Gene Page: FBXW8
Summary ?
| GeneID | 26259 |
| Symbol | FBXW8 |
| Synonyms | FBW6|FBW8|FBX29|FBXO29|FBXW6 |
| Description | F-box and WD repeat domain containing 8 |
| Reference | MIM:609073|HGNC:HGNC:13597|Ensembl:ENSG00000174989|HPRD:16427|Vega:OTTHUMG00000169329 |
| Gene type | protein-coding |
| Map location | 12q24.22 |
| Pascal p-value | 0.512 |
| Fetal beta | -0.298 |
| eGene | Frontal Cortex BA9 |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| PMID:cooccur | High-throughput literature-search | Systematic search in PubMed for genes co-occurring with SCZ keywords. A total of 3027 genes were included. | |
| Literature | High-throughput literature-search | Co-occurance with Schizophrenia keywords: schizophrenia,schizophrenias | Click to show details |
Section I. Genetics and epigenetics annotation
Section II. Transcriptome annotation
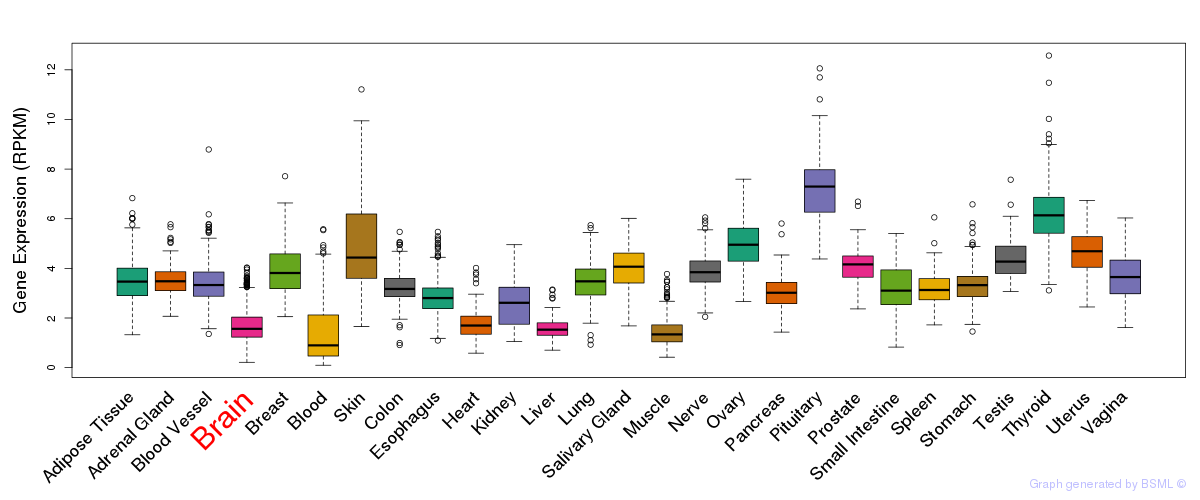
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| SEC11B | 0.87 | 0.82 |
| NUP35 | 0.84 | 0.86 |
| THOC4 | 0.84 | 0.85 |
| E2F5 | 0.83 | 0.85 |
| ERH | 0.83 | 0.89 |
| SFRS3 | 0.83 | 0.87 |
| AC026691.1 | 0.83 | 0.83 |
| KPNA2 | 0.83 | 0.80 |
| RANBP1 | 0.82 | 0.83 |
| DIABLO | 0.82 | 0.85 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| AF347015.33 | -0.66 | -0.79 |
| AF347015.27 | -0.66 | -0.77 |
| AF347015.8 | -0.64 | -0.77 |
| MT-CO2 | -0.64 | -0.75 |
| HLA-F | -0.64 | -0.73 |
| AF347015.15 | -0.63 | -0.78 |
| MT-CYB | -0.62 | -0.76 |
| AF347015.26 | -0.62 | -0.79 |
| TINAGL1 | -0.61 | -0.73 |
| AF347015.2 | -0.61 | -0.77 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| KEGG UBIQUITIN MEDIATED PROTEOLYSIS | 138 | 98 | All SZGR 2.0 genes in this pathway |
| REACTOME IMMUNE SYSTEM | 933 | 616 | All SZGR 2.0 genes in this pathway |
| REACTOME ADAPTIVE IMMUNE SYSTEM | 539 | 350 | All SZGR 2.0 genes in this pathway |
| REACTOME CLASS I MHC MEDIATED ANTIGEN PROCESSING PRESENTATION | 251 | 156 | All SZGR 2.0 genes in this pathway |
| REACTOME ANTIGEN PROCESSING UBIQUITINATION PROTEASOME DEGRADATION | 212 | 129 | All SZGR 2.0 genes in this pathway |
| MARTORIATI MDM4 TARGETS FETAL LIVER DN | 514 | 319 | All SZGR 2.0 genes in this pathway |
| GAVIN FOXP3 TARGETS CLUSTER P6 | 91 | 44 | All SZGR 2.0 genes in this pathway |
| MARTINEZ RB1 TARGETS DN | 543 | 317 | All SZGR 2.0 genes in this pathway |
| MARTINEZ TP53 TARGETS UP | 602 | 364 | All SZGR 2.0 genes in this pathway |
| MARTINEZ RB1 AND TP53 TARGETS UP | 601 | 369 | All SZGR 2.0 genes in this pathway |
| QI PLASMACYTOMA UP | 259 | 185 | All SZGR 2.0 genes in this pathway |
| FONTAINE FOLLICULAR THYROID ADENOMA UP | 75 | 43 | All SZGR 2.0 genes in this pathway |
| FONTAINE PAPILLARY THYROID CARCINOMA DN | 80 | 53 | All SZGR 2.0 genes in this pathway |
| TSUTSUMI FBXW8 TARGETS | 9 | 5 | All SZGR 2.0 genes in this pathway |