Gene Page: STX2
Summary ?
| GeneID | 2054 |
| Symbol | STX2 |
| Synonyms | EPIM|EPM|STX2A|STX2B|STX2C |
| Description | syntaxin 2 |
| Reference | MIM:132350|HGNC:HGNC:3403|Ensembl:ENSG00000111450|HPRD:11816|Vega:OTTHUMG00000168365 |
| Gene type | protein-coding |
| Map location | 12q24.33 |
| Pascal p-value | 0.401 |
| Sherlock p-value | 0.677 |
| Fetal beta | 1.846 |
| DMG | 1 (# studies) |
| Support | EXOCYTOSIS |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:GWASdb | Genome-wide Association Studies | GWASdb records for schizophrenia | |
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| DMG:Wockner_2014 | Genome-wide DNA methylation analysis | This dataset includes 4641 differentially methylated probes corresponding to 2929 unique genes between schizophrenia patients (n=24) and controls (n=24). | 1 |
Section I. Genetics and epigenetics annotation
 Differentially methylated gene
Differentially methylated gene
| Probe | Chromosome | Position | Nearest gene | P (dis) | Beta (dis) | FDR (dis) | Study |
|---|---|---|---|---|---|---|---|
| cg03785806 | 12 | 131304373 | STX2 | 5.07E-5 | 0.471 | 0.022 | DMG:Wockner_2014 |
Section II. Transcriptome annotation
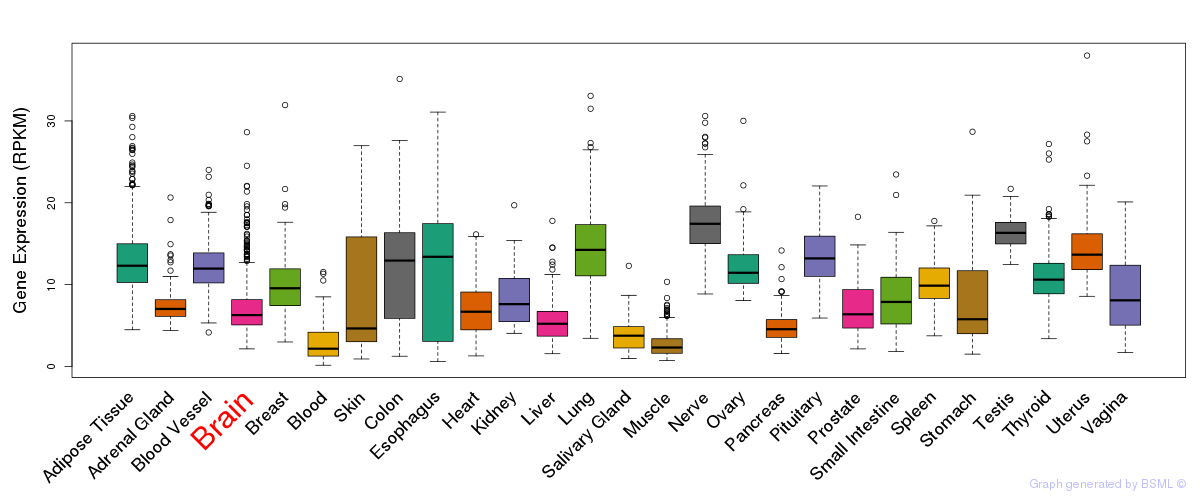
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
No co-expressed genes in brain regions
Section IV. Protein-protein interaction annotation
| Interactors | Aliases B | Official full name B | Experimental | Source | PubMed ID |
|---|---|---|---|---|---|
| SNAP23 | HsT17016 | SNAP23A | SNAP23B | synaptosomal-associated protein, 23kDa | Affinity Capture-Western Reconstituted Complex Two-hybrid | BioGRID | 8663154 |9168999 |12651853 |12828989 |
| SNAP25 | FLJ23079 | RIC-4 | RIC4 | SEC9 | SNAP | SNAP-25 | bA416N4.2 | dJ1068F16.2 | synaptosomal-associated protein, 25kDa | - | HPRD,BioGRID | 7768895 |
| STXBP1 | EIEE4 | MUNC18-1 | UNC18 | hUNC18 | rbSec1 | syntaxin binding protein 1 | Affinity Capture-Western Reconstituted Complex | BioGRID | 7768895 |12773094 |
| STXBP1 | EIEE4 | MUNC18-1 | UNC18 | hUNC18 | rbSec1 | syntaxin binding protein 1 | - | HPRD | 9045631 |
| STXBP2 | Hunc18b | MUNC18-2 | UNC18-2 | UNC18B | pp10122 | syntaxin binding protein 2 | Reconstituted Complex | BioGRID | 7768895 |
| STXBP2 | Hunc18b | MUNC18-2 | UNC18-2 | UNC18B | pp10122 | syntaxin binding protein 2 | - | HPRD | 12198139 |
| STXBP3 | MUNC18-3 | MUNC18C | PSP | UNC-18C | syntaxin binding protein 3 | Affinity Capture-Western | BioGRID | 12773094 |
| SYT1 | DKFZp781D2042 | P65 | SVP65 | SYT | synaptotagmin I | Reconstituted Complex | BioGRID | 10397765 |
| SYT4 | HsT1192 | KIAA1342 | synaptotagmin IV | - | HPRD | 10397765 |
| SYT7 | IPCA-7 | MGC150517 | PCANAP7 | SYT-VII | synaptotagmin VII | - | HPRD | 7791877 |
| VAMP2 | FLJ11460 | SYB2 | VAMP-2 | vesicle-associated membrane protein 2 (synaptobrevin 2) | Two-hybrid | BioGRID | 12828989 |
| VAMP3 | CEB | vesicle-associated membrane protein 3 (cellubrevin) | Two-hybrid | BioGRID | 12828989 |
| VAMP8 | EDB | vesicle-associated membrane protein 8 (endobrevin) | Affinity Capture-Western | BioGRID | 12828989 |
| VAPB | ALS8 | VAMP-B | VAMP-C | VAP-B | VAP-C | VAMP (vesicle-associated membrane protein)-associated protein B and C | Reconstituted Complex | BioGRID | 12651853 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| KEGG SNARE INTERACTIONS IN VESICULAR TRANSPORT | 38 | 25 | All SZGR 2.0 genes in this pathway |
| REACTOME BOTULINUM NEUROTOXICITY | 19 | 14 | All SZGR 2.0 genes in this pathway |
| REACTOME PROTEOLYTIC CLEAVAGE OF SNARE COMPLEX PROTEINS | 17 | 12 | All SZGR 2.0 genes in this pathway |
| REACTOME NEURONAL SYSTEM | 279 | 221 | All SZGR 2.0 genes in this pathway |
| CHARAFE BREAST CANCER LUMINAL VS MESENCHYMAL DN | 460 | 312 | All SZGR 2.0 genes in this pathway |
| PUJANA BRCA1 PCC NETWORK | 1652 | 1023 | All SZGR 2.0 genes in this pathway |
| PUJANA ATM PCC NETWORK | 1442 | 892 | All SZGR 2.0 genes in this pathway |
| IVANOVA HEMATOPOIESIS MATURE CELL | 293 | 160 | All SZGR 2.0 genes in this pathway |
| ULE SPLICING VIA NOVA2 | 43 | 38 | All SZGR 2.0 genes in this pathway |
| DOUGLAS BMI1 TARGETS UP | 566 | 371 | All SZGR 2.0 genes in this pathway |
| MATZUK SPERMATOCYTE | 72 | 55 | All SZGR 2.0 genes in this pathway |
| GRADE COLON VS RECTAL CANCER UP | 38 | 21 | All SZGR 2.0 genes in this pathway |
| BOYLAN MULTIPLE MYELOMA D CLUSTER DN | 40 | 26 | All SZGR 2.0 genes in this pathway |
| BOYLAN MULTIPLE MYELOMA C UP | 47 | 29 | All SZGR 2.0 genes in this pathway |
| HAN SATB1 TARGETS UP | 395 | 249 | All SZGR 2.0 genes in this pathway |
| PECE MAMMARY STEM CELL DN | 146 | 88 | All SZGR 2.0 genes in this pathway |
| ZWANG TRANSIENTLY UP BY 1ST EGF PULSE ONLY | 1839 | 928 | All SZGR 2.0 genes in this pathway |