Gene Page: MTMR10
Summary ?
| GeneID | 54893 |
| Symbol | MTMR10 |
| Synonyms | - |
| Description | myotubularin related protein 10 |
| Reference | HGNC:HGNC:25999|Ensembl:ENSG00000166912|HPRD:07895|Vega:OTTHUMG00000175662 |
| Gene type | protein-coding |
| Map location | 15q13.3 |
| Pascal p-value | 0.471 |
| Fetal beta | 0.71 |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CNV:YES | Copy number variation studies | Manual curation | |
| CV:PGCnp | Genome-wide Association Study | GWAS |
Section I. Genetics and epigenetics annotation
Section II. Transcriptome annotation
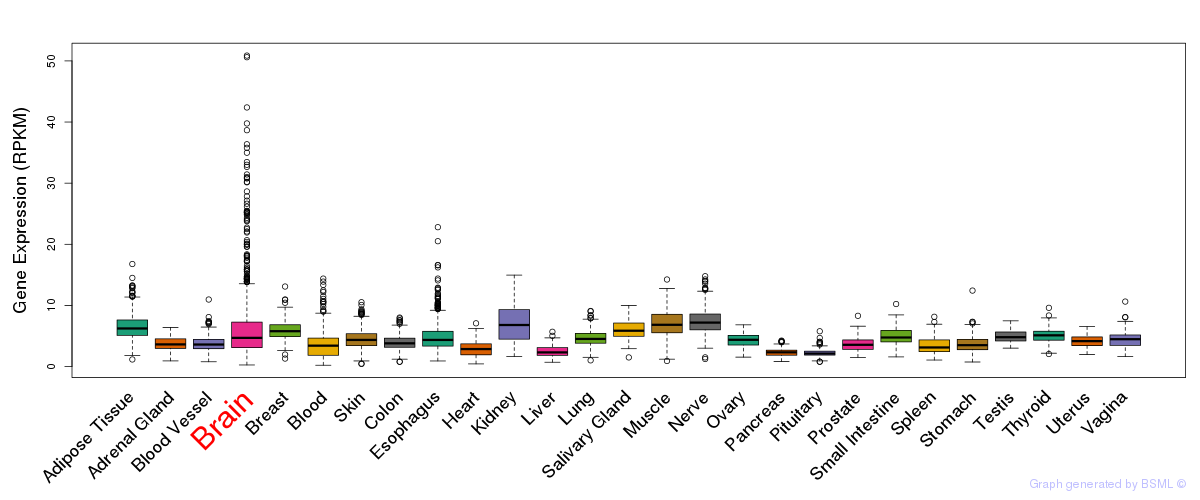
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| THYN1 | 0.66 | 0.67 |
| POLR2C | 0.65 | 0.64 |
| TCTN3 | 0.64 | 0.59 |
| GCA | 0.64 | 0.63 |
| TMEM116 | 0.64 | 0.62 |
| HADH | 0.64 | 0.63 |
| TXNDC12 | 0.63 | 0.56 |
| SHMT1 | 0.63 | 0.58 |
| TSPAN31 | 0.62 | 0.71 |
| RER1 | 0.62 | 0.60 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| AF347015.18 | -0.53 | -0.40 |
| AC100783.1 | -0.48 | -0.38 |
| AC010300.1 | -0.46 | -0.42 |
| AF347015.26 | -0.44 | -0.30 |
| MT-ATP8 | -0.43 | -0.30 |
| AC016705.1 | -0.43 | -0.32 |
| AF347015.8 | -0.42 | -0.28 |
| AF347015.2 | -0.42 | -0.27 |
| AF347015.15 | -0.40 | -0.27 |
| AC073957.1 | -0.40 | -0.34 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| RODRIGUES THYROID CARCINOMA POORLY DIFFERENTIATED DN | 805 | 505 | All SZGR 2.0 genes in this pathway |
| BERENJENO TRANSFORMED BY RHOA FOREVER DN | 31 | 23 | All SZGR 2.0 genes in this pathway |
| BERENJENO TRANSFORMED BY RHOA UP | 536 | 340 | All SZGR 2.0 genes in this pathway |
| RAMALHO STEMNESS UP | 206 | 118 | All SZGR 2.0 genes in this pathway |
| IVANOVA HEMATOPOIESIS STEM CELL AND PROGENITOR | 681 | 420 | All SZGR 2.0 genes in this pathway |
| KYNG DNA DAMAGE BY 4NQO OR GAMMA RADIATION | 15 | 13 | All SZGR 2.0 genes in this pathway |
| KYNG DNA DAMAGE UP | 226 | 164 | All SZGR 2.0 genes in this pathway |
| ROME INSULIN TARGETS IN MUSCLE UP | 442 | 263 | All SZGR 2.0 genes in this pathway |
| GOBERT OLIGODENDROCYTE DIFFERENTIATION DN | 1080 | 713 | All SZGR 2.0 genes in this pathway |