Gene Page: ZNF432
Summary ?
| GeneID | 9668 |
| Symbol | ZNF432 |
| Synonyms | - |
| Description | zinc finger protein 432 |
| Reference | HGNC:HGNC:20810|Ensembl:ENSG00000256087|HPRD:11716|Vega:OTTHUMG00000182552 |
| Gene type | protein-coding |
| Map location | 19q13.41 |
| Pascal p-value | 0.175 |
| Sherlock p-value | 0.352 |
| DMG | 2 (# studies) |
| eGene | Cerebellum Meta |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| DMG:Jaffe_2016 | Genome-wide DNA methylation analysis | This dataset includes 2,104 probes/CpGs associated with SZ patients (n=108) compared to 136 controls at Bonferroni-adjusted P < 0.05. | 2 |
| DMG:Nishioka_2013 | Genome-wide DNA methylation analysis | The authors investigated the methylation profiles of DNA in peripheral blood cells from 18 patients with first-episode schizophrenia (FESZ) and from 15 normal controls. | 2 |
Section I. Genetics and epigenetics annotation
 Differentially methylated gene
Differentially methylated gene
| Probe | Chromosome | Position | Nearest gene | P (dis) | Beta (dis) | FDR (dis) | Study |
|---|---|---|---|---|---|---|---|
| cg02729352 | 19 | 52552443 | ZNF432 | -0.019 | 0.92 | DMG:Nishioka_2013 | |
| cg11945235 | 19 | 52552025 | ZNF432 | 5.47E-9 | -0.017 | 3.01E-6 | DMG:Jaffe_2016 |
Section II. Transcriptome annotation
General gene expression (GTEx)
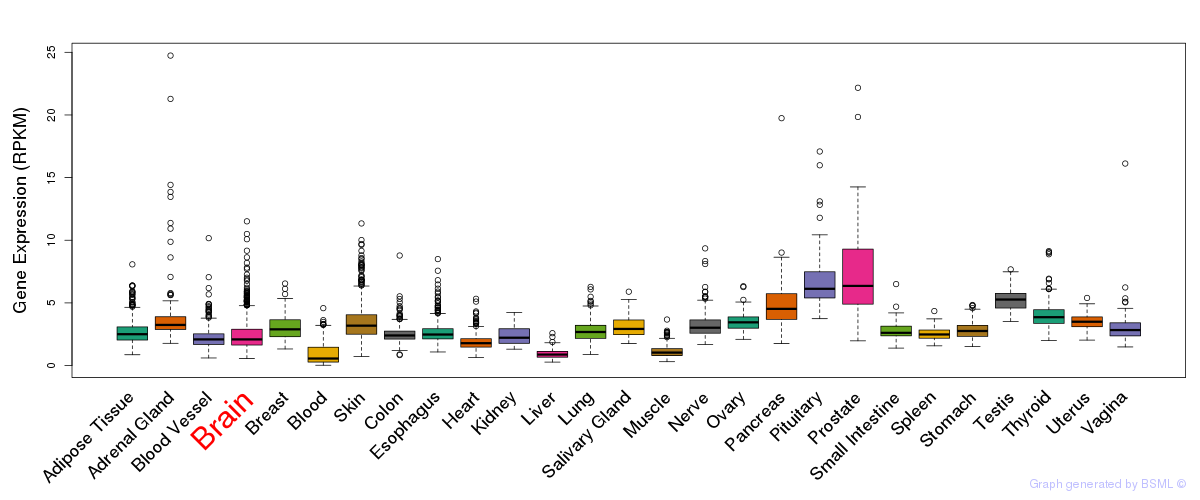
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| SLC35A4 | 0.77 | 0.75 |
| SLC44A2 | 0.75 | 0.75 |
| POMT2 | 0.74 | 0.72 |
| ARAF | 0.74 | 0.74 |
| SLC39A1 | 0.73 | 0.71 |
| ACAD10 | 0.70 | 0.66 |
| RBCK1 | 0.70 | 0.69 |
| PIGO | 0.70 | 0.66 |
| TTC38 | 0.70 | 0.73 |
| CRAT | 0.69 | 0.72 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| SEC62 | -0.50 | -0.53 |
| AF347015.21 | -0.46 | -0.41 |
| FAM159B | -0.41 | -0.44 |
| AC098691.2 | -0.38 | -0.32 |
| MT-CO2 | -0.38 | -0.32 |
| AF347015.18 | -0.37 | -0.34 |
| AF347015.8 | -0.36 | -0.31 |
| MT-ATP8 | -0.35 | -0.35 |
| IL32 | -0.35 | -0.31 |
| FAM133A | -0.35 | -0.35 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| REACTOME GENERIC TRANSCRIPTION PATHWAY | 352 | 181 | All SZGR 2.0 genes in this pathway |
| DOANE RESPONSE TO ANDROGEN UP | 184 | 125 | All SZGR 2.0 genes in this pathway |
| LIAO METASTASIS | 539 | 324 | All SZGR 2.0 genes in this pathway |
| CREIGHTON ENDOCRINE THERAPY RESISTANCE 2 | 473 | 224 | All SZGR 2.0 genes in this pathway |
| CREIGHTON ENDOCRINE THERAPY RESISTANCE 5 | 482 | 296 | All SZGR 2.0 genes in this pathway |
| CHEN HOXA5 TARGETS 9HR UP | 223 | 132 | All SZGR 2.0 genes in this pathway |
| PHONG TNF RESPONSE NOT VIA P38 | 337 | 236 | All SZGR 2.0 genes in this pathway |
| ZWANG TRANSIENTLY UP BY 1ST EGF PULSE ONLY | 1839 | 928 | All SZGR 2.0 genes in this pathway |