Gene Page: EXOC1
Summary ?
| GeneID | 55763 |
| Symbol | EXOC1 |
| Synonyms | BM-102|SEC3|SEC3L1|SEC3P |
| Description | exocyst complex component 1 |
| Reference | MIM:607879|HGNC:HGNC:30380|Ensembl:ENSG00000090989|HPRD:07615|Vega:OTTHUMG00000102165 |
| Gene type | protein-coding |
| Map location | 4q12 |
| Pascal p-value | 0.011 |
| Sherlock p-value | 0.533 |
| Fetal beta | 0.116 |
| eGene | Meta |
| Support | EXOCYTOSIS CompositeSet |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| PMID:cooccur | High-throughput literature-search | Systematic search in PubMed for genes co-occurring with SCZ keywords. A total of 3027 genes were included. | |
| Literature | High-throughput literature-search | Co-occurance with Schizophrenia keywords: schizophrenia,schizophrenias | Click to show details |
Section I. Genetics and epigenetics annotation
Section II. Transcriptome annotation
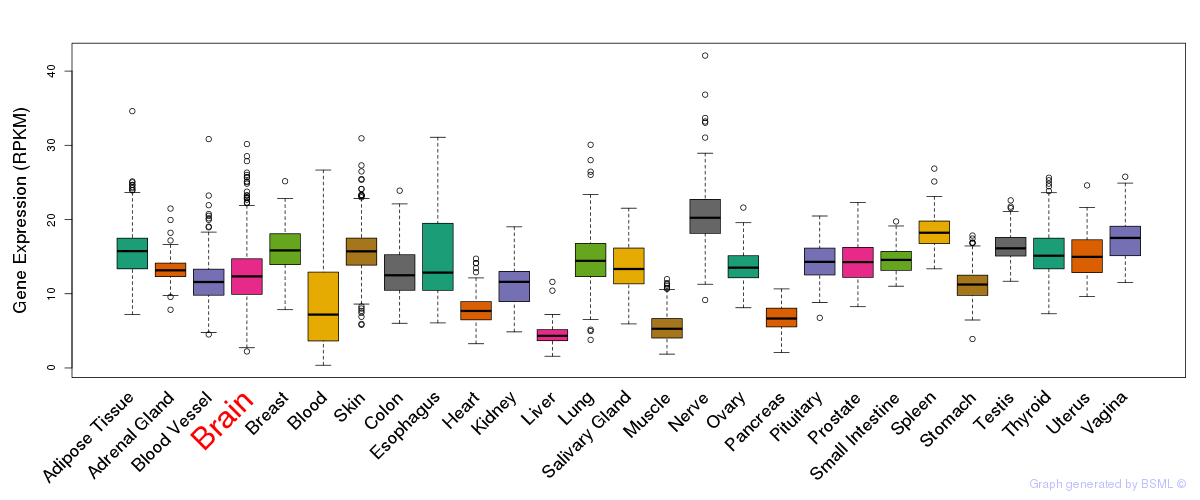
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| CARD11 | 0.74 | 0.44 |
| TTC39A | 0.73 | 0.47 |
| TANC1 | 0.73 | 0.53 |
| RGS3 | 0.72 | 0.48 |
| PTPN3 | 0.72 | 0.34 |
| ATP2A3 | 0.72 | 0.22 |
| AL133445.2 | 0.71 | 0.20 |
| PLCB4 | 0.71 | 0.29 |
| NTNG1 | 0.70 | 0.39 |
| SUSD2 | 0.70 | 0.33 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| IER5L | -0.22 | -0.21 |
| KLHL1 | -0.21 | -0.20 |
| GPR22 | -0.21 | -0.23 |
| MPPED1 | -0.21 | -0.23 |
| DPP4 | -0.21 | -0.22 |
| GPM6A | -0.21 | -0.25 |
| ZNF238 | -0.20 | -0.16 |
| EMID1 | -0.20 | -0.30 |
| ISLR2 | -0.20 | -0.19 |
| AC005768.1 | -0.20 | -0.27 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| PID INSULIN PATHWAY | 45 | 32 | All SZGR 2.0 genes in this pathway |
| PID ARF6 TRAFFICKING PATHWAY | 49 | 34 | All SZGR 2.0 genes in this pathway |
| REACTOME DIABETES PATHWAYS | 133 | 91 | All SZGR 2.0 genes in this pathway |
| REACTOME INSULIN SYNTHESIS AND PROCESSING | 21 | 15 | All SZGR 2.0 genes in this pathway |
| WANG LMO4 TARGETS DN | 352 | 225 | All SZGR 2.0 genes in this pathway |
| HAMAI APOPTOSIS VIA TRAIL UP | 584 | 356 | All SZGR 2.0 genes in this pathway |
| SHEN SMARCA2 TARGETS UP | 424 | 268 | All SZGR 2.0 genes in this pathway |
| BLALOCK ALZHEIMERS DISEASE DN | 1237 | 837 | All SZGR 2.0 genes in this pathway |
| VANASSE BCL2 TARGETS DN | 74 | 50 | All SZGR 2.0 genes in this pathway |
| FIGUEROA AML METHYLATION CLUSTER 6 UP | 140 | 81 | All SZGR 2.0 genes in this pathway |
| PILON KLF1 TARGETS DN | 1972 | 1213 | All SZGR 2.0 genes in this pathway |