Gene Page: SSTR5
Summary ?
| GeneID | 6755 |
| Symbol | SSTR5 |
| Synonyms | SS-5-R |
| Description | somatostatin receptor 5 |
| Reference | MIM:182455|HGNC:HGNC:11334|HPRD:01677| |
| Gene type | protein-coding |
| Map location | 16p13.3 |
| Pascal p-value | 0.958 |
| Fetal beta | -0.004 |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| PMID:cooccur | High-throughput literature-search | Systematic search in PubMed for genes co-occurring with SCZ keywords. A total of 3027 genes were included. | |
| Literature | High-throughput literature-search | Co-occurance with Schizophrenia keywords: schizophrenia,schizophrenic,schizophrenias | Click to show details |
Section I. Genetics and epigenetics annotation
Section II. Transcriptome annotation
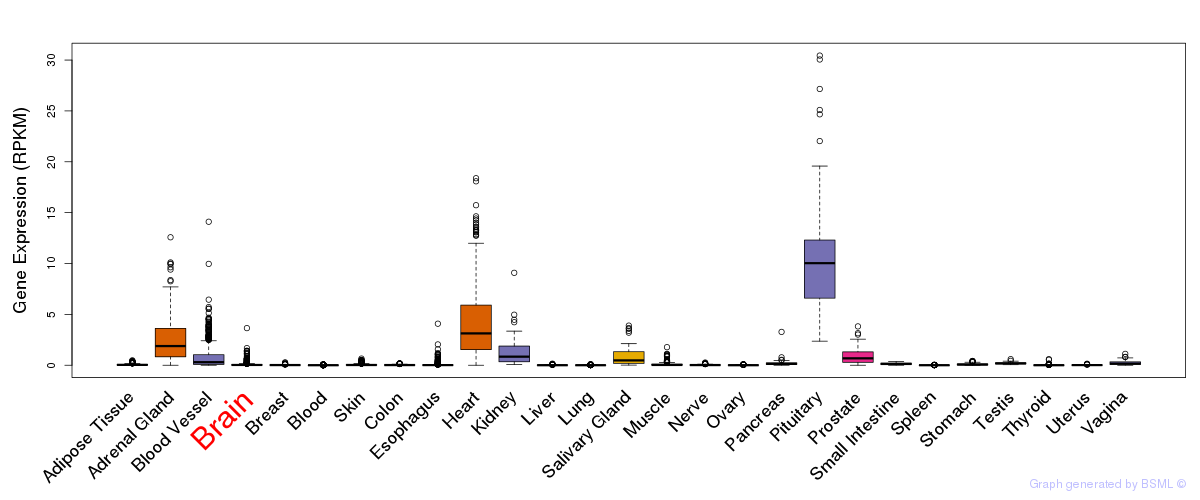
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| MSH5 | 0.58 | 0.50 |
| NPEPL1 | 0.57 | 0.42 |
| CCDC84 | 0.56 | 0.47 |
| PABPC1L | 0.56 | 0.46 |
| KIAA1407 | 0.56 | 0.48 |
| VARS2 | 0.55 | 0.44 |
| AC008686.2 | 0.55 | 0.39 |
| CILP | 0.55 | 0.35 |
| STAC3 | 0.55 | 0.43 |
| AGER | 0.55 | 0.44 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| TUBA1 | -0.33 | -0.37 |
| TPST2 | -0.33 | -0.37 |
| CCM2 | -0.33 | -0.36 |
| FAIM2 | -0.32 | -0.35 |
| PINK1 | -0.32 | -0.34 |
| HLA-C | -0.31 | -0.33 |
| PAH | -0.31 | -0.32 |
| SQSTM1 | -0.31 | -0.35 |
| ALDOA | -0.31 | -0.31 |
| FBXO2 | -0.31 | -0.33 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| KEGG NEUROACTIVE LIGAND RECEPTOR INTERACTION | 272 | 195 | All SZGR 2.0 genes in this pathway |
| REACTOME SIGNALING BY GPCR | 920 | 449 | All SZGR 2.0 genes in this pathway |
| REACTOME PEPTIDE LIGAND BINDING RECEPTORS | 188 | 108 | All SZGR 2.0 genes in this pathway |
| REACTOME CLASS A1 RHODOPSIN LIKE RECEPTORS | 305 | 177 | All SZGR 2.0 genes in this pathway |
| REACTOME GPCR DOWNSTREAM SIGNALING | 805 | 368 | All SZGR 2.0 genes in this pathway |
| REACTOME G ALPHA I SIGNALLING EVENTS | 195 | 114 | All SZGR 2.0 genes in this pathway |
| REACTOME GPCR LIGAND BINDING | 408 | 246 | All SZGR 2.0 genes in this pathway |
| SCHLOSSER SERUM RESPONSE DN | 712 | 443 | All SZGR 2.0 genes in this pathway |
| NIKOLSKY BREAST CANCER 16P13 AMPLICON | 120 | 49 | All SZGR 2.0 genes in this pathway |
| PENG GLUCOSE DEPRIVATION DN | 169 | 112 | All SZGR 2.0 genes in this pathway |
| MIKKELSEN MCV6 HCP WITH H3K27ME3 | 435 | 318 | All SZGR 2.0 genes in this pathway |
| MIKKELSEN MEF HCP WITH H3K27ME3 | 590 | 403 | All SZGR 2.0 genes in this pathway |
| KINNEY DNMT1 METHYLATION TARGETS | 14 | 6 | All SZGR 2.0 genes in this pathway |