Gene Page: ORMDL1
Summary ?
| GeneID | 94101 |
| Symbol | ORMDL1 |
| Synonyms | - |
| Description | ORMDL sphingolipid biosynthesis regulator 1 |
| Reference | MIM:610073|HGNC:HGNC:16036|Ensembl:ENSG00000128699|HPRD:15092|Vega:OTTHUMG00000132661 |
| Gene type | protein-coding |
| Map location | 2q32 |
| Pascal p-value | 0.249 |
| Fetal beta | 0.717 |
| DMG | 1 (# studies) |
| eGene | Caudate basal ganglia Cerebellar Hemisphere Cerebellum Frontal Cortex BA9 Hippocampus Hypothalamus Nucleus accumbens basal ganglia Myers' cis & trans Meta |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| DMG:Jaffe_2016 | Genome-wide DNA methylation analysis | This dataset includes 2,104 probes/CpGs associated with SZ patients (n=108) compared to 136 controls at Bonferroni-adjusted P < 0.05. | 2 |
| DNM:Guipponi_2014 | Whole Exome Sequencing analysis | 49 DNMs were identified by comparing the exome of 53 individuals with sporadic SCZ and of their non-affected parents |
Section I. Genetics and epigenetics annotation
 DNM table
DNM table
| Gene | Chromosome | Position | Ref | Alt | Transcript | AA change | Mutation type | Sift | CG46 | Trait | Study |
|---|---|---|---|---|---|---|---|---|---|---|---|
| ORMDL1 | G | A | NM_001128150 | p.K100K | synonymous | NA | NA | Schizophrenia | DNM:Guipponi_2014 |
 Differentially methylated gene
Differentially methylated gene
| Probe | Chromosome | Position | Nearest gene | P (dis) | Beta (dis) | FDR (dis) | Study |
|---|---|---|---|---|---|---|---|
| cg08891424 | 2 | 190648631 | ORMDL1 | 6.42E-10 | -0.013 | 9.43E-7 | DMG:Jaffe_2016 |
| cg26454191 | 2 | 190648463 | ORMDL1 | 7.2E-8 | -0.01 | 1.73E-5 | DMG:Jaffe_2016 |
 eQTL annotation
eQTL annotation
| SNP ID | Chromosome | Position | eGene | Gene Entrez ID | pvalue | qvalue | TSS distance | eQTL type |
|---|---|---|---|---|---|---|---|---|
| rs4667295 | chr2 | 190504389 | ORMDL1 | 94101 | 0.02 | cis | ||
| rs2289226 | chr2 | 190526204 | ORMDL1 | 94101 | 5.878E-4 | cis | ||
| rs7557519 | chr2 | 190583186 | ORMDL1 | 94101 | 0.05 | cis | ||
| rs1550388 | chr2 | 190603779 | ORMDL1 | 94101 | 0.01 | cis | ||
| rs2033870 | chr2 | 190603829 | ORMDL1 | 94101 | 0 | cis | ||
| rs4666783 | chr2 | 190612979 | ORMDL1 | 94101 | 0.01 | cis | ||
| rs3791767 | chr2 | 190639914 | ORMDL1 | 94101 | 0.02 | cis | ||
| rs7591929 | chr2 | 190643256 | ORMDL1 | 94101 | 5.737E-4 | cis | ||
| rs2289226 | chr2 | 190526204 | ORMDL1 | 94101 | 0.05 | trans | ||
| rs2033870 | chr2 | 190603829 | ORMDL1 | 94101 | 0.16 | trans | ||
| rs7591929 | chr2 | 190643256 | ORMDL1 | 94101 | 0.04 | trans | ||
| rs7495645 | chr15 | 92139578 | ORMDL1 | 94101 | 0.07 | trans |
Section II. Transcriptome annotation
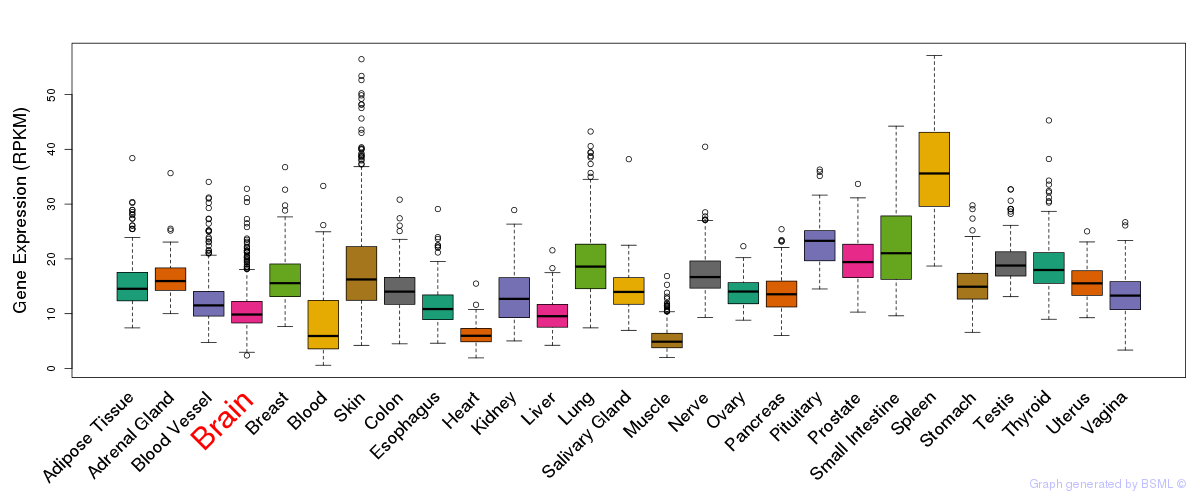
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
No co-expressed genes in brain regions
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| CHARAFE BREAST CANCER LUMINAL VS BASAL DN | 455 | 304 | All SZGR 2.0 genes in this pathway |
| SENESE HDAC3 TARGETS UP | 501 | 327 | All SZGR 2.0 genes in this pathway |
| KINSEY TARGETS OF EWSR1 FLII FUSION UP | 1278 | 748 | All SZGR 2.0 genes in this pathway |
| GRAESSMANN APOPTOSIS BY DOXORUBICIN DN | 1781 | 1082 | All SZGR 2.0 genes in this pathway |
| BENPORATH CYCLING GENES | 648 | 385 | All SZGR 2.0 genes in this pathway |
| UEDA CENTRAL CLOCK | 88 | 62 | All SZGR 2.0 genes in this pathway |
| JIANG HYPOXIA NORMAL | 311 | 205 | All SZGR 2.0 genes in this pathway |
| CREIGHTON ENDOCRINE THERAPY RESISTANCE 3 | 720 | 440 | All SZGR 2.0 genes in this pathway |
| CREIGHTON ENDOCRINE THERAPY RESISTANCE 5 | 482 | 296 | All SZGR 2.0 genes in this pathway |
| MARSON BOUND BY FOXP3 STIMULATED | 1022 | 619 | All SZGR 2.0 genes in this pathway |
| COLINA TARGETS OF 4EBP1 AND 4EBP2 | 356 | 214 | All SZGR 2.0 genes in this pathway |
| WANG RESPONSE TO GSK3 INHIBITOR SB216763 UP | 397 | 206 | All SZGR 2.0 genes in this pathway |
| BRUINS UVC RESPONSE LATE | 1137 | 655 | All SZGR 2.0 genes in this pathway |
| RATTENBACHER BOUND BY CELF1 | 467 | 251 | All SZGR 2.0 genes in this pathway |