Gene Page: SAMD4B
Summary ?
| GeneID | 55095 |
| Symbol | SAMD4B |
| Synonyms | SMGB|Smaug2 |
| Description | sterile alpha motif domain containing 4B |
| Reference | HGNC:HGNC:25492|Ensembl:ENSG00000179134|HPRD:10960|Vega:OTTHUMG00000182967 |
| Gene type | protein-coding |
| Map location | 19q13.2 |
| Pascal p-value | 0.15 |
| Sherlock p-value | 0.022 |
| eGene | Myers' cis & trans |
| Support | CompositeSet Darnell FMRP targets |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS |
Section I. Genetics and epigenetics annotation
 eQTL annotation
eQTL annotation
| SNP ID | Chromosome | Position | eGene | Gene Entrez ID | pvalue | qvalue | TSS distance | eQTL type |
|---|---|---|---|---|---|---|---|---|
| rs17839618 | chr5 | 128477438 | SAMD4B | 55095 | 0.12 | trans | ||
| rs8089917 | chr18 | 73363948 | SAMD4B | 55095 | 0.03 | trans |
Section II. Transcriptome annotation
General gene expression (GTEx)
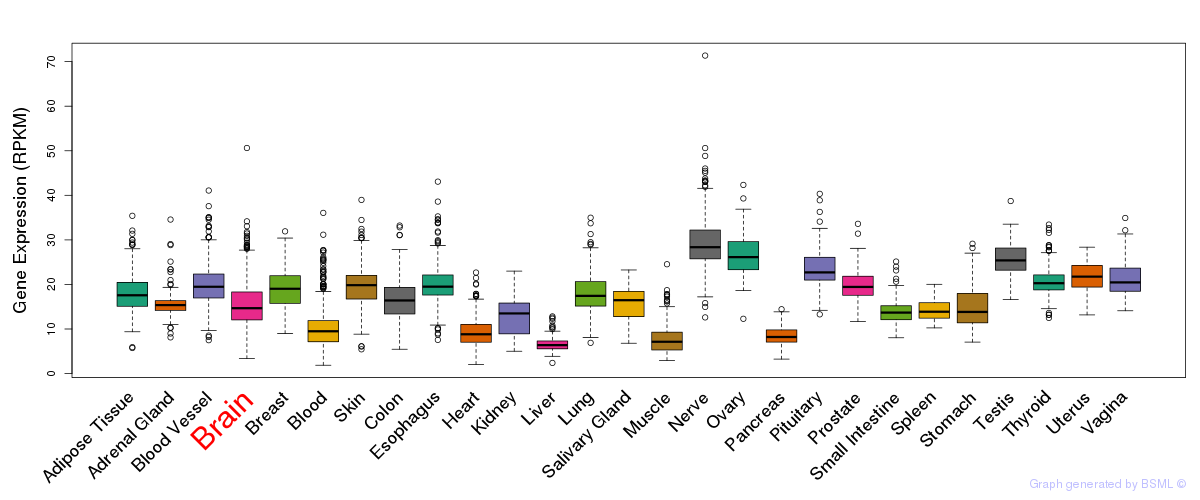
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| C17orf70 | 0.86 | 0.88 |
| TRIM46 | 0.86 | 0.87 |
| ISYNA1 | 0.85 | 0.85 |
| PODXL2 | 0.85 | 0.88 |
| FAM171A2 | 0.85 | 0.88 |
| DCAF15 | 0.85 | 0.90 |
| JUP | 0.85 | 0.86 |
| GTF2H4 | 0.84 | 0.85 |
| GPR172A | 0.84 | 0.89 |
| LRWD1 | 0.84 | 0.89 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| AF347015.27 | -0.70 | -0.83 |
| AF347015.31 | -0.69 | -0.81 |
| MT-CO2 | -0.68 | -0.81 |
| AF347015.33 | -0.68 | -0.81 |
| AF347015.8 | -0.67 | -0.81 |
| C5orf53 | -0.66 | -0.70 |
| MT-CYB | -0.66 | -0.80 |
| S100B | -0.64 | -0.74 |
| AF347015.15 | -0.63 | -0.78 |
| HLA-F | -0.63 | -0.67 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| OSMAN BLADDER CANCER DN | 406 | 230 | All SZGR 2.0 genes in this pathway |
| HEIDENBLAD AMPLIFIED IN PANCREATIC CANCER | 31 | 19 | All SZGR 2.0 genes in this pathway |
| KRIGE RESPONSE TO TOSEDOSTAT 6HR UP | 953 | 554 | All SZGR 2.0 genes in this pathway |
| KRIGE RESPONSE TO TOSEDOSTAT 24HR UP | 783 | 442 | All SZGR 2.0 genes in this pathway |
| MARSON BOUND BY FOXP3 STIMULATED | 1022 | 619 | All SZGR 2.0 genes in this pathway |
| KUUSELO PANCREATIC CANCER 19Q13 AMPLIFICATION | 35 | 22 | All SZGR 2.0 genes in this pathway |
| WANG TUMOR INVASIVENESS UP | 374 | 247 | All SZGR 2.0 genes in this pathway |
| YOSHIMURA MAPK8 TARGETS UP | 1305 | 895 | All SZGR 2.0 genes in this pathway |
| YOSHIMURA MAPK8 TARGETS DN | 366 | 257 | All SZGR 2.0 genes in this pathway |