Gene Page: FEM1A
Summary ?
| GeneID | 55527 |
| Symbol | FEM1A |
| Synonyms | EPRAP |
| Description | fem-1 homolog a (C. elegans) |
| Reference | MIM:613538|HGNC:HGNC:16934|HPRD:16888| |
| Gene type | protein-coding |
| Map location | 19p13.3 |
| Pascal p-value | 0.329 |
| Fetal beta | -0.516 |
| DMG | 1 (# studies) |
| eGene | Anterior cingulate cortex BA24 Caudate basal ganglia Cerebellar Hemisphere Cerebellum Cortex Frontal Cortex BA9 Nucleus accumbens basal ganglia Putamen basal ganglia Meta |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| DMG:Nishioka_2013 | Genome-wide DNA methylation analysis | The authors investigated the methylation profiles of DNA in peripheral blood cells from 18 patients with first-episode schizophrenia (FESZ) and from 15 normal controls. | 1 |
Section I. Genetics and epigenetics annotation
 Differentially methylated gene
Differentially methylated gene
| Probe | Chromosome | Position | Nearest gene | P (dis) | Beta (dis) | FDR (dis) | Study |
|---|---|---|---|---|---|---|---|
| cg26336164 | 19 | 4791680 | FEM1A | -0.022 | 0.65 | DMG:Nishioka_2013 |
 eQTL annotation
eQTL annotation
| SNP ID | Chromosome | Position | eGene | Gene Entrez ID | pvalue | qvalue | TSS distance | eQTL type |
|---|---|---|---|---|---|---|---|---|
| rs7252762 | 19 | 4797319 | FEM1A | ENSG00000141965.3 | 1.156E-7 | 0.01 | 5591 | gtex_brain_ba24 |
| rs4807641 | 19 | 4787763 | FEM1A | ENSG00000141965.3 | 4.009E-7 | 0 | -3965 | gtex_brain_putamen_basal |
| rs12609188 | 19 | 4788791 | FEM1A | ENSG00000141965.3 | 8.63E-8 | 0 | -2937 | gtex_brain_putamen_basal |
| rs11085099 | 19 | 4794257 | FEM1A | ENSG00000141965.3 | 1.727E-7 | 0 | 2529 | gtex_brain_putamen_basal |
| rs201690682 | 19 | 4794946 | FEM1A | ENSG00000141965.3 | 3.074E-8 | 0 | 3218 | gtex_brain_putamen_basal |
| rs201811248 | 19 | 4794947 | FEM1A | ENSG00000141965.3 | 3.122E-8 | 0 | 3219 | gtex_brain_putamen_basal |
| rs8111933 | 19 | 4795647 | FEM1A | ENSG00000141965.3 | 3.519E-9 | 0 | 3919 | gtex_brain_putamen_basal |
| rs7252762 | 19 | 4797319 | FEM1A | ENSG00000141965.3 | 6.87E-9 | 0 | 5591 | gtex_brain_putamen_basal |
| rs62113934 | 19 | 4801859 | FEM1A | ENSG00000141965.3 | 9.523E-8 | 0 | 10131 | gtex_brain_putamen_basal |
| rs143319456 | 19 | 4802424 | FEM1A | ENSG00000141965.3 | 4.81E-9 | 0 | 10696 | gtex_brain_putamen_basal |
| rs62113935 | 19 | 4802542 | FEM1A | ENSG00000141965.3 | 2.022E-7 | 0 | 10814 | gtex_brain_putamen_basal |
| rs8113352 | 19 | 4804212 | FEM1A | ENSG00000141965.3 | 2.075E-7 | 0 | 12484 | gtex_brain_putamen_basal |
| rs8102626 | 19 | 4804524 | FEM1A | ENSG00000141965.3 | 4.834E-7 | 0 | 12796 | gtex_brain_putamen_basal |
| rs75348862 | 19 | 4805003 | FEM1A | ENSG00000141965.3 | 4.959E-7 | 0 | 13275 | gtex_brain_putamen_basal |
| rs62113936 | 19 | 4805095 | FEM1A | ENSG00000141965.3 | 5.131E-7 | 0 | 13367 | gtex_brain_putamen_basal |
| rs7253085 | 19 | 4806381 | FEM1A | ENSG00000141965.3 | 1.089E-6 | 0 | 14653 | gtex_brain_putamen_basal |
| rs62113939 | 19 | 4806705 | FEM1A | ENSG00000141965.3 | 1.008E-6 | 0 | 14977 | gtex_brain_putamen_basal |
| rs62113940 | 19 | 4807247 | FEM1A | ENSG00000141965.3 | 9.553E-7 | 0 | 15519 | gtex_brain_putamen_basal |
| rs8110678 | 19 | 4807893 | FEM1A | ENSG00000141965.3 | 9.365E-7 | 0 | 16165 | gtex_brain_putamen_basal |
| rs8100815 | 19 | 4808305 | FEM1A | ENSG00000141965.3 | 4.453E-7 | 0 | 16577 | gtex_brain_putamen_basal |
Section II. Transcriptome annotation
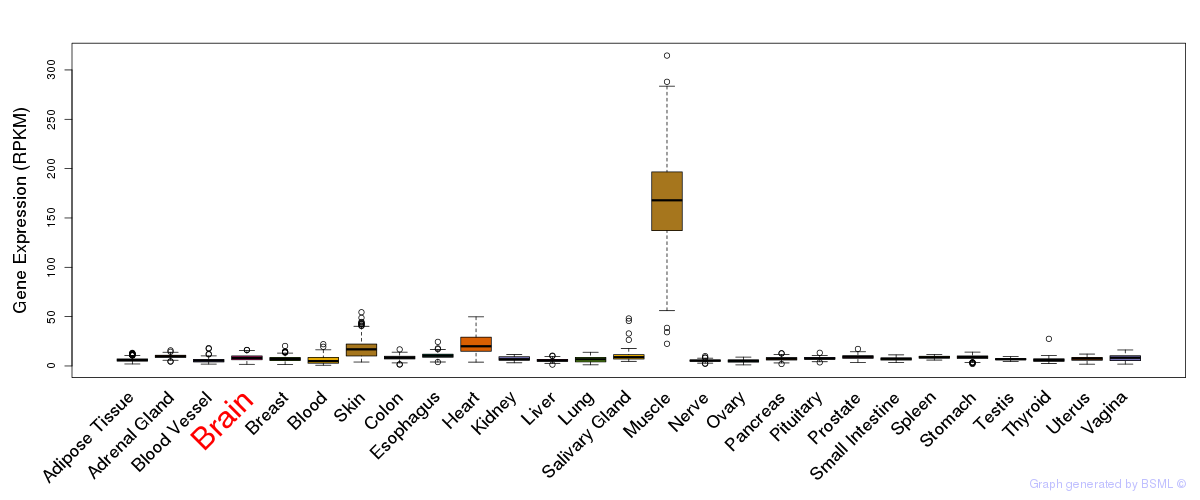
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| ARVCF | 0.91 | 0.88 |
| PRR14 | 0.91 | 0.88 |
| MED25 | 0.91 | 0.88 |
| ZNF446 | 0.90 | 0.90 |
| RALGDS | 0.90 | 0.88 |
| TELO2 | 0.89 | 0.87 |
| GRAMD1A | 0.89 | 0.84 |
| WIZ | 0.89 | 0.87 |
| TOR2A | 0.89 | 0.85 |
| KIAA1602 | 0.89 | 0.89 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| C5orf53 | -0.70 | -0.70 |
| ACOT13 | -0.66 | -0.68 |
| AF347015.27 | -0.66 | -0.73 |
| S100B | -0.64 | -0.69 |
| AF347015.31 | -0.64 | -0.72 |
| MT-CO2 | -0.62 | -0.72 |
| AF347015.21 | -0.61 | -0.79 |
| AF347015.33 | -0.60 | -0.68 |
| EAF2 | -0.60 | -0.66 |
| AF347015.8 | -0.60 | -0.71 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| BIDUS METASTASIS UP | 214 | 134 | All SZGR 2.0 genes in this pathway |
| BENPORATH NANOG TARGETS | 988 | 594 | All SZGR 2.0 genes in this pathway |
| BENPORATH SOX2 TARGETS | 734 | 436 | All SZGR 2.0 genes in this pathway |
| BYSTRYKH HEMATOPOIESIS STEM CELL QTL TRANS | 882 | 572 | All SZGR 2.0 genes in this pathway |
| VIETOR IFRD1 TARGETS | 23 | 13 | All SZGR 2.0 genes in this pathway |
| CHENG IMPRINTED BY ESTRADIOL | 110 | 68 | All SZGR 2.0 genes in this pathway |
| ACEVEDO NORMAL TISSUE ADJACENT TO LIVER TUMOR DN | 354 | 216 | All SZGR 2.0 genes in this pathway |
| BOCHKIS FOXA2 TARGETS | 425 | 261 | All SZGR 2.0 genes in this pathway |
| ZWANG TRANSIENTLY UP BY 1ST EGF PULSE ONLY | 1839 | 928 | All SZGR 2.0 genes in this pathway |