Gene Page: WDR59
Summary ?
| GeneID | 79726 |
| Symbol | WDR59 |
| Synonyms | FP977 |
| Description | WD repeat domain 59 |
| Reference | HGNC:HGNC:25706|HPRD:13351| |
| Gene type | protein-coding |
| Map location | 16q23.1 |
| Pascal p-value | 0.154 |
| Sherlock p-value | 0.295 |
| Fetal beta | 0.285 |
| eGene | Cerebellar Hemisphere Cerebellum Myers' cis & trans |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:GWASdb | Genome-wide Association Studies | GWASdb records for schizophrenia | |
| CV:PGCnp | Genome-wide Association Study | GWAS |
Section I. Genetics and epigenetics annotation
 eQTL annotation
eQTL annotation
| SNP ID | Chromosome | Position | eGene | Gene Entrez ID | pvalue | qvalue | TSS distance | eQTL type |
|---|---|---|---|---|---|---|---|---|
| rs163352 | chr3 | 3098040 | WDR59 | 79726 | 0.13 | trans |
Section II. Transcriptome annotation
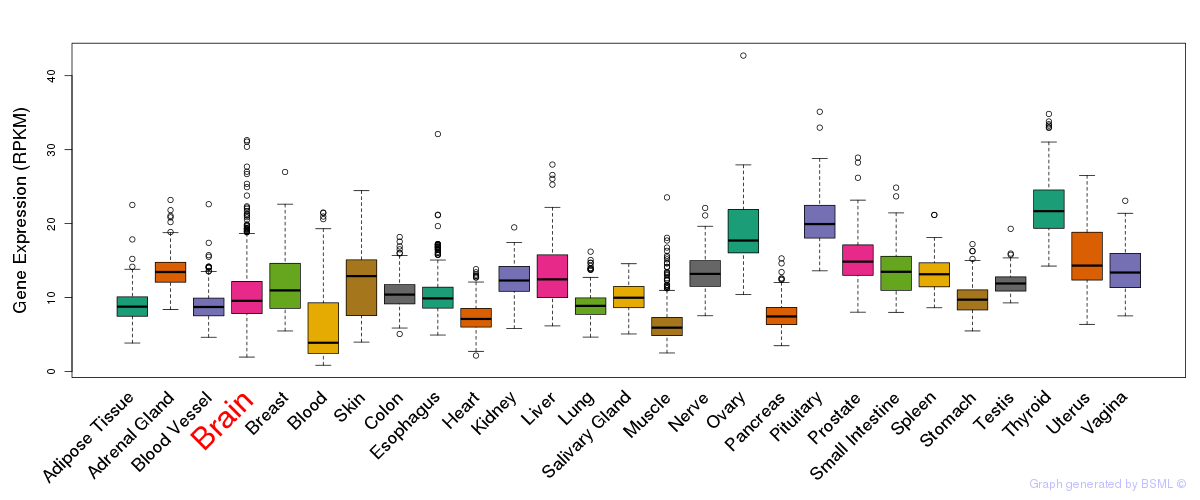
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| KIAA0317 | 0.92 | 0.93 |
| ULK2 | 0.92 | 0.94 |
| GNPTAB | 0.91 | 0.93 |
| NOLC1 | 0.90 | 0.92 |
| FBXW2 | 0.90 | 0.92 |
| TACC2 | 0.90 | 0.93 |
| FHOD3 | 0.90 | 0.92 |
| RNF145 | 0.90 | 0.92 |
| ALS2 | 0.89 | 0.93 |
| ACACA | 0.89 | 0.93 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| AF347015.31 | -0.79 | -0.83 |
| MT-CO2 | -0.78 | -0.84 |
| AF347015.27 | -0.77 | -0.81 |
| FXYD1 | -0.76 | -0.79 |
| AF347015.33 | -0.76 | -0.78 |
| MT-CYB | -0.75 | -0.79 |
| AF347015.8 | -0.74 | -0.81 |
| S100B | -0.74 | -0.77 |
| AF347015.21 | -0.74 | -0.85 |
| AC021016.1 | -0.73 | -0.79 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| MULLIGHAN MLL SIGNATURE 1 DN | 242 | 165 | All SZGR 2.0 genes in this pathway |
| MULLIGHAN MLL SIGNATURE 2 DN | 281 | 186 | All SZGR 2.0 genes in this pathway |
| ODONNELL TFRC TARGETS UP | 456 | 228 | All SZGR 2.0 genes in this pathway |
| PROVENZANI METASTASIS UP | 194 | 112 | All SZGR 2.0 genes in this pathway |
| MARSON BOUND BY FOXP3 STIMULATED | 1022 | 619 | All SZGR 2.0 genes in this pathway |
| FOSTER TOLERANT MACROPHAGE DN | 409 | 268 | All SZGR 2.0 genes in this pathway |
| OUILLETTE CLL 13Q14 DELETION UP | 74 | 40 | All SZGR 2.0 genes in this pathway |
| HOELZEL NF1 TARGETS DN | 115 | 73 | All SZGR 2.0 genes in this pathway |
| PURBEY TARGETS OF CTBP1 AND SATB1 UP | 87 | 52 | All SZGR 2.0 genes in this pathway |
| RATTENBACHER BOUND BY CELF1 | 467 | 251 | All SZGR 2.0 genes in this pathway |