Gene Page: OXT
Summary ?
| GeneID | 5020 |
| Symbol | OXT |
| Synonyms | OT|OT-NPI|OXT-NPI |
| Description | oxytocin/neurophysin I prepropeptide |
| Reference | MIM:167050|HGNC:HGNC:8528|Ensembl:ENSG00000101405|HPRD:08880|Vega:OTTHUMG00000031724 |
| Gene type | protein-coding |
| Map location | 20p13 |
| Pascal p-value | 0.244 |
| Fetal beta | -0.57 |
| eGene | Myers' cis & trans Meta |
| Support | NEUROTROPHIN SIGNALING Potential synaptic genes |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| PMID:cooccur | High-throughput literature-search | Systematic search in PubMed for genes co-occurring with SCZ keywords. A total of 3027 genes were included. | |
| Literature | High-throughput literature-search | Co-occurance with Schizophrenia keywords: schizophrenia,schizophrenic,schizophrenias | Click to show details |
Section I. Genetics and epigenetics annotation
 eQTL annotation
eQTL annotation
| SNP ID | Chromosome | Position | eGene | Gene Entrez ID | pvalue | qvalue | TSS distance | eQTL type |
|---|---|---|---|---|---|---|---|---|
| rs2494875 | chr1 | 50652317 | OXT | 5020 | 0.04 | trans | ||
| rs6688407 | chr1 | 111796110 | OXT | 5020 | 0.04 | trans | ||
| rs17047497 | chr2 | 56347610 | OXT | 5020 | 0.03 | trans | ||
| rs11679895 | chr2 | 104914347 | OXT | 5020 | 0.13 | trans | ||
| rs17203661 | chr4 | 8527262 | OXT | 5020 | 0.18 | trans | ||
| rs13146859 | chr4 | 8528027 | OXT | 5020 | 0.19 | trans | ||
| rs17211228 | chr5 | 81742122 | OXT | 5020 | 0.06 | trans | ||
| rs2546197 | chr5 | 95172280 | OXT | 5020 | 0.15 | trans | ||
| rs9371139 | chr6 | 169955205 | OXT | 5020 | 0.07 | trans | ||
| rs4391437 | chr8 | 9373515 | OXT | 5020 | 0.08 | trans | ||
| rs6601322 | chr8 | 9375852 | OXT | 5020 | 0.02 | trans | ||
| rs17062725 | chr9 | 79168620 | OXT | 5020 | 1.405E-5 | trans | ||
| rs17824714 | chr12 | 65764276 | OXT | 5020 | 0.13 | trans | ||
| rs6581637 | chr12 | 65852923 | OXT | 5020 | 0.13 | trans | ||
| rs12585923 | chr13 | 76625287 | OXT | 5020 | 0.15 | trans | ||
| rs1330442 | chr13 | 76680920 | OXT | 5020 | 0.04 | trans | ||
| rs1499605 | chr15 | 63706224 | OXT | 5020 | 0.08 | trans | ||
| rs289811 | chr15 | 63710672 | OXT | 5020 | 0.02 | trans | ||
| rs16953493 | chr17 | 3714057 | OXT | 5020 | 0.05 | trans | ||
| rs16953498 | chr17 | 3719619 | OXT | 5020 | 0.06 | trans | ||
| rs1183148 | chr17 | 3733846 | OXT | 5020 | 0.04 | trans | ||
| rs7272436 | chr20 | 52558686 | OXT | 5020 | 0.16 | trans |
Section II. Transcriptome annotation
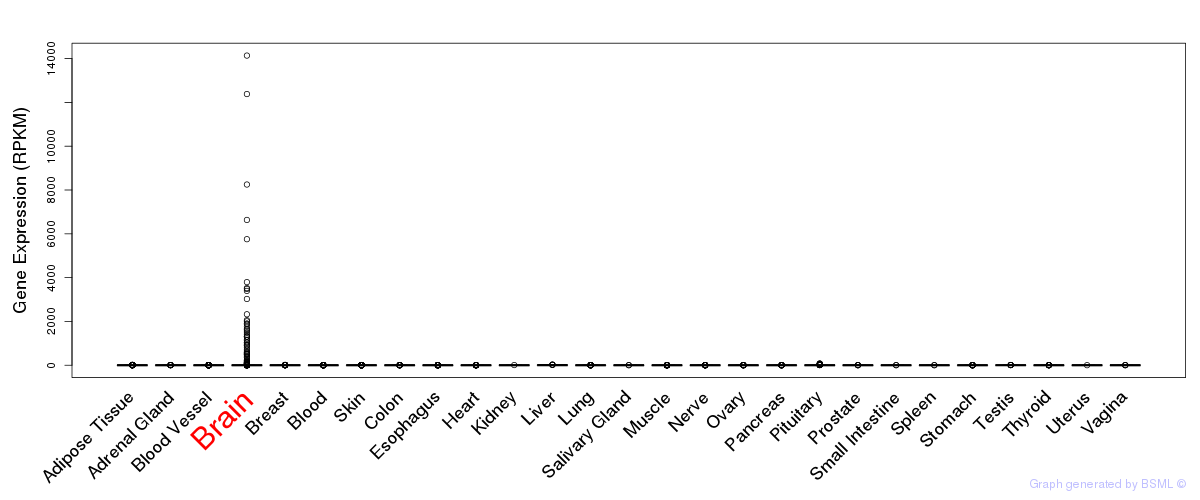
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| RAB9A | 0.68 | 0.72 |
| AKR7A2 | 0.68 | 0.66 |
| FUCA2 | 0.65 | 0.66 |
| MRPL40 | 0.65 | 0.62 |
| TMEM59 | 0.64 | 0.66 |
| FAM96A | 0.64 | 0.54 |
| C2orf28 | 0.64 | 0.62 |
| SRI | 0.64 | 0.64 |
| GRHPR | 0.63 | 0.63 |
| SDHC | 0.63 | 0.62 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| JAG2 | -0.41 | -0.40 |
| BAT2D1 | -0.40 | -0.43 |
| CACNA1C | -0.40 | -0.35 |
| HIVEP3 | -0.39 | -0.33 |
| CACNA1B | -0.39 | -0.34 |
| ADCY1 | -0.39 | -0.29 |
| XKR4 | -0.37 | -0.30 |
| MYT1L | -0.37 | -0.31 |
| AL391628.1 | -0.36 | -0.38 |
| TNRC6C | -0.36 | -0.34 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| REACTOME GASTRIN CREB SIGNALLING PATHWAY VIA PKC AND MAPK | 205 | 136 | All SZGR 2.0 genes in this pathway |
| REACTOME SIGNALING BY GPCR | 920 | 449 | All SZGR 2.0 genes in this pathway |
| REACTOME PEPTIDE LIGAND BINDING RECEPTORS | 188 | 108 | All SZGR 2.0 genes in this pathway |
| REACTOME CLASS A1 RHODOPSIN LIKE RECEPTORS | 305 | 177 | All SZGR 2.0 genes in this pathway |
| REACTOME G ALPHA Q SIGNALLING EVENTS | 184 | 116 | All SZGR 2.0 genes in this pathway |
| REACTOME GPCR DOWNSTREAM SIGNALING | 805 | 368 | All SZGR 2.0 genes in this pathway |
| REACTOME GPCR LIGAND BINDING | 408 | 246 | All SZGR 2.0 genes in this pathway |
| HOEBEKE LYMPHOID STEM CELL DN | 86 | 59 | All SZGR 2.0 genes in this pathway |
| LEE AGING NEOCORTEX UP | 89 | 59 | All SZGR 2.0 genes in this pathway |
| MATZUK POST-IMPLANTATION AND POST-PARTUM | 14 | 13 | All SZGR 2.0 genes in this pathway |
| YOSHIMURA MAPK8 TARGETS UP | 1305 | 895 | All SZGR 2.0 genes in this pathway |
| MEISSNER NPC HCP WITH H3 UNMETHYLATED | 536 | 296 | All SZGR 2.0 genes in this pathway |
| YAMASHITA LIVER CANCER WITH EPCAM UP | 53 | 25 | All SZGR 2.0 genes in this pathway |
| MEISSNER BRAIN HCP WITH H3K4ME2 AND H3K27ME3 | 59 | 35 | All SZGR 2.0 genes in this pathway |
| MIKKELSEN MEF HCP WITH H3K27ME3 | 590 | 403 | All SZGR 2.0 genes in this pathway |