Gene Page: GRID2
Summary ?
| GeneID | 2895 |
| Symbol | GRID2 |
| Synonyms | GluD2|SCAR18 |
| Description | glutamate ionotropic receptor delta type subunit 2 |
| Reference | MIM:602368|HGNC:HGNC:4576|Ensembl:ENSG00000152208|HPRD:06781|Vega:OTTHUMG00000130975 |
| Gene type | protein-coding |
| Map location | 4q22 |
| Pascal p-value | 0.01 |
| TADA p-value | 0.017 |
| Fetal beta | -0.289 |
| eGene | Frontal Cortex BA9 Myers' cis & trans |
| Support | CompositeSet Potential synaptic genes |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| DNM:Fromer_2014 | Whole Exome Sequencing analysis | This study reported a WES study of 623 schizophrenia trios, reporting DNMs using genomic DNA. | |
| PMID:cooccur | High-throughput literature-search | Systematic search in PubMed for genes co-occurring with SCZ keywords. A total of 3027 genes were included. | |
| Expression | Meta-analysis of gene expression | P value: 1.638 | |
| Literature | High-throughput literature-search | Co-occurance with Schizophrenia keywords: schizophrenia,schizophrenic,schizophrenias | Click to show details |
| GO_Annotation | Mapping neuro-related keywords to Gene Ontology annotations | Hits with neuro-related keywords: 7 |
Section I. Genetics and epigenetics annotation
 DNM table
DNM table
| Gene | Chromosome | Position | Ref | Alt | Transcript | AA change | Mutation type | Sift | CG46 | Trait | Study |
|---|---|---|---|---|---|---|---|---|---|---|---|
| GRID2 | chr4 | 94411901 | C | G | NM_001510 | p.657T>S | missense | Schizophrenia | DNM:Fromer_2014 |
 eQTL annotation
eQTL annotation
| SNP ID | Chromosome | Position | eGene | Gene Entrez ID | pvalue | qvalue | TSS distance | eQTL type |
|---|---|---|---|---|---|---|---|---|
| rs7370564 | 0 | GRID2 | 2895 | 0.17 | trans |
Section II. Transcriptome annotation
General gene expression (GTEx)
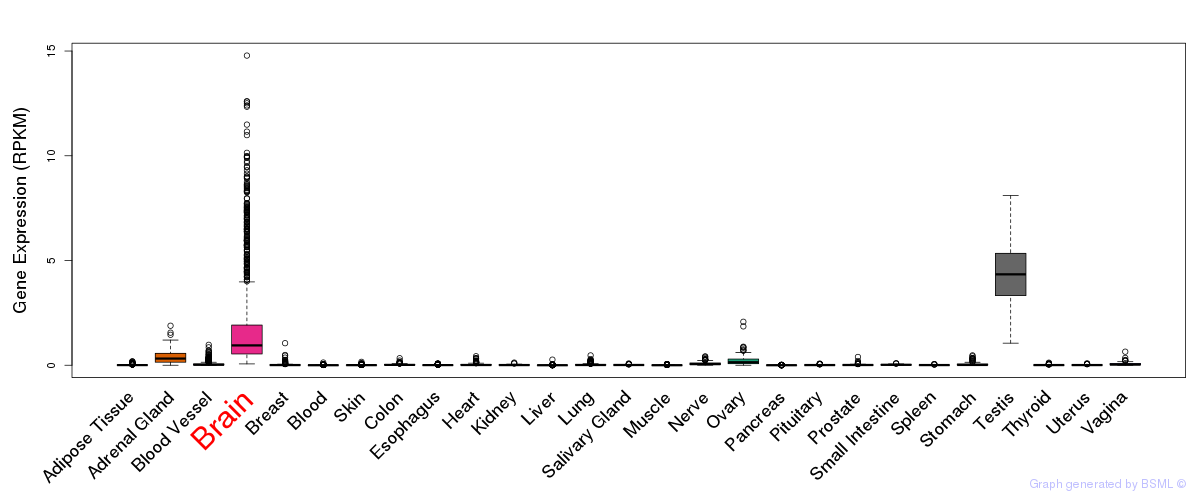
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| SLC7A6 | 0.88 | 0.85 |
| NOL6 | 0.86 | 0.88 |
| MFHAS1 | 0.86 | 0.83 |
| SSBP3 | 0.85 | 0.90 |
| SEZ6 | 0.84 | 0.81 |
| CTNND2 | 0.84 | 0.81 |
| RP1-21O18.1 | 0.84 | 0.79 |
| LZTS1 | 0.83 | 0.79 |
| PTPRS | 0.83 | 0.84 |
| ZNF142 | 0.83 | 0.84 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| SERPINB6 | -0.65 | -0.71 |
| AF347015.31 | -0.61 | -0.74 |
| AL139819.3 | -0.61 | -0.74 |
| AF347015.27 | -0.61 | -0.76 |
| MT-CO2 | -0.60 | -0.75 |
| AF347015.33 | -0.60 | -0.73 |
| BDH2 | -0.60 | -0.70 |
| AF347015.8 | -0.59 | -0.75 |
| MT-CYB | -0.59 | -0.73 |
| S100A16 | -0.58 | -0.71 |
Section III. Gene Ontology annotation
| Molecular function | GO term | Evidence | Neuro keywords | PubMed ID |
|---|---|---|---|---|
| GO:0004970 | ionotropic glutamate receptor activity | IEA | glutamate (GO term level: 7) | - |
| GO:0005234 | extracellular-glutamate-gated ion channel activity | IEA | glutamate (GO term level: 11) | - |
| GO:0004872 | receptor activity | IEA | - | |
| GO:0005515 | protein binding | IEA | - | |
| GO:0005216 | ion channel activity | IEA | - | |
| Biological process | GO term | Evidence | Neuro keywords | PubMed ID |
| GO:0060079 | regulation of excitatory postsynaptic membrane potential | IEA | neuron, Synap (GO term level: 10) | - |
| GO:0007215 | glutamate signaling pathway | TAS | glutamate (GO term level: 7) | 9465309 |
| GO:0035249 | synaptic transmission, glutamatergic | IEA | neuron, glutamate, Synap, Neurotransmitter (GO term level: 8) | - |
| GO:0006811 | ion transport | IEA | - | |
| GO:0060134 | prepulse inhibition | IEA | - | |
| Cellular component | GO term | Evidence | Neuro keywords | PubMed ID |
| GO:0019717 | synaptosome | IEA | Synap, Brain (GO term level: 7) | - |
| GO:0045211 | postsynaptic membrane | IEA | Synap, Neurotransmitter (GO term level: 5) | - |
| GO:0045202 | synapse | IEA | neuron, Synap, Neurotransmitter, Glial (GO term level: 2) | - |
| GO:0016021 | integral to membrane | IEA | - | |
| GO:0005886 | plasma membrane | IEA | - | |
| GO:0005887 | integral to plasma membrane | TAS | 9465309 | |
| GO:0030054 | cell junction | IEA | - |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| KEGG NEUROACTIVE LIGAND RECEPTOR INTERACTION | 272 | 195 | All SZGR 2.0 genes in this pathway |
| KEGG LONG TERM DEPRESSION | 70 | 53 | All SZGR 2.0 genes in this pathway |
| HATADA METHYLATED IN LUNG CANCER UP | 390 | 236 | All SZGR 2.0 genes in this pathway |
| BENPORATH OCT4 TARGETS | 290 | 172 | All SZGR 2.0 genes in this pathway |
| BENPORATH SOX2 TARGETS | 734 | 436 | All SZGR 2.0 genes in this pathway |
| MOOTHA HUMAN MITODB 6 2002 | 429 | 260 | All SZGR 2.0 genes in this pathway |
| HOFFMANN IMMATURE TO MATURE B LYMPHOCYTE DN | 50 | 36 | All SZGR 2.0 genes in this pathway |
| VERHAAK GLIOBLASTOMA PRONEURAL | 177 | 132 | All SZGR 2.0 genes in this pathway |
| GOBERT OLIGODENDROCYTE DIFFERENTIATION DN | 1080 | 713 | All SZGR 2.0 genes in this pathway |