Gene Page: CEP290
Summary ?
| GeneID | 80184 |
| Symbol | CEP290 |
| Synonyms | 3H11Ag|BBS14|CT87|JBTS5|LCA10|MKS4|NPHP6|POC3|SLSN6|rd16 |
| Description | centrosomal protein 290kDa |
| Reference | MIM:610142|HGNC:HGNC:29021|Ensembl:ENSG00000198707|HPRD:08585|HPRD:10856|Vega:OTTHUMG00000169871 |
| Gene type | protein-coding |
| Map location | 12q21.32 |
| Pascal p-value | 0.049 |
| Sherlock p-value | 0.128 |
| Fetal beta | 1.059 |
| eGene | Cerebellum Myers' cis & trans |
| Support | CompositeSet |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| DNM:Fromer_2014 | Whole Exome Sequencing analysis | This study reported a WES study of 623 schizophrenia trios, reporting DNMs using genomic DNA. | |
| PMID:cooccur | High-throughput literature-search | Systematic search in PubMed for genes co-occurring with SCZ keywords. A total of 3027 genes were included. | |
| Literature | High-throughput literature-search | Co-occurance with Schizophrenia keywords: schizophrenia,schizophrenias | Click to show details |
| GO_Annotation | Mapping neuro-related keywords to Gene Ontology annotations | Hits with neuro-related keywords: 2 |
Section I. Genetics and epigenetics annotation
 DNM table
DNM table
| Gene | Chromosome | Position | Ref | Alt | Transcript | AA change | Mutation type | Sift | CG46 | Trait | Study |
|---|---|---|---|---|---|---|---|---|---|---|---|
| CEP290 | chr12 | 88508231 | G | C | NM_025114 | p.673S>C | missense | Schizophrenia | DNM:Fromer_2014 |
 eQTL annotation
eQTL annotation
| SNP ID | Chromosome | Position | eGene | Gene Entrez ID | pvalue | qvalue | TSS distance | eQTL type |
|---|---|---|---|---|---|---|---|---|
| snp_a-1904613 | 0 | CEP290 | 80184 | 0.17 | trans | |||
| rs4562302 | chr8 | 60173690 | CEP290 | 80184 | 0.11 | trans |
Section II. Transcriptome annotation
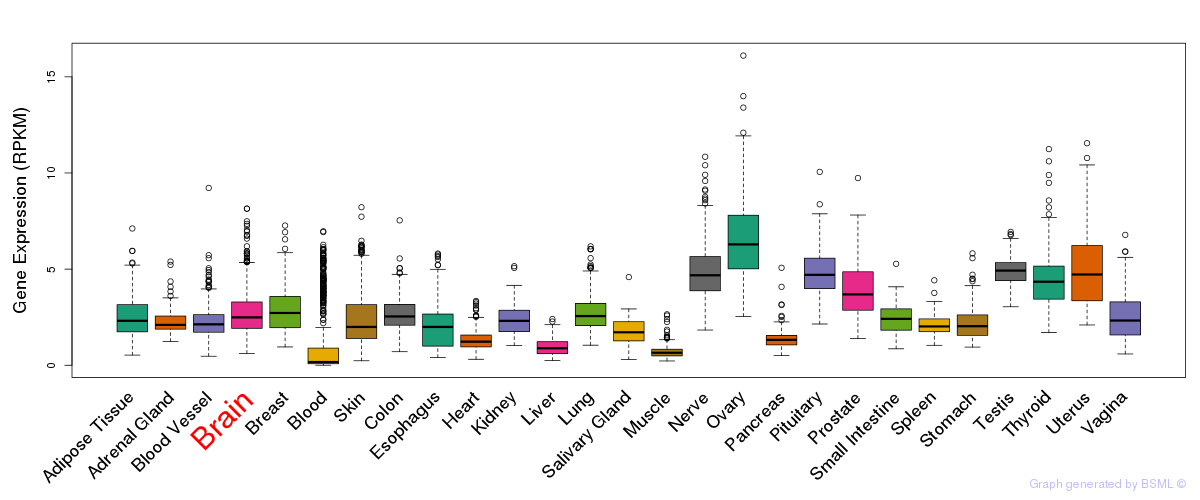
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
No co-expressed genes in brain regions
Section III. Gene Ontology annotation
| Molecular function | GO term | Evidence | Neuro keywords | PubMed ID |
|---|---|---|---|---|
| GO:0005515 | protein binding | IPI | 16682973 | |
| GO:0051011 | microtubule minus-end binding | IDA | 16682973 | |
| GO:0016563 | transcription activator activity | IDA | 16682973 | |
| Biological process | GO term | Evidence | Neuro keywords | PubMed ID |
| GO:0030902 | hindbrain development | ISS | Brain (GO term level: 8) | 16682973 |
| GO:0030030 | cell projection organization | IEA | axon (GO term level: 7) | - |
| GO:0042462 | eye photoreceptor cell development | ISS | 16682973 | |
| GO:0030916 | otic vesicle formation | ISS | 16682973 | |
| GO:0015031 | protein transport | ISS | - | |
| GO:0048793 | pronephros development | ISS | 16682973 | |
| Cellular component | GO term | Evidence | Neuro keywords | PubMed ID |
| GO:0000930 | gamma-tubulin complex | IDA | 16682973 | |
| GO:0005813 | centrosome | IDA | 14654843 | |
| GO:0005829 | cytosol | IDA | 16682973 | |
| GO:0005634 | nucleus | IDA | 16682973 | |
| GO:0005737 | cytoplasm | IDA | 14654843 | |
| GO:0009986 | cell surface | IDA | 12493773 | |
| GO:0005929 | cilium | IEA | - | |
| GO:0032391 | photoreceptor connecting cilium | ISS | 16682973 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| REACTOME CELL CYCLE | 421 | 253 | All SZGR 2.0 genes in this pathway |
| REACTOME CELL CYCLE MITOTIC | 325 | 185 | All SZGR 2.0 genes in this pathway |
| REACTOME RECRUITMENT OF MITOTIC CENTROSOME PROTEINS AND COMPLEXES | 66 | 43 | All SZGR 2.0 genes in this pathway |
| REACTOME LOSS OF NLP FROM MITOTIC CENTROSOMES | 59 | 38 | All SZGR 2.0 genes in this pathway |
| REACTOME MITOTIC G2 G2 M PHASES | 81 | 50 | All SZGR 2.0 genes in this pathway |
| SENGUPTA NASOPHARYNGEAL CARCINOMA WITH LMP1 UP | 408 | 247 | All SZGR 2.0 genes in this pathway |
| HORIUCHI WTAP TARGETS UP | 306 | 188 | All SZGR 2.0 genes in this pathway |
| LEE NEURAL CREST STEM CELL DN | 118 | 79 | All SZGR 2.0 genes in this pathway |
| HAMAI APOPTOSIS VIA TRAIL UP | 584 | 356 | All SZGR 2.0 genes in this pathway |
| PUJANA BRCA2 PCC NETWORK | 423 | 265 | All SZGR 2.0 genes in this pathway |
| NUYTTEN NIPP1 TARGETS UP | 769 | 437 | All SZGR 2.0 genes in this pathway |
| BROWNE HCMV INFECTION 14HR DN | 298 | 200 | All SZGR 2.0 genes in this pathway |
| AGUIRRE PANCREATIC CANCER COPY NUMBER DN | 238 | 145 | All SZGR 2.0 genes in this pathway |
| PURBEY TARGETS OF CTBP1 AND SATB1 UP | 87 | 52 | All SZGR 2.0 genes in this pathway |
| ZWANG CLASS 1 TRANSIENTLY INDUCED BY EGF | 516 | 308 | All SZGR 2.0 genes in this pathway |