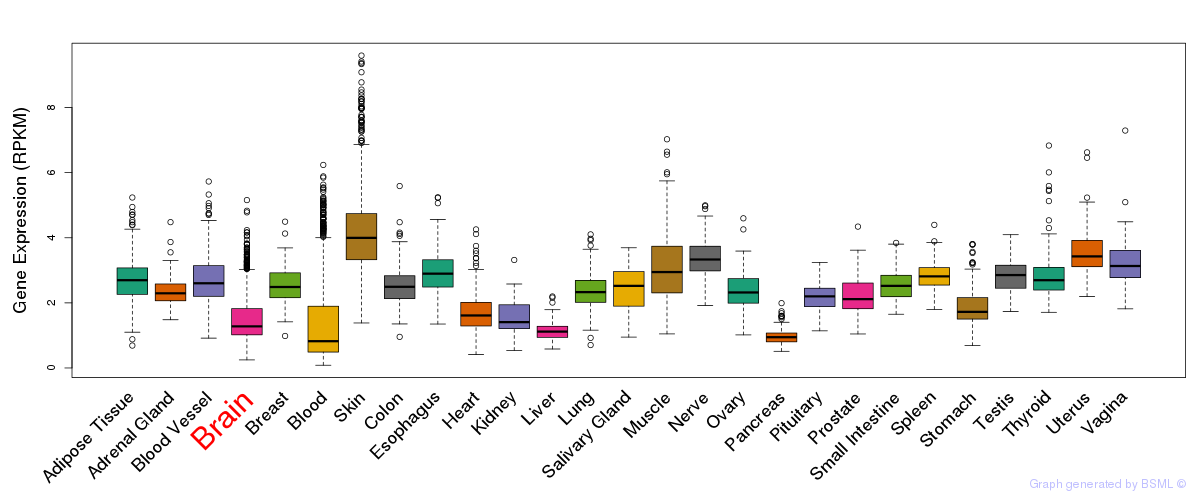
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| CRYBG3 | 0.87 | 0.85 |
| KIAA0494 | 0.87 | 0.87 |
| PDE8A | 0.86 | 0.82 |
| SPG20 | 0.85 | 0.84 |
| DOCK5 | 0.85 | 0.78 |
| ATP10B | 0.85 | 0.85 |
| MTMR10 | 0.84 | 0.80 |
| WIPF1 | 0.84 | 0.83 |
| SLC12A2 | 0.84 | 0.78 |
| ABCA8 | 0.84 | 0.78 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| C1orf86 | -0.47 | -0.54 |
| NR2C2AP | -0.47 | -0.50 |
| CCDC107 | -0.46 | -0.51 |
| AC011491.1 | -0.46 | -0.51 |
| AC174470.1 | -0.46 | -0.53 |
| RPL36 | -0.45 | -0.63 |
| CCDC28B | -0.45 | -0.58 |
| SH2B2 | -0.45 | -0.52 |
| MED19 | -0.45 | -0.50 |
| NOL12 | -0.44 | -0.47 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| KEGG LYSOSOME | 121 | 83 | All SZGR 2.0 genes in this pathway |
| REACTOME MEMBRANE TRAFFICKING | 129 | 74 | All SZGR 2.0 genes in this pathway |
| REACTOME TRANS GOLGI NETWORK VESICLE BUDDING | 60 | 31 | All SZGR 2.0 genes in this pathway |
| REACTOME GOLGI ASSOCIATED VESICLE BIOGENESIS | 53 | 27 | All SZGR 2.0 genes in this pathway |
| WANG LMO4 TARGETS DN | 352 | 225 | All SZGR 2.0 genes in this pathway |
| BENPORATH MYC MAX TARGETS | 775 | 494 | All SZGR 2.0 genes in this pathway |
| KANG IMMORTALIZED BY TERT UP | 89 | 61 | All SZGR 2.0 genes in this pathway |
| CUI TCF21 TARGETS 2 DN | 830 | 547 | All SZGR 2.0 genes in this pathway |
| LIN NPAS4 TARGETS UP | 163 | 100 | All SZGR 2.0 genes in this pathway |
| GYORFFY DOXORUBICIN RESISTANCE | 56 | 34 | All SZGR 2.0 genes in this pathway |
| DANG BOUND BY MYC | 1103 | 714 | All SZGR 2.0 genes in this pathway |
| FORTSCHEGGER PHF8 TARGETS DN | 784 | 464 | All SZGR 2.0 genes in this pathway |