Gene Page: TSPAN18
Summary ?
| GeneID | 90139 |
| Symbol | TSPAN18 |
| Synonyms | TSPAN |
| Description | tetraspanin 18 |
| Reference | HGNC:HGNC:20660|Ensembl:ENSG00000157570|HPRD:17435|Vega:OTTHUMG00000163883 |
| Gene type | protein-coding |
| Map location | 11p11.2 |
| Pascal p-value | 0.222 |
| DMG | 1 (# studies) |
| eGene | Cerebellum Myers' cis & trans |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:GWAScat | Genome-wide Association Studies | This data set includes 560 SNPs associated with schizophrenia. A total of 486 genes were mapped to these SNPs within 50kb. | |
| CV:GWASdb | Genome-wide Association Studies | GWASdb records for schizophrenia | |
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| DMG:Wockner_2014 | Genome-wide DNA methylation analysis | This dataset includes 4641 differentially methylated probes corresponding to 2929 unique genes between schizophrenia patients (n=24) and controls (n=24). | 1 |
| PMID:cooccur | High-throughput literature-search | Systematic search in PubMed for genes co-occurring with SCZ keywords. A total of 3027 genes were included. | |
| Literature | High-throughput literature-search | Co-occurance with Schizophrenia keywords: schizophrenia,schizophrenias | Click to show details |
Section I. Genetics and epigenetics annotation
 Differentially methylated gene
Differentially methylated gene
| Probe | Chromosome | Position | Nearest gene | P (dis) | Beta (dis) | FDR (dis) | Study |
|---|---|---|---|---|---|---|---|
| cg13989519 | 11 | 44820674 | TSPAN18 | 2.372E-4 | -0.407 | 0.037 | DMG:Wockner_2014 |
 eQTL annotation
eQTL annotation
| SNP ID | Chromosome | Position | eGene | Gene Entrez ID | pvalue | qvalue | TSS distance | eQTL type |
|---|---|---|---|---|---|---|---|---|
| rs6687849 | chr1 | 175904808 | TSPAN18 | 90139 | 0.18 | trans | ||
| rs2502827 | chr1 | 176044216 | TSPAN18 | 90139 | 0.03 | trans | ||
| rs17572651 | chr1 | 218943612 | TSPAN18 | 90139 | 0.08 | trans | ||
| rs6753473 | chr2 | 26526418 | TSPAN18 | 90139 | 0.01 | trans | ||
| rs17028363 | chr2 | 41913161 | TSPAN18 | 90139 | 0.15 | trans | ||
| rs16829545 | chr2 | 151977407 | TSPAN18 | 90139 | 2.308E-30 | trans | ||
| rs16841750 | chr2 | 158288461 | TSPAN18 | 90139 | 3.988E-5 | trans | ||
| rs16863816 | chr2 | 177292762 | TSPAN18 | 90139 | 0.13 | trans | ||
| rs7584986 | chr2 | 184111432 | TSPAN18 | 90139 | 1.898E-4 | trans | ||
| rs6741060 | chr2 | 217169088 | TSPAN18 | 90139 | 0.15 | trans | ||
| rs10183246 | chr2 | 237187882 | TSPAN18 | 90139 | 0.11 | trans | ||
| rs9810143 | chr3 | 5060209 | TSPAN18 | 90139 | 3.33E-5 | trans | ||
| rs6797307 | chr3 | 8601563 | TSPAN18 | 90139 | 0.01 | trans | ||
| rs6773248 | chr3 | 174790333 | TSPAN18 | 90139 | 0.11 | trans | ||
| rs6797728 | chr3 | 174790417 | TSPAN18 | 90139 | 0.11 | trans | ||
| rs12642340 | chr4 | 35390674 | TSPAN18 | 90139 | 0.13 | trans | ||
| rs17008119 | chr4 | 125321425 | TSPAN18 | 90139 | 0.06 | trans | ||
| rs7692715 | chr4 | 125335209 | TSPAN18 | 90139 | 0.01 | trans | ||
| rs4488887 | chr4 | 177695833 | TSPAN18 | 90139 | 0.12 | trans | ||
| rs10474151 | chr5 | 84286402 | TSPAN18 | 90139 | 0.04 | trans | ||
| rs13360496 | 0 | TSPAN18 | 90139 | 0 | trans | |||
| rs13191953 | chr6 | 24151042 | TSPAN18 | 90139 | 0.14 | trans | ||
| rs12196880 | chr6 | 24313746 | TSPAN18 | 90139 | 0.07 | trans | ||
| rs10806988 | chr6 | 24341768 | TSPAN18 | 90139 | 0.12 | trans | ||
| rs16890367 | chr6 | 38078448 | TSPAN18 | 90139 | 0 | trans | ||
| rs12706918 | chr7 | 80266147 | TSPAN18 | 90139 | 0.06 | trans | ||
| rs6996695 | chr8 | 77540580 | TSPAN18 | 90139 | 0.1 | trans | ||
| rs3118341 | chr9 | 25185518 | TSPAN18 | 90139 | 0.01 | trans | ||
| rs9406868 | chr9 | 25223372 | TSPAN18 | 90139 | 0.07 | trans | ||
| rs11139334 | chr9 | 84209393 | TSPAN18 | 90139 | 0.02 | trans | ||
| rs2393316 | chr10 | 59333070 | TSPAN18 | 90139 | 0.01 | trans | ||
| rs17124813 | chr10 | 110347464 | TSPAN18 | 90139 | 0.06 | trans | ||
| rs7972699 | chr12 | 105155290 | TSPAN18 | 90139 | 0.17 | trans | ||
| rs7488439 | chr12 | 105155814 | TSPAN18 | 90139 | 0.05 | trans | ||
| rs17036489 | chr12 | 105292283 | TSPAN18 | 90139 | 0.02 | trans | ||
| rs16955618 | chr15 | 29937543 | TSPAN18 | 90139 | 1.787E-32 | trans | ||
| rs12914345 | chr15 | 77670400 | TSPAN18 | 90139 | 0.1 | trans | ||
| rs17711752 | chr16 | 34436184 | TSPAN18 | 90139 | 0.2 | trans | ||
| rs11873184 | chr18 | 1584081 | TSPAN18 | 90139 | 0.01 | trans | ||
| rs7274477 | chr20 | 1676942 | TSPAN18 | 90139 | 0.02 | trans | ||
| rs1041786 | chr21 | 22617710 | TSPAN18 | 90139 | 5.398E-7 | trans | ||
| rs16990575 | chrX | 32691327 | TSPAN18 | 90139 | 0 | trans | ||
| rs16993189 | chrX | 144666166 | TSPAN18 | 90139 | 0.17 | trans |
Section II. Transcriptome annotation
General gene expression (GTEx)
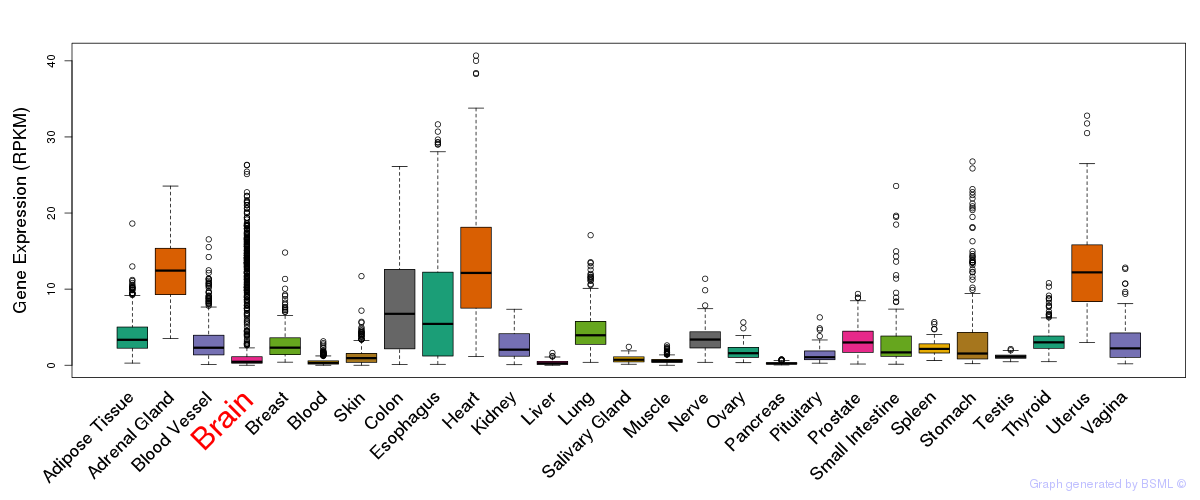
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
No co-expressed genes in brain regions
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| LIU PROSTATE CANCER DN | 481 | 290 | All SZGR 2.0 genes in this pathway |
| THUM SYSTOLIC HEART FAILURE UP | 423 | 283 | All SZGR 2.0 genes in this pathway |
| WANG ESOPHAGUS CANCER VS NORMAL UP | 121 | 72 | All SZGR 2.0 genes in this pathway |
| SCHAEFFER PROSTATE DEVELOPMENT 48HR DN | 428 | 306 | All SZGR 2.0 genes in this pathway |
| SHEPARD CRUSH AND BURN MUTANT UP | 197 | 110 | All SZGR 2.0 genes in this pathway |
| SHEPARD BMYB MORPHOLINO UP | 205 | 126 | All SZGR 2.0 genes in this pathway |
| RODWELL AGING KIDNEY UP | 487 | 303 | All SZGR 2.0 genes in this pathway |
| TOYOTA TARGETS OF MIR34B AND MIR34C | 463 | 262 | All SZGR 2.0 genes in this pathway |
| MEISSNER NPC HCP WITH H3K4ME3 AND H3K27ME3 | 142 | 103 | All SZGR 2.0 genes in this pathway |
| MEISSNER BRAIN HCP WITH H3K4ME3 AND H3K27ME3 | 1069 | 729 | All SZGR 2.0 genes in this pathway |
| MIKKELSEN NPC HCP WITH H3K4ME3 AND H3K27ME3 | 210 | 148 | All SZGR 2.0 genes in this pathway |
| HOELZEL NF1 TARGETS UP | 139 | 93 | All SZGR 2.0 genes in this pathway |