Gene Page: ZDBF2
Summary ?
| GeneID | 57683 |
| Symbol | ZDBF2 |
| Synonyms | - |
| Description | zinc finger DBF-type containing 2 |
| Reference | HGNC:HGNC:29313|Ensembl:ENSG00000204186|Vega:OTTHUMG00000154648 |
| Gene type | protein-coding |
| Map location | 2q33.3 |
| Pascal p-value | 0.045 |
| Sherlock p-value | 0.018 |
| Support | Ascano FMRP targets |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| GSMA_IIA | Genome scan meta-analysis (All samples) | Psr: 0.00916 | |
| GSMA_IIE | Genome scan meta-analysis (European-ancestry samples) | Psr: 0.01016 |
Section I. Genetics and epigenetics annotation
Section II. Transcriptome annotation
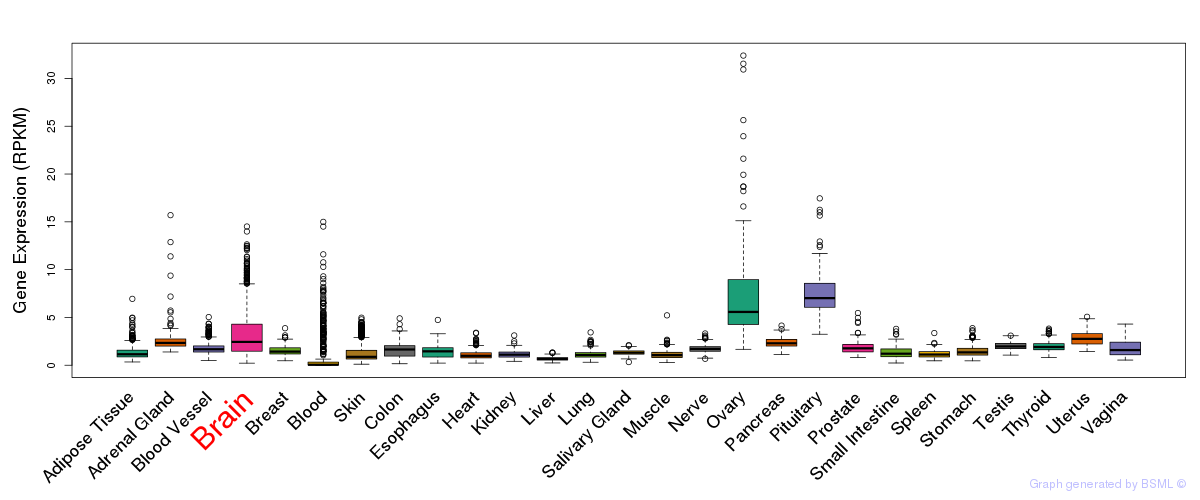
General gene expression (GTEx)
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| SIX5 | 0.85 | 0.84 |
| JUB | 0.82 | 0.74 |
| NFATC4 | 0.79 | 0.75 |
| ERBB2 | 0.79 | 0.77 |
| AL161668.2 | 0.79 | 0.77 |
| SOX9 | 0.78 | 0.71 |
| EPHB4 | 0.78 | 0.75 |
| CRB2 | 0.78 | 0.65 |
| SMO | 0.78 | 0.74 |
| TRIP10 | 0.78 | 0.78 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| EPHX4 | -0.34 | -0.42 |
| FBXW7 | -0.33 | -0.39 |
| NPM2 | -0.33 | -0.34 |
| SERPINI1 | -0.33 | -0.40 |
| GLRX2 | -0.33 | -0.44 |
| ABCC12 | -0.32 | -0.41 |
| CHN1 | -0.31 | -0.27 |
| LNX1 | -0.31 | -0.35 |
| KCNV1 | -0.31 | -0.37 |
| NRSN1 | -0.31 | -0.36 |
Section III. Gene Ontology annotation
| Molecular function | GO term | Evidence | Neuro keywords | PubMed ID |
|---|---|---|---|---|
| GO:0003676 | nucleic acid binding | IEA | - | |
| GO:0008270 | zinc ion binding | IEA | - | |
| GO:0046872 | metal ion binding | IEA | - |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| DODD NASOPHARYNGEAL CARCINOMA DN | 1375 | 806 | All SZGR 2.0 genes in this pathway |
| IVANOVA HEMATOPOIESIS STEM CELL | 254 | 164 | All SZGR 2.0 genes in this pathway |
| ACEVEDO LIVER CANCER WITH H3K9ME3 UP | 141 | 75 | All SZGR 2.0 genes in this pathway |
| WONG ADULT TISSUE STEM MODULE | 721 | 492 | All SZGR 2.0 genes in this pathway |