Gene Page: ATF7IP2
Summary ?
| GeneID | 80063 |
| Symbol | ATF7IP2 |
| Synonyms | MCAF2 |
| Description | activating transcription factor 7 interacting protein 2 |
| Reference | MIM:613645|HGNC:HGNC:20397|Ensembl:ENSG00000166669|HPRD:16517|Vega:OTTHUMG00000129749 |
| Gene type | protein-coding |
| Map location | 16p13.13 |
| Pascal p-value | 0.437 |
| Sherlock p-value | 0.13 |
| Fetal beta | -0.885 |
| eGene | Cerebellum Nucleus accumbens basal ganglia Myers' cis & trans |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| PMID:cooccur | High-throughput literature-search | Systematic search in PubMed for genes co-occurring with SCZ keywords. A total of 3027 genes were included. | |
| Literature | High-throughput literature-search | Co-occurance with Schizophrenia keywords: schizophrenia,schizophrenias | Click to show details |
Section I. Genetics and epigenetics annotation
 eQTL annotation
eQTL annotation
| SNP ID | Chromosome | Position | eGene | Gene Entrez ID | pvalue | qvalue | TSS distance | eQTL type |
|---|---|---|---|---|---|---|---|---|
| rs12029998 | 0 | ATF7IP2 | 80063 | 6.313E-4 | trans | |||
| rs12072589 | chr1 | 102074474 | ATF7IP2 | 80063 | 0.03 | trans | ||
| rs11164251 | chr1 | 102079419 | ATF7IP2 | 80063 | 0.05 | trans | ||
| rs6707600 | chr2 | 24521765 | ATF7IP2 | 80063 | 0.11 | trans | ||
| rs1406236 | chr2 | 29499441 | ATF7IP2 | 80063 | 0.12 | trans | ||
| rs1406237 | chr2 | 29499624 | ATF7IP2 | 80063 | 0.12 | trans | ||
| rs4666189 | chr2 | 29502689 | ATF7IP2 | 80063 | 0.12 | trans | ||
| rs16840841 | chr2 | 157641525 | ATF7IP2 | 80063 | 0.03 | trans | ||
| rs7560489 | chr2 | 158145388 | ATF7IP2 | 80063 | 0 | trans | ||
| rs7593761 | chr2 | 197275602 | ATF7IP2 | 80063 | 0.12 | trans | ||
| rs16847759 | chr3 | 168869746 | ATF7IP2 | 80063 | 0.03 | trans | ||
| rs12641054 | chr4 | 113071694 | ATF7IP2 | 80063 | 0.01 | trans | ||
| rs11956817 | chr5 | 98842089 | ATF7IP2 | 80063 | 0.12 | trans | ||
| rs1422148 | chr5 | 107526837 | ATF7IP2 | 80063 | 0 | trans | ||
| rs12659121 | chr5 | 107660002 | ATF7IP2 | 80063 | 0 | trans | ||
| rs17161316 | chr5 | 107756153 | ATF7IP2 | 80063 | 0 | trans | ||
| rs12524898 | chr6 | 67967596 | ATF7IP2 | 80063 | 0.15 | trans | ||
| rs10969802 | chr9 | 30636427 | ATF7IP2 | 80063 | 0.07 | trans | ||
| rs10969816 | chr9 | 30661611 | ATF7IP2 | 80063 | 0.07 | trans | ||
| rs12350524 | chr9 | 30743421 | ATF7IP2 | 80063 | 0.07 | trans | ||
| rs12343068 | chr9 | 30783524 | ATF7IP2 | 80063 | 0.07 | trans | ||
| rs10511868 | chr9 | 30816504 | ATF7IP2 | 80063 | 0.07 | trans | ||
| rs10868543 | chr9 | 89852686 | ATF7IP2 | 80063 | 1.01E-8 | trans | ||
| rs10776917 | chr9 | 137858478 | ATF7IP2 | 80063 | 0.16 | trans | ||
| rs1335812 | chr10 | 24119275 | ATF7IP2 | 80063 | 0.13 | trans | ||
| rs602658 | chr10 | 36007778 | ATF7IP2 | 80063 | 0.01 | trans | ||
| rs11018448 | chr11 | 88793318 | ATF7IP2 | 80063 | 0.07 | trans | ||
| rs8181526 | chr11 | 88797520 | ATF7IP2 | 80063 | 0.03 | trans | ||
| rs17109555 | chr14 | 40132840 | ATF7IP2 | 80063 | 0.13 | trans | ||
| rs10492782 | chr16 | 13210707 | ATF7IP2 | 80063 | 0 | trans |
Section II. Transcriptome annotation
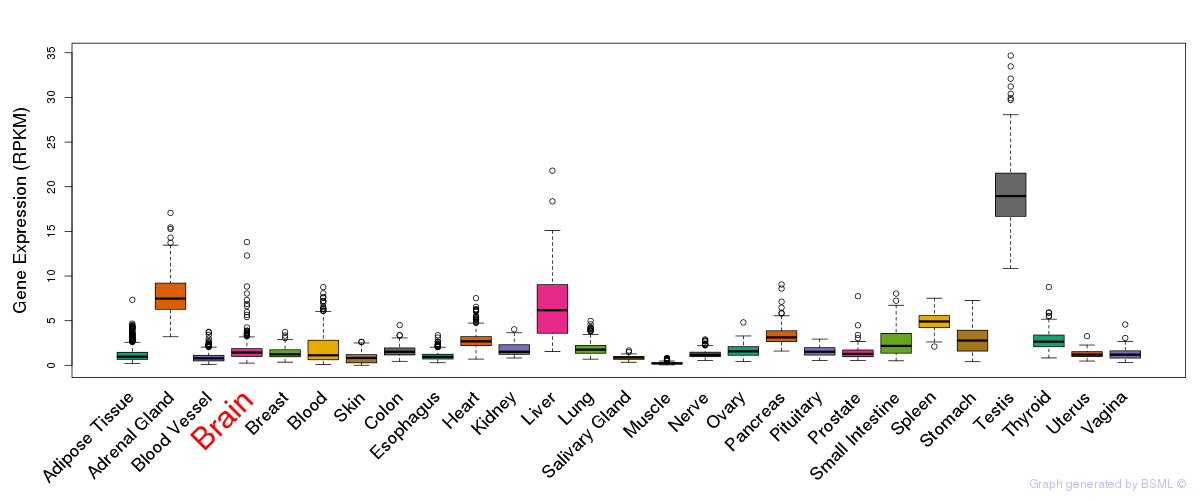
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| MRPL1 | 0.62 | 0.59 |
| C10orf84 | 0.59 | 0.57 |
| GGH | 0.58 | 0.52 |
| GLRX3 | 0.57 | 0.55 |
| TXNDC9 | 0.57 | 0.55 |
| OLA1 | 0.56 | 0.54 |
| NFU1 | 0.56 | 0.53 |
| C3orf14 | 0.56 | 0.58 |
| PPP2R3C | 0.56 | 0.54 |
| NIPSNAP3A | 0.56 | 0.53 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| PPP1R13L | -0.37 | -0.40 |
| MT-CYB | -0.36 | -0.40 |
| AF347015.26 | -0.35 | -0.41 |
| AF347015.2 | -0.35 | -0.41 |
| AF347015.8 | -0.35 | -0.38 |
| TRIM56 | -0.34 | -0.37 |
| CASKIN2 | -0.34 | -0.34 |
| ENG | -0.34 | -0.35 |
| MT-CO2 | -0.34 | -0.37 |
| FAM38A | -0.34 | -0.34 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| EBAUER TARGETS OF PAX3 FOXO1 FUSION UP | 207 | 128 | All SZGR 2.0 genes in this pathway |
| RICKMAN TUMOR DIFFERENTIATED WELL VS POORLY UP | 236 | 139 | All SZGR 2.0 genes in this pathway |
| ROZANOV MMP14 TARGETS UP | 266 | 171 | All SZGR 2.0 genes in this pathway |
| SUZUKI RESPONSE TO TSA AND DECITABINE 1B | 23 | 14 | All SZGR 2.0 genes in this pathway |
| HELLER SILENCED BY METHYLATION UP | 282 | 183 | All SZGR 2.0 genes in this pathway |
| HELLER HDAC TARGETS SILENCED BY METHYLATION UP | 461 | 298 | All SZGR 2.0 genes in this pathway |
| MARTINEZ RB1 TARGETS DN | 543 | 317 | All SZGR 2.0 genes in this pathway |
| MARTINEZ TP53 TARGETS DN | 593 | 372 | All SZGR 2.0 genes in this pathway |
| MARTINEZ RB1 AND TP53 TARGETS DN | 591 | 366 | All SZGR 2.0 genes in this pathway |
| LEE DIFFERENTIATING T LYMPHOCYTE | 200 | 115 | All SZGR 2.0 genes in this pathway |
| WANG RESPONSE TO GSK3 INHIBITOR SB216763 UP | 397 | 206 | All SZGR 2.0 genes in this pathway |
| SMIRNOV RESPONSE TO IR 2HR DN | 55 | 35 | All SZGR 2.0 genes in this pathway |