Gene Page: SPR
Summary ?
| GeneID | 6697 |
| Symbol | SPR |
| Synonyms | SDR38C1 |
| Description | sepiapterin reductase (7,8-dihydrobiopterin:NADP+ oxidoreductase) |
| Reference | MIM:182125|HGNC:HGNC:11257|Ensembl:ENSG00000116096|HPRD:01632|Vega:OTTHUMG00000129777 |
| Gene type | protein-coding |
| Map location | 2p14-p12 |
| Pascal p-value | 0.011 |
| Fetal beta | -1.433 |
| eGene | Myers' cis & trans |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| PMID:cooccur | High-throughput literature-search | Systematic search in PubMed for genes co-occurring with SCZ keywords. A total of 3027 genes were included. | |
| Literature | High-throughput literature-search | Co-occurance with Schizophrenia keywords: schizophrenia,schizophrenic,schizophrenias | Click to show details |
Section I. Genetics and epigenetics annotation
 eQTL annotation
eQTL annotation
| SNP ID | Chromosome | Position | eGene | Gene Entrez ID | pvalue | qvalue | TSS distance | eQTL type |
|---|---|---|---|---|---|---|---|---|
| rs17022908 | chr2 | 38783057 | SPR | 6697 | 0.13 | trans | ||
| rs7607798 | chr2 | 102043108 | SPR | 6697 | 0.13 | trans | ||
| rs10178499 | chr2 | 127937434 | SPR | 6697 | 0.17 | trans | ||
| rs2461463 | chr2 | 128708062 | SPR | 6697 | 0.07 | trans | ||
| rs1397611 | chr4 | 120225347 | SPR | 6697 | 0.16 | trans | ||
| rs11722940 | chr4 | 120233215 | SPR | 6697 | 0.13 | trans | ||
| rs17595790 | chr4 | 120276219 | SPR | 6697 | 0.12 | trans | ||
| rs7706288 | chr5 | 141864248 | SPR | 6697 | 0 | trans | ||
| rs1898661 | chr5 | 153338309 | SPR | 6697 | 0.12 | trans | ||
| rs11996557 | chr8 | 102901392 | SPR | 6697 | 0.15 | trans | ||
| rs16868715 | chr8 | 102922408 | SPR | 6697 | 0.08 | trans | ||
| rs10739219 | chr9 | 108938328 | SPR | 6697 | 0.18 | trans | ||
| rs10980452 | chr9 | 113332327 | SPR | 6697 | 0.14 | trans | ||
| rs6478513 | chr9 | 124238226 | SPR | 6697 | 0.11 | trans | ||
| rs17152865 | chr10 | 13171768 | SPR | 6697 | 0.04 | trans | ||
| snp_a-2055321 | 0 | SPR | 6697 | 0.17 | trans | |||
| rs11030327 | chr11 | 3956554 | SPR | 6697 | 0.07 | trans | ||
| rs17007452 | chr12 | 81243404 | SPR | 6697 | 0.01 | trans | ||
| rs3899870 | chr13 | 97205717 | SPR | 6697 | 0.02 | trans | ||
| rs9300361 | chr13 | 97210272 | SPR | 6697 | 0.14 | trans | ||
| rs3784005 | chr14 | 76077230 | SPR | 6697 | 0.05 | trans | ||
| rs17757723 | chr14 | 79323591 | SPR | 6697 | 0.06 | trans | ||
| rs2829739 | chr21 | 26791883 | SPR | 6697 | 0.06 | trans | ||
| rs216780 | chr21 | 27324318 | SPR | 6697 | 0.16 | trans | ||
| rs2829996 | chr21 | 27326755 | SPR | 6697 | 0.06 | trans | ||
| rs1001022 | chr22 | 26403487 | SPR | 6697 | 0.04 | trans |
Section II. Transcriptome annotation
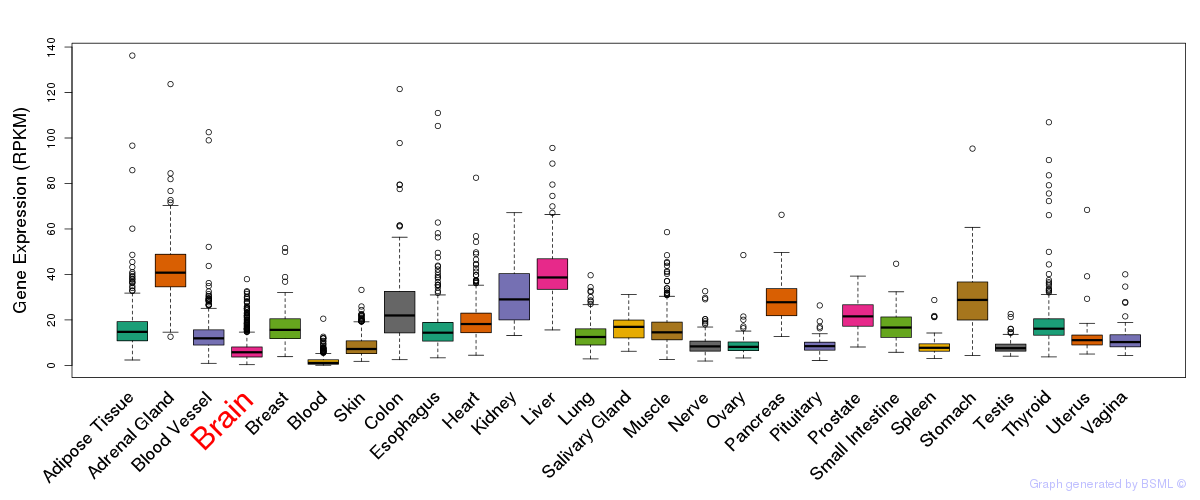
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| TAP1 | 0.72 | 0.47 |
| IRF9 | 0.71 | 0.43 |
| MX1 | 0.70 | 0.40 |
| IFIT3 | 0.70 | 0.38 |
| IFITM3 | 0.70 | 0.35 |
| SP110 | 0.69 | 0.54 |
| IFIT1 | 0.69 | 0.31 |
| PLSCR1 | 0.69 | 0.45 |
| IFI44L | 0.69 | 0.46 |
| PARP9 | 0.68 | 0.52 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| OR2W3 | -0.35 | -0.38 |
| PRRT2 | -0.35 | -0.37 |
| PKNOX2 | -0.35 | -0.36 |
| LRRC4C | -0.35 | -0.38 |
| SLITRK5 | -0.35 | -0.36 |
| CSMD1 | -0.34 | -0.36 |
| KIAA1239 | -0.34 | -0.40 |
| MYLIP | -0.34 | -0.38 |
| ARPP-21 | -0.34 | -0.38 |
| CHD5 | -0.34 | -0.36 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| KEGG FOLATE BIOSYNTHESIS | 11 | 10 | All SZGR 2.0 genes in this pathway |
| REACTOME TETRAHYDROBIOPTERIN BH4 SYNTHESIS RECYCLING SALVAGE AND REGULATION | 13 | 9 | All SZGR 2.0 genes in this pathway |
| REACTOME ENOS ACTIVATION AND REGULATION | 20 | 14 | All SZGR 2.0 genes in this pathway |
| WATANABE RECTAL CANCER RADIOTHERAPY RESPONSIVE DN | 92 | 51 | All SZGR 2.0 genes in this pathway |
| GRAESSMANN APOPTOSIS BY DOXORUBICIN UP | 1142 | 669 | All SZGR 2.0 genes in this pathway |
| RASHI RESPONSE TO IONIZING RADIATION 3 | 48 | 31 | All SZGR 2.0 genes in this pathway |
| DACOSTA UV RESPONSE VIA ERCC3 UP | 309 | 199 | All SZGR 2.0 genes in this pathway |
| FERREIRA EWINGS SARCOMA UNSTABLE VS STABLE DN | 98 | 59 | All SZGR 2.0 genes in this pathway |
| SCHAEFFER PROSTATE DEVELOPMENT 6HR DN | 514 | 330 | All SZGR 2.0 genes in this pathway |
| IVANOVA HEMATOPOIESIS EARLY PROGENITOR | 532 | 309 | All SZGR 2.0 genes in this pathway |
| BROWNE HCMV INFECTION 24HR UP | 148 | 96 | All SZGR 2.0 genes in this pathway |
| DOUGLAS BMI1 TARGETS UP | 566 | 371 | All SZGR 2.0 genes in this pathway |
| KRIGE RESPONSE TO TOSEDOSTAT 6HR DN | 911 | 527 | All SZGR 2.0 genes in this pathway |
| KRIGE RESPONSE TO TOSEDOSTAT 24HR DN | 1011 | 592 | All SZGR 2.0 genes in this pathway |
| BONOME OVARIAN CANCER SURVIVAL OPTIMAL DEBULKING | 246 | 152 | All SZGR 2.0 genes in this pathway |
| TOOKER GEMCITABINE RESISTANCE DN | 122 | 84 | All SZGR 2.0 genes in this pathway |
| YOSHIMURA MAPK8 TARGETS UP | 1305 | 895 | All SZGR 2.0 genes in this pathway |