Gene Page: ACCS
Summary ?
| GeneID | 84680 |
| Symbol | ACCS |
| Synonyms | ACS|PHACS |
| Description | 1-aminocyclopropane-1-carboxylate synthase homolog (inactive) |
| Reference | MIM:608405|HGNC:HGNC:23989|Ensembl:ENSG00000110455|HPRD:10524|Vega:OTTHUMG00000166427 |
| Gene type | protein-coding |
| Map location | 11p11 |
| Pascal p-value | 0.088 |
| Fetal beta | -0.2 |
| eGene | Caudate basal ganglia Cerebellar Hemisphere Cerebellum Cortex Frontal Cortex BA9 Hippocampus Hypothalamus Nucleus accumbens basal ganglia Putamen basal ganglia Myers' cis & trans Meta |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| PMID:cooccur | High-throughput literature-search | Systematic search in PubMed for genes co-occurring with SCZ keywords. A total of 3027 genes were included. | |
| Literature | High-throughput literature-search | Co-occurance with Schizophrenia keywords: schizophrenia,schizophrenic,schizophrenias | Click to show details |
Section I. Genetics and epigenetics annotation
 eQTL annotation
eQTL annotation
| SNP ID | Chromosome | Position | eGene | Gene Entrez ID | pvalue | qvalue | TSS distance | eQTL type |
|---|---|---|---|---|---|---|---|---|
| rs10994209 | chr10 | 61877705 | ACCS | 84680 | 0.18 | trans | ||
| rs178501 | 11 | 44081716 | ACCS | ENSG00000110455.9 | 3.621E-8 | 0 | -5759 | gtex_brain_putamen_basal |
| rs178502 | 11 | 44082356 | ACCS | ENSG00000110455.9 | 2.386E-6 | 0 | -5119 | gtex_brain_putamen_basal |
| rs178504 | 11 | 44083650 | ACCS | ENSG00000110455.9 | 1.731E-9 | 0 | -3825 | gtex_brain_putamen_basal |
| . | 11 | 44084573 | ACCS | ENSG00000110455.9 | 2.613E-6 | 0 | -2902 | gtex_brain_putamen_basal |
| rs178505 | 11 | 44084849 | ACCS | ENSG00000110455.9 | 7.967E-10 | 0 | -2626 | gtex_brain_putamen_basal |
| rs178506 | 11 | 44085369 | ACCS | ENSG00000110455.9 | 4.106E-9 | 0 | -2106 | gtex_brain_putamen_basal |
| rs2074038 | 11 | 44087989 | ACCS | ENSG00000110455.9 | 2.407E-6 | 0 | 514 | gtex_brain_putamen_basal |
| rs3923808 | 11 | 44117122 | ACCS | ENSG00000110455.9 | 1.766E-6 | 0 | 29647 | gtex_brain_putamen_basal |
| rs12365753 | 11 | 44117424 | ACCS | ENSG00000110455.9 | 1.625E-6 | 0 | 29949 | gtex_brain_putamen_basal |
| rs10838235 | 11 | 44131354 | ACCS | ENSG00000110455.9 | 2.762E-6 | 0 | 43879 | gtex_brain_putamen_basal |
| rs7102268 | 11 | 44144643 | ACCS | ENSG00000110455.9 | 1.195E-6 | 0 | 57168 | gtex_brain_putamen_basal |
Section II. Transcriptome annotation
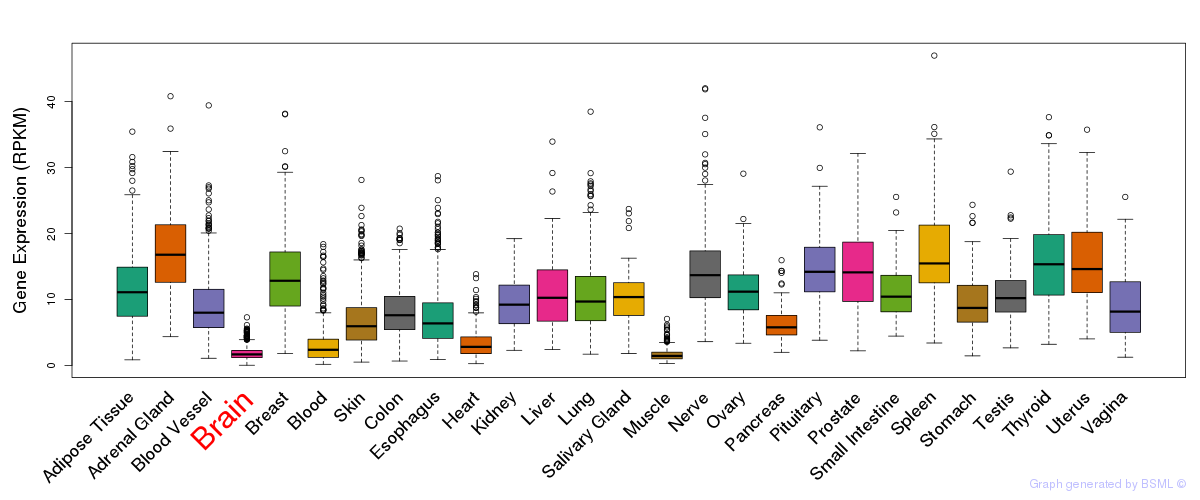
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
No co-expressed genes in brain regions
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| ACEVEDO LIVER CANCER DN | 540 | 340 | All SZGR 2.0 genes in this pathway |
| ACEVEDO LIVER TUMOR VS NORMAL ADJACENT TISSUE DN | 274 | 165 | All SZGR 2.0 genes in this pathway |
| YOSHIMURA MAPK8 TARGETS UP | 1305 | 895 | All SZGR 2.0 genes in this pathway |
| CHANG CORE SERUM RESPONSE DN | 209 | 137 | All SZGR 2.0 genes in this pathway |
| BRUINS UVC RESPONSE LATE | 1137 | 655 | All SZGR 2.0 genes in this pathway |