Gene Page: HSPA6
Summary ?
| GeneID | 3310 |
| Symbol | HSPA6 |
| Synonyms | HSP70B' |
| Description | heat shock protein family A (Hsp70) member 6 |
| Reference | MIM:140555|HGNC:HGNC:5239|Ensembl:ENSG00000173110|HPRD:00775|Vega:OTTHUMG00000034461 |
| Gene type | protein-coding |
| Map location | 1q23 |
| Fetal beta | -0.853 |
| DMG | 1 (# studies) |
| eGene | Myers' cis & trans |
| Support | CompositeSet |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| DMG:Wockner_2014 | Genome-wide DNA methylation analysis | This dataset includes 4641 differentially methylated probes corresponding to 2929 unique genes between schizophrenia patients (n=24) and controls (n=24). | 1 |
| GSMA_IIA | Genome scan meta-analysis (All samples) | Psr: 0.00814 |
Section I. Genetics and epigenetics annotation
 Differentially methylated gene
Differentially methylated gene
| Probe | Chromosome | Position | Nearest gene | P (dis) | Beta (dis) | FDR (dis) | Study |
|---|---|---|---|---|---|---|---|
| cg18761894 | 1 | 161495979 | HSPA6 | 2.59E-5 | 0.307 | 0.017 | DMG:Wockner_2014 |
 eQTL annotation
eQTL annotation
| SNP ID | Chromosome | Position | eGene | Gene Entrez ID | pvalue | qvalue | TSS distance | eQTL type |
|---|---|---|---|---|---|---|---|---|
| rs7547399 | chr1 | 162134746 | HSPA6 | 3310 | 0.01 | cis | ||
| rs16858695 | chr1 | 162193737 | HSPA6 | 3310 | 0.02 | cis | ||
| rs12029998 | 0 | HSPA6 | 3310 | 2.892E-6 | trans | |||
| rs10733033 | chr1 | 22310261 | HSPA6 | 3310 | 2.117E-4 | trans | ||
| rs7543090 | chr1 | 32231154 | HSPA6 | 3310 | 0.02 | trans | ||
| rs10493635 | chr1 | 80417215 | HSPA6 | 3310 | 0.15 | trans | ||
| rs17129266 | chr1 | 86969109 | HSPA6 | 3310 | 0.03 | trans | ||
| rs6593651 | chr1 | 95772660 | HSPA6 | 3310 | 0.04 | trans | ||
| rs17120268 | chr1 | 99829635 | HSPA6 | 3310 | 0.1 | trans | ||
| rs12072589 | chr1 | 102074474 | HSPA6 | 3310 | 0.16 | trans | ||
| rs11164251 | chr1 | 102079419 | HSPA6 | 3310 | 0.17 | trans | ||
| rs6683395 | chr1 | 108385120 | HSPA6 | 3310 | 4.31E-4 | trans | ||
| rs10802055 | 0 | HSPA6 | 3310 | 0.02 | trans | |||
| rs12047301 | chr1 | 209094528 | HSPA6 | 3310 | 0.09 | trans | ||
| rs12565778 | chr1 | 213724810 | HSPA6 | 3310 | 0.11 | trans | ||
| rs16838848 | chr1 | 239900045 | HSPA6 | 3310 | 0.11 | trans | ||
| rs10178561 | chr2 | 20484388 | HSPA6 | 3310 | 0.11 | trans | ||
| rs10195153 | chr2 | 20551992 | HSPA6 | 3310 | 0 | trans | ||
| rs16987610 | chr2 | 20605248 | HSPA6 | 3310 | 1.152E-4 | trans | ||
| rs17044609 | chr2 | 23231427 | HSPA6 | 3310 | 0.16 | trans | ||
| rs6707600 | chr2 | 24521765 | HSPA6 | 3310 | 1.036E-7 | trans | ||
| rs2702079 | chr2 | 24587570 | HSPA6 | 3310 | 0.19 | trans | ||
| rs1406236 | chr2 | 29499441 | HSPA6 | 3310 | 0.04 | trans | ||
| rs1406237 | chr2 | 29499624 | HSPA6 | 3310 | 0.04 | trans | ||
| rs4666189 | chr2 | 29502689 | HSPA6 | 3310 | 0.04 | trans | ||
| rs10170219 | chr2 | 74803516 | HSPA6 | 3310 | 0.08 | trans | ||
| rs10196599 | chr2 | 76957490 | HSPA6 | 3310 | 0.13 | trans | ||
| rs222826 | chr2 | 146878531 | HSPA6 | 3310 | 0.03 | trans | ||
| snp_a-1854914 | 0 | HSPA6 | 3310 | 0.01 | trans | |||
| rs16840841 | chr2 | 157641525 | HSPA6 | 3310 | 0.01 | trans | ||
| rs7560489 | chr2 | 158145388 | HSPA6 | 3310 | 0.01 | trans | ||
| rs7593761 | chr2 | 197275602 | HSPA6 | 3310 | 2.051E-8 | trans | ||
| rs1882580 | chr2 | 227518447 | HSPA6 | 3310 | 0.01 | trans | ||
| rs10510602 | chr3 | 28023926 | HSPA6 | 3310 | 0.16 | trans | ||
| rs12107591 | chr3 | 36751197 | HSPA6 | 3310 | 0.1 | trans | ||
| rs9842768 | chr3 | 36751493 | HSPA6 | 3310 | 0.04 | trans | ||
| rs9842956 | chr3 | 36751621 | HSPA6 | 3310 | 0.06 | trans | ||
| rs2844434 | chr3 | 38390883 | HSPA6 | 3310 | 0.01 | trans | ||
| rs17055195 | chr3 | 55369565 | HSPA6 | 3310 | 0 | trans | ||
| rs17055216 | chr3 | 55381947 | HSPA6 | 3310 | 0 | trans | ||
| rs17055218 | chr3 | 55382206 | HSPA6 | 3310 | 0 | trans | ||
| rs17055224 | chr3 | 55383487 | HSPA6 | 3310 | 0 | trans | ||
| rs17055232 | chr3 | 55385671 | HSPA6 | 3310 | 0 | trans | ||
| rs17055233 | chr3 | 55385691 | HSPA6 | 3310 | 0 | trans | ||
| rs6445730 | chr3 | 55386217 | HSPA6 | 3310 | 0 | trans | ||
| rs747266 | chr3 | 55387379 | HSPA6 | 3310 | 0 | trans | ||
| snp_a-2042818 | 0 | HSPA6 | 3310 | 0.01 | trans | |||
| rs2365463 | chr3 | 66686263 | HSPA6 | 3310 | 0.02 | trans | ||
| rs9883532 | chr3 | 73489258 | HSPA6 | 3310 | 0.02 | trans | ||
| rs1840211 | chr3 | 95854993 | HSPA6 | 3310 | 0.14 | trans | ||
| rs2614197 | chr3 | 113311172 | HSPA6 | 3310 | 0.03 | trans | ||
| rs16847759 | chr3 | 168869746 | HSPA6 | 3310 | 3.186E-4 | trans | ||
| rs2193745 | chr3 | 173964139 | HSPA6 | 3310 | 0.11 | trans | ||
| rs1805608 | chr3 | 180771972 | HSPA6 | 3310 | 3.549E-5 | trans | ||
| rs7630486 | chr3 | 189801990 | HSPA6 | 3310 | 0 | trans | ||
| rs4692018 | chr4 | 26980718 | HSPA6 | 3310 | 0.04 | trans | ||
| rs17086547 | chr4 | 57119142 | HSPA6 | 3310 | 0.06 | trans | ||
| rs10000289 | chr4 | 96437605 | HSPA6 | 3310 | 0.14 | trans | ||
| rs7694882 | chr4 | 112309223 | HSPA6 | 3310 | 0.11 | trans | ||
| rs12641054 | chr4 | 113071694 | HSPA6 | 3310 | 3.141E-17 | trans | ||
| rs9995457 | chr4 | 128299300 | HSPA6 | 3310 | 0.17 | trans | ||
| rs17056206 | chr4 | 171733180 | HSPA6 | 3310 | 0.09 | trans | ||
| rs4398573 | chr4 | 188733644 | HSPA6 | 3310 | 0 | trans | ||
| rs883525 | chr5 | 6495578 | HSPA6 | 3310 | 0.06 | trans | ||
| rs11948869 | chr5 | 6503004 | HSPA6 | 3310 | 0.06 | trans | ||
| rs1844691 | chr5 | 23869998 | HSPA6 | 3310 | 0.16 | trans | ||
| rs6871776 | chr5 | 24036641 | HSPA6 | 3310 | 0.03 | trans | ||
| rs7716468 | chr5 | 24087602 | HSPA6 | 3310 | 0.16 | trans | ||
| rs10065849 | chr5 | 30592364 | HSPA6 | 3310 | 0.04 | trans | ||
| rs1697073 | chr5 | 30907267 | HSPA6 | 3310 | 0.03 | trans | ||
| rs13328131 | chr5 | 35698425 | HSPA6 | 3310 | 0.02 | trans | ||
| rs7717250 | chr5 | 59486425 | HSPA6 | 3310 | 0.06 | trans | ||
| rs16893719 | chr5 | 64647810 | HSPA6 | 3310 | 0.14 | trans | ||
| rs7379895 | 0 | HSPA6 | 3310 | 0.18 | trans | |||
| rs2289344 | chr5 | 91628223 | HSPA6 | 3310 | 0.01 | trans | ||
| rs11956817 | chr5 | 98842089 | HSPA6 | 3310 | 1.381E-5 | trans | ||
| rs1422148 | chr5 | 107526837 | HSPA6 | 3310 | 0 | trans | ||
| rs4463170 | chr5 | 107627192 | HSPA6 | 3310 | 0 | trans | ||
| rs12659121 | chr5 | 107660002 | HSPA6 | 3310 | 0 | trans | ||
| rs17161316 | chr5 | 107756153 | HSPA6 | 3310 | 0 | trans | ||
| rs17147528 | chr5 | 120605704 | HSPA6 | 3310 | 0.03 | trans | ||
| rs7718064 | chr5 | 160556767 | HSPA6 | 3310 | 0.01 | trans | ||
| rs9378593 | chr6 | 1100505 | HSPA6 | 3310 | 0.14 | trans | ||
| rs200200 | chr6 | 5443639 | HSPA6 | 3310 | 0.01 | trans | ||
| rs12663430 | chr6 | 8060533 | HSPA6 | 3310 | 0.11 | trans | ||
| rs9392965 | chr6 | 8141268 | HSPA6 | 3310 | 0.05 | trans | ||
| rs502759 | chr6 | 18468463 | HSPA6 | 3310 | 0.16 | trans | ||
| rs17583852 | chr6 | 32912587 | HSPA6 | 3310 | 0.03 | trans | ||
| rs9381009 | chr6 | 40918100 | HSPA6 | 3310 | 0.13 | trans | ||
| rs3125579 | chr6 | 98908957 | HSPA6 | 3310 | 0.01 | trans | ||
| rs9321410 | chr6 | 133998132 | HSPA6 | 3310 | 0.01 | trans | ||
| rs9402819 | chr6 | 136848169 | HSPA6 | 3310 | 0.06 | trans | ||
| rs9389400 | chr6 | 136854803 | HSPA6 | 3310 | 0.06 | trans | ||
| rs6918404 | chr6 | 139297610 | HSPA6 | 3310 | 0.03 | trans | ||
| rs197485 | chr6 | 143068177 | HSPA6 | 3310 | 1.389E-6 | trans | ||
| rs1424935 | chr6 | 143699145 | HSPA6 | 3310 | 0.11 | trans | ||
| rs3807504 | chr7 | 16684098 | HSPA6 | 3310 | 0.06 | trans | ||
| rs17150652 | chr7 | 25012054 | HSPA6 | 3310 | 0.1 | trans | ||
| rs343083 | chr7 | 35571887 | HSPA6 | 3310 | 0.17 | trans | ||
| rs7808547 | chr7 | 83753166 | HSPA6 | 3310 | 0 | trans | ||
| rs3779572 | chr7 | 90608515 | HSPA6 | 3310 | 7.699E-4 | trans | ||
| rs1208907 | chr7 | 90654714 | HSPA6 | 3310 | 0.07 | trans | ||
| rs3779585 | chr7 | 90719297 | HSPA6 | 3310 | 1.604E-4 | trans | ||
| snp_a-2030753 | 0 | HSPA6 | 3310 | 5.549E-4 | trans | |||
| rs879428 | chr7 | 90737968 | HSPA6 | 3310 | 1.604E-4 | trans | ||
| rs2113546 | chr7 | 134953754 | HSPA6 | 3310 | 0.03 | trans | ||
| rs17133801 | chr7 | 146787365 | HSPA6 | 3310 | 6.712E-8 | trans | ||
| rs718119 | chr8 | 3159708 | HSPA6 | 3310 | 0.02 | trans | ||
| rs3896041 | chr8 | 3160879 | HSPA6 | 3310 | 0.18 | trans | ||
| rs17066062 | chr8 | 3272540 | HSPA6 | 3310 | 0.11 | trans | ||
| rs11998339 | chr8 | 8726545 | HSPA6 | 3310 | 0.12 | trans | ||
| rs17695317 | chr8 | 18489042 | HSPA6 | 3310 | 0.19 | trans | ||
| rs16885771 | chr8 | 36898251 | HSPA6 | 3310 | 0.17 | trans | ||
| rs16918960 | chr8 | 54175049 | HSPA6 | 3310 | 0.18 | trans | ||
| rs6992596 | chr8 | 54453189 | HSPA6 | 3310 | 0.02 | trans | ||
| rs16920160 | chr8 | 55308290 | HSPA6 | 3310 | 3.758E-4 | trans | ||
| rs1431586 | chr8 | 64390728 | HSPA6 | 3310 | 0 | trans | ||
| rs4263735 | chr8 | 77931285 | HSPA6 | 3310 | 0.01 | trans | ||
| rs1483373 | chr8 | 90186442 | HSPA6 | 3310 | 0.03 | trans | ||
| rs16876804 | chr8 | 108825477 | HSPA6 | 3310 | 0.05 | trans | ||
| rs10491582 | chr9 | 953598 | HSPA6 | 3310 | 0.03 | trans | ||
| rs2121246 | chr9 | 12037953 | HSPA6 | 3310 | 1.603E-6 | trans | ||
| rs11999552 | chr9 | 30351246 | HSPA6 | 3310 | 0.03 | trans | ||
| rs7032605 | chr9 | 30426130 | HSPA6 | 3310 | 0.1 | trans | ||
| rs10969802 | chr9 | 30636427 | HSPA6 | 3310 | 0.02 | trans | ||
| rs10969816 | chr9 | 30661611 | HSPA6 | 3310 | 0.02 | trans | ||
| rs12350524 | chr9 | 30743421 | HSPA6 | 3310 | 0.02 | trans | ||
| rs12343068 | chr9 | 30783524 | HSPA6 | 3310 | 0.02 | trans | ||
| rs10511868 | chr9 | 30816504 | HSPA6 | 3310 | 0.02 | trans | ||
| rs3758354 | chr9 | 75764564 | HSPA6 | 3310 | 0.16 | trans | ||
| rs914715 | chr9 | 82310897 | HSPA6 | 3310 | 0.13 | trans | ||
| rs7026368 | chr9 | 82675015 | HSPA6 | 3310 | 0.04 | trans | ||
| rs11140349 | chr9 | 86675586 | HSPA6 | 3310 | 0.01 | trans | ||
| rs10868543 | chr9 | 89852686 | HSPA6 | 3310 | 1.558E-18 | trans | ||
| rs17053159 | chr9 | 90198374 | HSPA6 | 3310 | 0 | trans | ||
| rs3802458 | chr9 | 97741273 | HSPA6 | 3310 | 0.18 | trans | ||
| rs3802457 | chr9 | 97741335 | HSPA6 | 3310 | 1.391E-5 | trans | ||
| rs10993509 | chr9 | 98071989 | HSPA6 | 3310 | 0.09 | trans | ||
| rs10816779 | chr9 | 111946102 | HSPA6 | 3310 | 0.04 | trans | ||
| rs2210142 | chr9 | 119716771 | HSPA6 | 3310 | 0 | trans | ||
| rs7868621 | chr9 | 132130970 | HSPA6 | 3310 | 0.06 | trans | ||
| rs10776917 | chr9 | 137858478 | HSPA6 | 3310 | 9.068E-11 | trans | ||
| rs1335812 | chr10 | 24119275 | HSPA6 | 3310 | 0.02 | trans | ||
| rs602658 | chr10 | 36007778 | HSPA6 | 3310 | 0.04 | trans | ||
| rs1598793 | chr10 | 79127993 | HSPA6 | 3310 | 0.05 | trans | ||
| rs7090588 | chr10 | 85717555 | HSPA6 | 3310 | 0 | trans | ||
| rs12247436 | chr10 | 85770974 | HSPA6 | 3310 | 0.18 | trans | ||
| rs1902680 | chr10 | 88049777 | HSPA6 | 3310 | 0 | trans | ||
| rs17109578 | chr10 | 90351347 | HSPA6 | 3310 | 0 | trans | ||
| rs12218424 | chr10 | 92958797 | HSPA6 | 3310 | 0 | trans | ||
| rs11195998 | chr10 | 114335289 | HSPA6 | 3310 | 0.01 | trans | ||
| rs11196000 | chr10 | 114337116 | HSPA6 | 3310 | 0.01 | trans | ||
| rs3124736 | chr10 | 115497111 | HSPA6 | 3310 | 0.19 | trans | ||
| rs10787671 | chr10 | 118189794 | HSPA6 | 3310 | 0.13 | trans | ||
| rs1431486 | chr10 | 118193131 | HSPA6 | 3310 | 0.02 | trans | ||
| rs17104648 | chr10 | 124869902 | HSPA6 | 3310 | 1.451E-4 | trans | ||
| rs6578285 | chr11 | 1380763 | HSPA6 | 3310 | 0 | trans | ||
| rs11824618 | chr11 | 4188729 | HSPA6 | 3310 | 0.01 | trans | ||
| rs7948587 | chr11 | 7784541 | HSPA6 | 3310 | 0.07 | trans | ||
| rs12286689 | chr11 | 7853932 | HSPA6 | 3310 | 0.07 | trans | ||
| rs11022545 | chr11 | 12933682 | HSPA6 | 3310 | 0.1 | trans | ||
| rs16932222 | chr11 | 15875436 | HSPA6 | 3310 | 1.13E-4 | trans | ||
| rs11823900 | chr11 | 15887003 | HSPA6 | 3310 | 6.214E-10 | trans | ||
| rs11603947 | chr11 | 33486575 | HSPA6 | 3310 | 0.15 | trans | ||
| rs7928546 | chr11 | 36102893 | HSPA6 | 3310 | 0.18 | trans | ||
| rs17138435 | chr11 | 79192986 | HSPA6 | 3310 | 0.07 | trans | ||
| rs1499026 | chr11 | 88433481 | HSPA6 | 3310 | 0.08 | trans | ||
| rs10501694 | chr11 | 88560150 | HSPA6 | 3310 | 0.01 | trans | ||
| rs11018448 | chr11 | 88793318 | HSPA6 | 3310 | 0.13 | trans | ||
| rs8181526 | chr11 | 88797520 | HSPA6 | 3310 | 0.04 | trans | ||
| rs2164522 | chr11 | 89190488 | HSPA6 | 3310 | 0.06 | trans | ||
| rs11018625 | chr11 | 89202568 | HSPA6 | 3310 | 0.01 | trans | ||
| rs11018629 | chr11 | 89206963 | HSPA6 | 3310 | 0.07 | trans | ||
| rs614128 | chr11 | 89214113 | HSPA6 | 3310 | 0.03 | trans | ||
| snp_a-2212958 | 0 | HSPA6 | 3310 | 0.17 | trans | |||
| rs17133063 | chr11 | 98991111 | HSPA6 | 3310 | 0.08 | trans | ||
| rs17119029 | chr11 | 115857373 | HSPA6 | 3310 | 0.01 | trans | ||
| rs1632799 | chr11 | 115918300 | HSPA6 | 3310 | 0 | trans | ||
| rs4759436 | chr12 | 3083708 | HSPA6 | 3310 | 0.08 | trans | ||
| rs6489409 | chr12 | 3083951 | HSPA6 | 3310 | 0.04 | trans | ||
| rs7953537 | chr12 | 44693287 | HSPA6 | 3310 | 0.06 | trans | ||
| rs17094560 | chr12 | 44769767 | HSPA6 | 3310 | 0.06 | trans | ||
| rs1344796 | chr12 | 76397104 | HSPA6 | 3310 | 8.041E-4 | trans | ||
| rs17046341 | chr12 | 79472791 | HSPA6 | 3310 | 0 | trans | ||
| rs11114260 | chr12 | 80241492 | HSPA6 | 3310 | 0.01 | trans | ||
| rs12304649 | chr12 | 115144452 | HSPA6 | 3310 | 7.987E-4 | trans | ||
| rs12320582 | chr12 | 115154435 | HSPA6 | 3310 | 0.03 | trans | ||
| rs17077917 | chr13 | 33412452 | HSPA6 | 3310 | 0.01 | trans | ||
| rs10492401 | chr13 | 33489671 | HSPA6 | 3310 | 8.988E-4 | trans | ||
| rs9566314 | chr13 | 38767683 | HSPA6 | 3310 | 0 | trans | ||
| rs17070045 | chr13 | 58400458 | HSPA6 | 3310 | 0 | trans | ||
| rs17235614 | chr13 | 97953192 | HSPA6 | 3310 | 0.19 | trans | ||
| rs16960790 | chr13 | 103589970 | HSPA6 | 3310 | 0.12 | trans | ||
| rs12866183 | chr13 | 103605460 | HSPA6 | 3310 | 0.08 | trans | ||
| rs17330244 | chr13 | 103612897 | HSPA6 | 3310 | 0.08 | trans | ||
| rs16966394 | chr13 | 105986280 | HSPA6 | 3310 | 0.07 | trans | ||
| rs7332984 | chr13 | 108328677 | HSPA6 | 3310 | 0 | trans | ||
| rs17109555 | chr14 | 40132840 | HSPA6 | 3310 | 0.01 | trans | ||
| rs17105827 | chr14 | 72488090 | HSPA6 | 3310 | 0.18 | trans | ||
| rs758913 | chr14 | 73264460 | HSPA6 | 3310 | 0.1 | trans | ||
| rs1642278 | chr14 | 87273027 | HSPA6 | 3310 | 0.09 | trans | ||
| rs4503732 | chr15 | 46642262 | HSPA6 | 3310 | 0.05 | trans | ||
| rs16960698 | chr15 | 48523283 | HSPA6 | 3310 | 0 | trans | ||
| rs6494052 | chr15 | 59295924 | HSPA6 | 3310 | 0.03 | trans | ||
| rs16941048 | chr15 | 59416283 | HSPA6 | 3310 | 0.17 | trans | ||
| rs8039102 | chr15 | 66510617 | HSPA6 | 3310 | 0.03 | trans | ||
| rs2278598 | chr15 | 89174747 | HSPA6 | 3310 | 0.05 | trans | ||
| rs10152821 | chr15 | 89180884 | HSPA6 | 3310 | 0.06 | trans | ||
| rs9920555 | chr15 | 98417871 | HSPA6 | 3310 | 0.08 | trans | ||
| rs276614 | chr16 | 13166182 | HSPA6 | 3310 | 0.12 | trans | ||
| rs10492782 | chr16 | 13210707 | HSPA6 | 3310 | 1.071E-14 | trans | ||
| rs7190264 | chr16 | 23685116 | HSPA6 | 3310 | 0 | trans | ||
| rs755295 | chr16 | 27806317 | HSPA6 | 3310 | 0.04 | trans | ||
| rs17821412 | chr16 | 57938754 | HSPA6 | 3310 | 0.03 | trans | ||
| rs4888697 | chr16 | 77963301 | HSPA6 | 3310 | 0.15 | trans | ||
| rs8062753 | chr16 | 78711589 | HSPA6 | 3310 | 0.16 | trans | ||
| rs8082342 | chr17 | 8719237 | HSPA6 | 3310 | 0.18 | trans | ||
| rs16944595 | chr17 | 11242834 | HSPA6 | 3310 | 3.799E-4 | trans | ||
| rs16944600 | chr17 | 11245098 | HSPA6 | 3310 | 0.06 | trans | ||
| rs16944601 | chr17 | 11245219 | HSPA6 | 3310 | 0.11 | trans | ||
| rs4924758 | chr17 | 18367377 | HSPA6 | 3310 | 0.01 | trans | ||
| rs792799 | chr17 | 51176163 | HSPA6 | 3310 | 0.14 | trans | ||
| rs16970485 | chr17 | 75722431 | HSPA6 | 3310 | 0.07 | trans | ||
| rs1811086 | chr17 | 76173021 | HSPA6 | 3310 | 0.1 | trans | ||
| rs9956108 | chr18 | 3042415 | HSPA6 | 3310 | 5.298E-4 | trans | ||
| rs10468647 | chr18 | 3868774 | HSPA6 | 3310 | 5.171E-6 | trans | ||
| rs9807505 | chr18 | 4259102 | HSPA6 | 3310 | 0.06 | trans | ||
| rs3744977 | chr18 | 8826082 | HSPA6 | 3310 | 0.12 | trans | ||
| snp_a-2036900 | 0 | HSPA6 | 3310 | 3.783E-5 | trans | |||
| rs7227140 | chr18 | 10696798 | HSPA6 | 3310 | 0.01 | trans | ||
| rs9950823 | chr18 | 47526879 | HSPA6 | 3310 | 0.06 | trans | ||
| rs16951384 | chr18 | 47527116 | HSPA6 | 3310 | 0 | trans | ||
| rs17734946 | chr18 | 54987388 | HSPA6 | 3310 | 0.17 | trans | ||
| rs1942853 | chr18 | 55195680 | HSPA6 | 3310 | 0.16 | trans | ||
| rs7227933 | chr18 | 57382213 | HSPA6 | 3310 | 0.14 | trans | ||
| rs17082636 | chr18 | 68098567 | HSPA6 | 3310 | 0.04 | trans | ||
| rs3935656 | chr18 | 70947038 | HSPA6 | 3310 | 0.05 | trans | ||
| rs4802946 | 0 | HSPA6 | 3310 | 0 | trans | |||
| rs1673892 | chr19 | 52974222 | HSPA6 | 3310 | 2.314E-4 | trans | ||
| rs979158 | chr19 | 52976380 | HSPA6 | 3310 | 6.406E-4 | trans | ||
| rs4815748 | chr20 | 4822039 | HSPA6 | 3310 | 0.09 | trans | ||
| rs362582 | chr20 | 10254286 | HSPA6 | 3310 | 0.01 | trans | ||
| rs362592 | chr20 | 10272491 | HSPA6 | 3310 | 0.19 | trans | ||
| rs362991 | chr20 | 10276280 | HSPA6 | 3310 | 0.14 | trans | ||
| rs363004 | chr20 | 10279796 | HSPA6 | 3310 | 0.13 | trans | ||
| rs8120542 | chr20 | 24395092 | HSPA6 | 3310 | 0 | trans | ||
| rs8114637 | chr20 | 24395207 | HSPA6 | 3310 | 0 | trans | ||
| rs4536719 | chr20 | 24397114 | HSPA6 | 3310 | 0.12 | trans | ||
| rs9974024 | chr20 | 24397274 | HSPA6 | 3310 | 0 | trans | ||
| rs6515493 | chr20 | 24401635 | HSPA6 | 3310 | 0 | trans | ||
| rs6049703 | chr20 | 24432425 | HSPA6 | 3310 | 3.969E-4 | trans | ||
| rs1158857 | chr20 | 37334090 | HSPA6 | 3310 | 0.07 | trans | ||
| rs914136 | chr21 | 30848367 | HSPA6 | 3310 | 0.16 | trans | ||
| rs2832343 | chr21 | 30849712 | HSPA6 | 3310 | 0.09 | trans | ||
| rs963073 | chr21 | 30867451 | HSPA6 | 3310 | 0.16 | trans | ||
| rs7275620 | chr21 | 30888876 | HSPA6 | 3310 | 0.13 | trans | ||
| rs16994521 | chr21 | 38208009 | HSPA6 | 3310 | 0.15 | trans | ||
| rs1894251 | chr22 | 22440498 | HSPA6 | 3310 | 0.03 | trans | ||
| rs1925925 | chrX | 69625446 | HSPA6 | 3310 | 0.14 | trans | ||
| rs6524459 | chrX | 84704458 | HSPA6 | 3310 | 8.398E-7 | trans | ||
| rs1293493 | chrX | 122175994 | HSPA6 | 3310 | 9.945E-4 | trans | ||
| rs9306818 | chrX | 143403931 | HSPA6 | 3310 | 0.05 | trans |
Section II. Transcriptome annotation
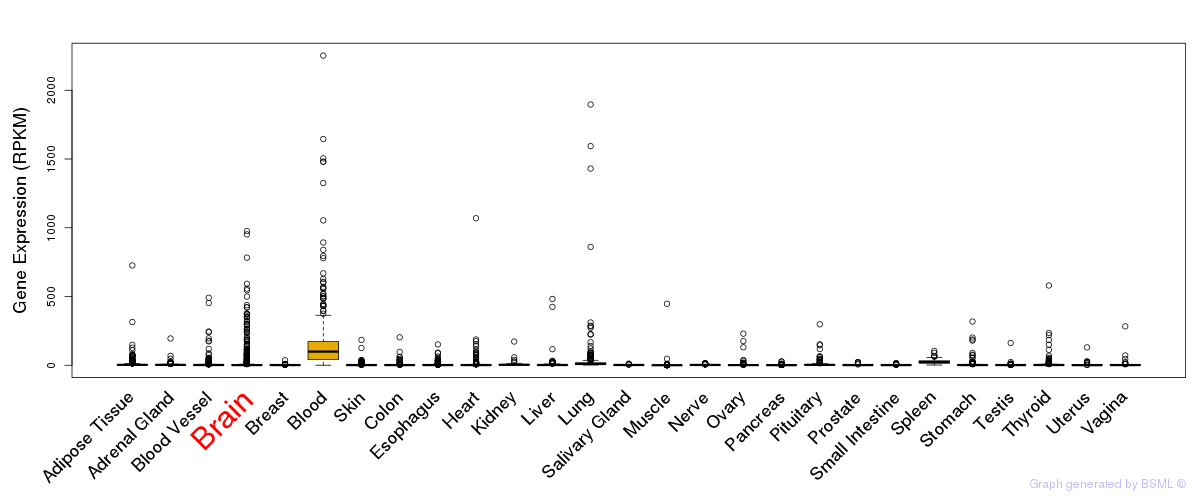
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
No co-expressed genes in brain regions
Section III. Gene Ontology annotation
| Molecular function | GO term | Evidence | Neuro keywords | PubMed ID |
|---|---|---|---|---|
| GO:0000166 | nucleotide binding | IEA | - | |
| GO:0005524 | ATP binding | IEA | - | |
| Biological process | GO term | Evidence | Neuro keywords | PubMed ID |
| GO:0006986 | response to unfolded protein | TAS | 2327978 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| KEGG SPLICEOSOME | 128 | 72 | All SZGR 2.0 genes in this pathway |
| KEGG MAPK SIGNALING PATHWAY | 267 | 205 | All SZGR 2.0 genes in this pathway |
| KEGG ENDOCYTOSIS | 183 | 132 | All SZGR 2.0 genes in this pathway |
| KEGG ANTIGEN PROCESSING AND PRESENTATION | 89 | 65 | All SZGR 2.0 genes in this pathway |
| NAKAMURA TUMOR ZONE PERIPHERAL VS CENTRAL DN | 634 | 384 | All SZGR 2.0 genes in this pathway |
| LU TUMOR VASCULATURE UP | 29 | 17 | All SZGR 2.0 genes in this pathway |
| TURASHVILI BREAST LOBULAR CARCINOMA VS DUCTAL NORMAL UP | 69 | 38 | All SZGR 2.0 genes in this pathway |
| FULCHER INFLAMMATORY RESPONSE LECTIN VS LPS UP | 579 | 346 | All SZGR 2.0 genes in this pathway |
| NOJIMA SFRP2 TARGETS UP | 31 | 23 | All SZGR 2.0 genes in this pathway |
| SMIRNOV CIRCULATING ENDOTHELIOCYTES IN CANCER UP | 158 | 103 | All SZGR 2.0 genes in this pathway |
| OSWALD HEMATOPOIETIC STEM CELL IN COLLAGEN GEL DN | 275 | 168 | All SZGR 2.0 genes in this pathway |
| GARGALOVIC RESPONSE TO OXIDIZED PHOSPHOLIPIDS BLUE UP | 136 | 80 | All SZGR 2.0 genes in this pathway |
| KIM WT1 TARGETS 8HR UP | 164 | 122 | All SZGR 2.0 genes in this pathway |
| KIM WT1 TARGETS 12HR UP | 162 | 116 | All SZGR 2.0 genes in this pathway |
| CONCANNON APOPTOSIS BY EPOXOMICIN UP | 239 | 157 | All SZGR 2.0 genes in this pathway |
| DUNNE TARGETS OF AML1 MTG8 FUSION UP | 52 | 32 | All SZGR 2.0 genes in this pathway |
| KAN RESPONSE TO ARSENIC TRIOXIDE | 123 | 80 | All SZGR 2.0 genes in this pathway |
| HWANG PROSTATE CANCER MARKERS | 28 | 19 | All SZGR 2.0 genes in this pathway |
| PEREZ TP63 TARGETS | 355 | 243 | All SZGR 2.0 genes in this pathway |
| MAHADEVAN RESPONSE TO MP470 DN | 19 | 11 | All SZGR 2.0 genes in this pathway |
| FARMER BREAST CANCER APOCRINE VS BASAL | 330 | 217 | All SZGR 2.0 genes in this pathway |
| DAUER STAT3 TARGETS DN | 50 | 34 | All SZGR 2.0 genes in this pathway |
| BUYTAERT PHOTODYNAMIC THERAPY STRESS UP | 811 | 508 | All SZGR 2.0 genes in this pathway |
| BENPORATH SUZ12 TARGETS | 1038 | 678 | All SZGR 2.0 genes in this pathway |
| BENPORATH EED TARGETS | 1062 | 725 | All SZGR 2.0 genes in this pathway |
| BENPORATH ES WITH H3K27ME3 | 1118 | 744 | All SZGR 2.0 genes in this pathway |
| BENPORATH PRC2 TARGETS | 652 | 441 | All SZGR 2.0 genes in this pathway |
| LEI MYB TARGETS | 318 | 215 | All SZGR 2.0 genes in this pathway |
| MOREAUX MULTIPLE MYELOMA BY TACI UP | 412 | 249 | All SZGR 2.0 genes in this pathway |
| BROWNE HCMV INFECTION 18HR UP | 178 | 111 | All SZGR 2.0 genes in this pathway |
| KAYO CALORIE RESTRICTION MUSCLE UP | 95 | 64 | All SZGR 2.0 genes in this pathway |
| BACOLOD RESISTANCE TO ALKYLATING AGENTS UP | 26 | 17 | All SZGR 2.0 genes in this pathway |
| TRACEY RESISTANCE TO IFNA2 DN | 32 | 23 | All SZGR 2.0 genes in this pathway |
| BILD MYC ONCOGENIC SIGNATURE | 206 | 117 | All SZGR 2.0 genes in this pathway |
| HELLER SILENCED BY METHYLATION DN | 105 | 67 | All SZGR 2.0 genes in this pathway |
| HELLER HDAC TARGETS DN | 292 | 189 | All SZGR 2.0 genes in this pathway |
| HELLER HDAC TARGETS SILENCED BY METHYLATION DN | 281 | 179 | All SZGR 2.0 genes in this pathway |
| RIZKI TUMOR INVASIVENESS 3D DN | 270 | 181 | All SZGR 2.0 genes in this pathway |
| SMID BREAST CANCER BASAL UP | 648 | 398 | All SZGR 2.0 genes in this pathway |
| BOQUEST STEM CELL CULTURED VS FRESH UP | 425 | 298 | All SZGR 2.0 genes in this pathway |
| PODAR RESPONSE TO ADAPHOSTIN UP | 147 | 98 | All SZGR 2.0 genes in this pathway |
| LU TUMOR ENDOTHELIAL MARKERS UP | 22 | 14 | All SZGR 2.0 genes in this pathway |
| YAGI AML WITH INV 16 TRANSLOCATION | 422 | 277 | All SZGR 2.0 genes in this pathway |
| NAKAYAMA SOFT TISSUE TUMORS PCA1 UP | 76 | 46 | All SZGR 2.0 genes in this pathway |
| CHANDRAN METASTASIS UP | 221 | 135 | All SZGR 2.0 genes in this pathway |
| WIERENGA STAT5A TARGETS UP | 217 | 131 | All SZGR 2.0 genes in this pathway |
| WIERENGA STAT5A TARGETS GROUP1 | 136 | 76 | All SZGR 2.0 genes in this pathway |
| FORTSCHEGGER PHF8 TARGETS DN | 784 | 464 | All SZGR 2.0 genes in this pathway |
| ZWANG TRANSIENTLY UP BY 2ND EGF PULSE ONLY | 1725 | 838 | All SZGR 2.0 genes in this pathway |