Gene Page: ZIC4
Summary ?
| GeneID | 84107 |
| Symbol | ZIC4 |
| Synonyms | - |
| Description | Zic family member 4 |
| Reference | MIM:608948|HGNC:HGNC:20393|Ensembl:ENSG00000174963|HPRD:12340|Vega:OTTHUMG00000037281 |
| Gene type | protein-coding |
| Map location | 3q24 |
| Pascal p-value | 0.001 |
| Fetal beta | 0.225 |
| DMG | 1 (# studies) |
| eGene | Myers' cis & trans |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| DMG:Wockner_2014 | Genome-wide DNA methylation analysis | This dataset includes 4641 differentially methylated probes corresponding to 2929 unique genes between schizophrenia patients (n=24) and controls (n=24). | 2 |
| PMID:cooccur | High-throughput literature-search | Systematic search in PubMed for genes co-occurring with SCZ keywords. A total of 3027 genes were included. | |
| Literature | High-throughput literature-search | Co-occurance with Schizophrenia keywords: schizophrenia,schizophrenias | Click to show details |
Section I. Genetics and epigenetics annotation
 Differentially methylated gene
Differentially methylated gene
| Probe | Chromosome | Position | Nearest gene | P (dis) | Beta (dis) | FDR (dis) | Study |
|---|---|---|---|---|---|---|---|
| cg18292434 | 3 | 147112541 | ZIC4 | 4.72E-6 | 0.473 | 0.01 | DMG:Wockner_2014 |
| cg12892506 | 3 | 147113918 | ZIC4 | 3.153E-4 | 0.692 | 0.04 | DMG:Wockner_2014 |
 eQTL annotation
eQTL annotation
| SNP ID | Chromosome | Position | eGene | Gene Entrez ID | pvalue | qvalue | TSS distance | eQTL type |
|---|---|---|---|---|---|---|---|---|
| rs13447530 | chr1 | 91983798 | ZIC4 | 84107 | 0.19 | trans | ||
| rs16829545 | chr2 | 151977407 | ZIC4 | 84107 | 1.667E-9 | trans | ||
| rs7584986 | chr2 | 184111432 | ZIC4 | 84107 | 0.12 | trans | ||
| rs6741060 | chr2 | 217169088 | ZIC4 | 84107 | 0.06 | trans | ||
| rs9810143 | chr3 | 5060209 | ZIC4 | 84107 | 0.01 | trans | ||
| rs6996695 | chr8 | 77540580 | ZIC4 | 84107 | 0.2 | trans | ||
| rs16955618 | chr15 | 29937543 | ZIC4 | 84107 | 3.727E-9 | trans | ||
| rs17681878 | chr18 | 34767223 | ZIC4 | 84107 | 0.01 | trans |
Section II. Transcriptome annotation
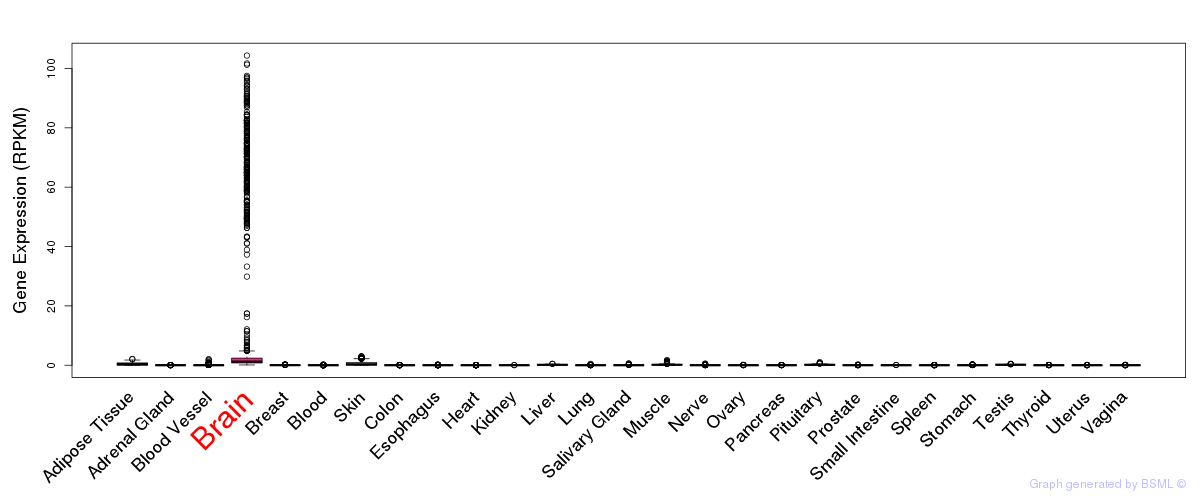
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
No co-expressed genes in brain regions
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| BENPORATH SUZ12 TARGETS | 1038 | 678 | All SZGR 2.0 genes in this pathway |
| BENPORATH EED TARGETS | 1062 | 725 | All SZGR 2.0 genes in this pathway |
| BENPORATH ES WITH H3K27ME3 | 1118 | 744 | All SZGR 2.0 genes in this pathway |
| BENPORATH PRC2 TARGETS | 652 | 441 | All SZGR 2.0 genes in this pathway |
| KONDO PROSTATE CANCER WITH H3K27ME3 | 196 | 93 | All SZGR 2.0 genes in this pathway |
| MIKKELSEN MEF ICP WITH H3K27ME3 | 206 | 108 | All SZGR 2.0 genes in this pathway |
| MIKKELSEN IPS ICP WITH H3K27ME3 | 54 | 32 | All SZGR 2.0 genes in this pathway |
| MIKKELSEN ES ICP WITH H3K27ME3 | 42 | 27 | All SZGR 2.0 genes in this pathway |
| MARTENS TRETINOIN RESPONSE UP | 857 | 456 | All SZGR 2.0 genes in this pathway |
| BRUINS UVC RESPONSE VIA TP53 GROUP A | 898 | 516 | All SZGR 2.0 genes in this pathway |