Gene Page: CHRNA3
Summary ?
| GeneID | 1136 |
| Symbol | CHRNA3 |
| Synonyms | LNCR2|NACHRA3|PAOD2 |
| Description | cholinergic receptor nicotinic alpha 3 subunit |
| Reference | MIM:118503|HGNC:HGNC:1957|Ensembl:ENSG00000080644|HPRD:07511|Vega:OTTHUMG00000143863 |
| Gene type | protein-coding |
| Map location | 15q24 |
| Pascal p-value | 4.988E-9 |
| eGene | Caudate basal ganglia Nucleus accumbens basal ganglia Putamen basal ganglia Myers' cis & trans |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:GWAScat | Genome-wide Association Studies | This data set includes 560 SNPs associated with schizophrenia. A total of 486 genes were mapped to these SNPs within 50kb. | |
| CV:PGC128 | Genome-wide Association Study | A multi-stage schizophrenia GWAS of up to 36,989 cases and 113,075 controls. Reported by the Schizophrenia Working Group of PGC. 128 independent associations spanning 108 loci | |
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| PMID:cooccur | High-throughput literature-search | Systematic search in PubMed for genes co-occurring with SCZ keywords. A total of 3027 genes were included. | |
| Literature | High-throughput literature-search | Co-occurance with Schizophrenia keywords: schizophrenia,schizophrenic,schizophrenias | Click to show details |
| GO_Annotation | Mapping neuro-related keywords to Gene Ontology annotations | Hits with neuro-related keywords: 16 |
Section I. Genetics and epigenetics annotation
 CV:PGC128
CV:PGC128
| SNP ID | Chromosome | Position | Allele | P | Function | Gene | Up/Down Distance |
|---|---|---|---|---|---|---|---|
| rs8042374 | chr15 | 78908032 | AG | 1.865E-12 | intronic | CHRNA3 |
 eQTL annotation
eQTL annotation
| SNP ID | Chromosome | Position | eGene | Gene Entrez ID | pvalue | qvalue | TSS distance | eQTL type |
|---|---|---|---|---|---|---|---|---|
| rs17130103 | chr1 | 88150541 | CHRNA3 | 1136 | 0.11 | trans | ||
| rs16823145 | chr1 | 184397914 | CHRNA3 | 1136 | 0.04 | trans | ||
| rs6666913 | chr1 | 237092144 | CHRNA3 | 1136 | 0.13 | trans | ||
| rs7596511 | chr2 | 120010898 | CHRNA3 | 1136 | 0.01 | trans | ||
| rs2324466 | chr3 | 76100753 | CHRNA3 | 1136 | 0.15 | trans | ||
| rs17013713 | chr3 | 76105150 | CHRNA3 | 1136 | 0.05 | trans | ||
| rs17013722 | chr3 | 76107656 | CHRNA3 | 1136 | 0.15 | trans | ||
| rs17013724 | chr3 | 76107684 | CHRNA3 | 1136 | 0.15 | trans | ||
| rs16831155 | chr3 | 173574763 | CHRNA3 | 1136 | 0.15 | trans | ||
| rs10212684 | chr4 | 102822506 | CHRNA3 | 1136 | 0.1 | trans | ||
| rs9986296 | chr5 | 3199318 | CHRNA3 | 1136 | 0.12 | trans | ||
| rs348150 | chr8 | 73371619 | CHRNA3 | 1136 | 0.1 | trans | ||
| rs16938220 | chr8 | 73377940 | CHRNA3 | 1136 | 0.07 | trans | ||
| rs17233214 | chr11 | 21367388 | CHRNA3 | 1136 | 0.13 | trans | ||
| rs2069410 | chr12 | 56364644 | CHRNA3 | 1136 | 0.01 | trans | ||
| rs17834315 | chr13 | 37991091 | CHRNA3 | 1136 | 0.06 | trans | ||
| rs10498433 | chr14 | 52289530 | CHRNA3 | 1136 | 0.01 | trans | ||
| rs12900783 | 15 | 78815523 | CHRNA3 | ENSG00000080644.11 | 1.393E-6 | 0 | 98114 | gtex_brain_putamen_basal |
| rs12594391 | 15 | 78816240 | CHRNA3 | ENSG00000080644.11 | 1.354E-6 | 0 | 97397 | gtex_brain_putamen_basal |
| rs12910969 | 15 | 78816980 | CHRNA3 | ENSG00000080644.11 | 1.323E-6 | 0 | 96657 | gtex_brain_putamen_basal |
| rs12910726 | 15 | 78817034 | CHRNA3 | ENSG00000080644.11 | 1.318E-6 | 0 | 96603 | gtex_brain_putamen_basal |
| rs1504545 | 15 | 78818471 | CHRNA3 | ENSG00000080644.11 | 1.259E-6 | 0 | 95166 | gtex_brain_putamen_basal |
| rs952215 | 15 | 78819153 | CHRNA3 | ENSG00000080644.11 | 1.231E-6 | 0 | 94484 | gtex_brain_putamen_basal |
| rs952216 | 15 | 78819202 | CHRNA3 | ENSG00000080644.11 | 1.229E-6 | 0 | 94435 | gtex_brain_putamen_basal |
| rs12902493 | 15 | 78819275 | CHRNA3 | ENSG00000080644.11 | 1.226E-6 | 0 | 94362 | gtex_brain_putamen_basal |
| rs12903129 | 15 | 78819606 | CHRNA3 | ENSG00000080644.11 | 1.214E-6 | 0 | 94031 | gtex_brain_putamen_basal |
| rs11636131 | 15 | 78821606 | CHRNA3 | ENSG00000080644.11 | 1.15E-6 | 0 | 92031 | gtex_brain_putamen_basal |
| rs11632604 | 15 | 78821914 | CHRNA3 | ENSG00000080644.11 | 1.141E-6 | 0 | 91723 | gtex_brain_putamen_basal |
| rs12910289 | 15 | 78822065 | CHRNA3 | ENSG00000080644.11 | 1.044E-6 | 0 | 91572 | gtex_brain_putamen_basal |
| rs35773115 | 15 | 78822189 | CHRNA3 | ENSG00000080644.11 | 2.036E-6 | 0 | 91448 | gtex_brain_putamen_basal |
| rs12910100 | 15 | 78822667 | CHRNA3 | ENSG00000080644.11 | 1.13E-6 | 0 | 90970 | gtex_brain_putamen_basal |
| rs12915428 | 15 | 78823368 | CHRNA3 | ENSG00000080644.11 | 1.124E-6 | 0 | 90269 | gtex_brain_putamen_basal |
| rs12900612 | 15 | 78823993 | CHRNA3 | ENSG00000080644.11 | 1.094E-6 | 0 | 89644 | gtex_brain_putamen_basal |
| rs1847530 | 15 | 78824031 | CHRNA3 | ENSG00000080644.11 | 1.119E-6 | 0 | 89606 | gtex_brain_putamen_basal |
| rs1504546 | 15 | 78824235 | CHRNA3 | ENSG00000080644.11 | 1.116E-6 | 0 | 89402 | gtex_brain_putamen_basal |
| rs145042403 | 15 | 78824287 | CHRNA3 | ENSG00000080644.11 | 1.116E-6 | 0 | 89350 | gtex_brain_putamen_basal |
| rs11630349 | 15 | 78824608 | CHRNA3 | ENSG00000080644.11 | 1.115E-6 | 0 | 89029 | gtex_brain_putamen_basal |
| rs12906284 | 15 | 78825105 | CHRNA3 | ENSG00000080644.11 | 1.115E-6 | 0 | 88532 | gtex_brain_putamen_basal |
| rs12907425 | 15 | 78825149 | CHRNA3 | ENSG00000080644.11 | 1.115E-6 | 0 | 88488 | gtex_brain_putamen_basal |
| rs12906846 | 15 | 78825295 | CHRNA3 | ENSG00000080644.11 | 1.115E-6 | 0 | 88342 | gtex_brain_putamen_basal |
| rs12907169 | 15 | 78825498 | CHRNA3 | ENSG00000080644.11 | 1.115E-6 | 0 | 88139 | gtex_brain_putamen_basal |
| rs12906951 | 15 | 78825562 | CHRNA3 | ENSG00000080644.11 | 1.115E-6 | 0 | 88075 | gtex_brain_putamen_basal |
| rs56007453 | 15 | 78826239 | CHRNA3 | ENSG00000080644.11 | 1.115E-6 | 0 | 87398 | gtex_brain_putamen_basal |
| rs17407257 | 15 | 78826249 | CHRNA3 | ENSG00000080644.11 | 1.115E-6 | 0 | 87388 | gtex_brain_putamen_basal |
| rs12916999 | 15 | 78826912 | CHRNA3 | ENSG00000080644.11 | 1.115E-6 | 0 | 86725 | gtex_brain_putamen_basal |
| rs12915366 | 15 | 78831753 | CHRNA3 | ENSG00000080644.11 | 1.115E-6 | 0 | 81884 | gtex_brain_putamen_basal |
| rs12915652 | 15 | 78831928 | CHRNA3 | ENSG00000080644.11 | 1.115E-6 | 0 | 81709 | gtex_brain_putamen_basal |
| rs12916483 | 15 | 78832397 | CHRNA3 | ENSG00000080644.11 | 1.115E-6 | 0 | 81240 | gtex_brain_putamen_basal |
| rs3813572 | 15 | 78832588 | CHRNA3 | ENSG00000080644.11 | 1.115E-6 | 0 | 81049 | gtex_brain_putamen_basal |
| rs3813571 | 15 | 78832792 | CHRNA3 | ENSG00000080644.11 | 1.115E-6 | 0 | 80845 | gtex_brain_putamen_basal |
| rs55690619 | 15 | 78833612 | CHRNA3 | ENSG00000080644.11 | 1.28E-7 | 0 | 80025 | gtex_brain_putamen_basal |
| rs4886571 | 15 | 78833758 | CHRNA3 | ENSG00000080644.11 | 1.28E-7 | 0 | 79879 | gtex_brain_putamen_basal |
| rs4243083 | 15 | 78833830 | CHRNA3 | ENSG00000080644.11 | 1.324E-7 | 0 | 79807 | gtex_brain_putamen_basal |
| rs2292117 | 15 | 78834689 | CHRNA3 | ENSG00000080644.11 | 1.28E-7 | 0 | 78948 | gtex_brain_putamen_basal |
| rs11858230 | 15 | 78835552 | CHRNA3 | ENSG00000080644.11 | 1.28E-7 | 0 | 78085 | gtex_brain_putamen_basal |
| rs8025429 | 15 | 78836362 | CHRNA3 | ENSG00000080644.11 | 1.28E-7 | 0 | 77275 | gtex_brain_putamen_basal |
| rs67437837 | 15 | 78836631 | CHRNA3 | ENSG00000080644.11 | 1.308E-7 | 0 | 77006 | gtex_brain_putamen_basal |
| rs76712448 | 15 | 78836719 | CHRNA3 | ENSG00000080644.11 | 3.5E-7 | 0 | 76918 | gtex_brain_putamen_basal |
| rs75754317 | 15 | 78836721 | CHRNA3 | ENSG00000080644.11 | 3.424E-7 | 0 | 76916 | gtex_brain_putamen_basal |
| rs78406067 | 15 | 78836723 | CHRNA3 | ENSG00000080644.11 | 3.424E-7 | 0 | 76914 | gtex_brain_putamen_basal |
| rs77434238 | 15 | 78836726 | CHRNA3 | ENSG00000080644.11 | 3.532E-7 | 0 | 76911 | gtex_brain_putamen_basal |
| rs28392948 | 15 | 78836866 | CHRNA3 | ENSG00000080644.11 | 1.28E-7 | 0 | 76771 | gtex_brain_putamen_basal |
| rs4886572 | 15 | 78837452 | CHRNA3 | ENSG00000080644.11 | 1.28E-7 | 0 | 76185 | gtex_brain_putamen_basal |
| rs4887062 | 15 | 78837801 | CHRNA3 | ENSG00000080644.11 | 1.28E-7 | 0 | 75836 | gtex_brain_putamen_basal |
| rs2869544 | 15 | 78839400 | CHRNA3 | ENSG00000080644.11 | 6.625E-8 | 0 | 74237 | gtex_brain_putamen_basal |
| rs4887063 | 15 | 78839715 | CHRNA3 | ENSG00000080644.11 | 6.625E-8 | 0 | 73922 | gtex_brain_putamen_basal |
| rs8053 | 15 | 78841220 | CHRNA3 | ENSG00000080644.11 | 6.625E-8 | 0 | 72417 | gtex_brain_putamen_basal |
| rs1979907 | 15 | 78842239 | CHRNA3 | ENSG00000080644.11 | 6.625E-8 | 0 | 71398 | gtex_brain_putamen_basal |
| rs1979906 | 15 | 78842289 | CHRNA3 | ENSG00000080644.11 | 6.625E-8 | 0 | 71348 | gtex_brain_putamen_basal |
| rs1979905 | 15 | 78842374 | CHRNA3 | ENSG00000080644.11 | 6.625E-8 | 0 | 71263 | gtex_brain_putamen_basal |
| rs397760194 | 15 | 78842835 | CHRNA3 | ENSG00000080644.11 | 1.638E-8 | 0 | 70802 | gtex_brain_putamen_basal |
| rs4887064 | 15 | 78842847 | CHRNA3 | ENSG00000080644.11 | 6.625E-8 | 0 | 70790 | gtex_brain_putamen_basal |
| rs12907966 | 15 | 78843051 | CHRNA3 | ENSG00000080644.11 | 1.346E-6 | 0 | 70586 | gtex_brain_putamen_basal |
| rs1504548 | 15 | 78843246 | CHRNA3 | ENSG00000080644.11 | 6.625E-8 | 0 | 70391 | gtex_brain_putamen_basal |
| rs1979908 | 15 | 78843564 | CHRNA3 | ENSG00000080644.11 | 6.625E-8 | 0 | 70073 | gtex_brain_putamen_basal |
| rs880395 | 15 | 78844356 | CHRNA3 | ENSG00000080644.11 | 6.625E-8 | 0 | 69281 | gtex_brain_putamen_basal |
| rs905740 | 15 | 78844386 | CHRNA3 | ENSG00000080644.11 | 6.625E-8 | 0 | 69251 | gtex_brain_putamen_basal |
| rs7164030 | 15 | 78844661 | CHRNA3 | ENSG00000080644.11 | 6.625E-8 | 0 | 68976 | gtex_brain_putamen_basal |
| rs4275821 | 15 | 78849541 | CHRNA3 | ENSG00000080644.11 | 2.143E-11 | 0 | 64096 | gtex_brain_putamen_basal |
| rs7173512 | 15 | 78849914 | CHRNA3 | ENSG00000080644.11 | 2.11E-11 | 0 | 63723 | gtex_brain_putamen_basal |
| rs3829787 | 15 | 78856266 | CHRNA3 | ENSG00000080644.11 | 2.409E-10 | 0 | 57371 | gtex_brain_putamen_basal |
| rs871058 | 15 | 78858491 | CHRNA3 | ENSG00000080644.11 | 4.928E-8 | 0 | 55146 | gtex_brain_putamen_basal |
| rs57945453 | 15 | 78862845 | CHRNA3 | ENSG00000080644.11 | 2.018E-10 | 0 | 50792 | gtex_brain_putamen_basal |
| rs588765 | 15 | 78865425 | CHRNA3 | ENSG00000080644.11 | 4.202E-8 | 0 | 48212 | gtex_brain_putamen_basal |
| rs61012457 | 15 | 78865694 | CHRNA3 | ENSG00000080644.11 | 2.106E-10 | 0 | 47943 | gtex_brain_putamen_basal |
| rs6495306 | 15 | 78865893 | CHRNA3 | ENSG00000080644.11 | 4.18E-8 | 0 | 47744 | gtex_brain_putamen_basal |
| rs2456019 | 15 | 78868489 | CHRNA3 | ENSG00000080644.11 | 6.291E-12 | 0 | 45148 | gtex_brain_putamen_basal |
| rs601079 | 15 | 78869579 | CHRNA3 | ENSG00000080644.11 | 2.16E-7 | 0 | 44058 | gtex_brain_putamen_basal |
| rs495956 | 15 | 78869930 | CHRNA3 | ENSG00000080644.11 | 1.084E-10 | 0 | 43707 | gtex_brain_putamen_basal |
| rs495090 | 15 | 78870003 | CHRNA3 | ENSG00000080644.11 | 1.087E-10 | 0 | 43634 | gtex_brain_putamen_basal |
| rs471889 | 15 | 78870235 | CHRNA3 | ENSG00000080644.11 | 1.09E-10 | 0 | 43402 | gtex_brain_putamen_basal |
| rs680244 | 15 | 78871288 | CHRNA3 | ENSG00000080644.11 | 1.632E-7 | 0 | 42349 | gtex_brain_putamen_basal |
| rs10709956 | 15 | 78872834 | CHRNA3 | ENSG00000080644.11 | 1.763E-7 | 0 | 40803 | gtex_brain_putamen_basal |
| rs621849 | 15 | 78872861 | CHRNA3 | ENSG00000080644.11 | 2.164E-7 | 0 | 40776 | gtex_brain_putamen_basal |
| rs692780 | 15 | 78876505 | CHRNA3 | ENSG00000080644.11 | 6.649E-12 | 0 | 37132 | gtex_brain_putamen_basal |
| rs11637635 | 15 | 78877150 | CHRNA3 | ENSG00000080644.11 | 6.745E-12 | 0 | 36487 | gtex_brain_putamen_basal |
| rs481134 | 15 | 78877563 | CHRNA3 | ENSG00000080644.11 | 4.276E-8 | 0 | 36074 | gtex_brain_putamen_basal |
| rs5813927 | 15 | 78878100 | CHRNA3 | ENSG00000080644.11 | 4.178E-8 | 0 | 35537 | gtex_brain_putamen_basal |
| rs555018 | 15 | 78879242 | CHRNA3 | ENSG00000080644.11 | 2.913E-8 | 0 | 34395 | gtex_brain_putamen_basal |
| rs647041 | 15 | 78880481 | CHRNA3 | ENSG00000080644.11 | 1.319E-8 | 0 | 33156 | gtex_brain_putamen_basal |
| rs17408276 | 15 | 78881618 | CHRNA3 | ENSG00000080644.11 | 2.146E-10 | 0 | 32019 | gtex_brain_putamen_basal |
| rs584135 | 15 | 78883628 | CHRNA3 | ENSG00000080644.11 | 2.061E-12 | 0 | 30009 | gtex_brain_putamen_basal |
| rs514743 | 15 | 78884227 | CHRNA3 | ENSG00000080644.11 | 1.722E-12 | 0 | 29410 | gtex_brain_putamen_basal |
| rs615470 | 15 | 78885988 | CHRNA3 | ENSG00000080644.11 | 8.884E-12 | 0 | 27649 | gtex_brain_putamen_basal |
| rs660652 | 15 | 78887832 | CHRNA3 | ENSG00000080644.11 | 1.718E-12 | 0 | 25805 | gtex_brain_putamen_basal |
| rs472054 | 15 | 78887994 | CHRNA3 | ENSG00000080644.11 | 1.718E-12 | 0 | 25643 | gtex_brain_putamen_basal |
| rs398057851 | 15 | 78889782 | CHRNA3 | ENSG00000080644.11 | 2.934E-8 | 0 | 23855 | gtex_brain_putamen_basal |
| rs6495307 | 15 | 78890321 | CHRNA3 | ENSG00000080644.11 | 2.934E-8 | 0 | 23316 | gtex_brain_putamen_basal |
| rs12911602 | 15 | 78891449 | CHRNA3 | ENSG00000080644.11 | 2.934E-8 | 0 | 22188 | gtex_brain_putamen_basal |
| rs62010327 | 15 | 78892784 | CHRNA3 | ENSG00000080644.11 | 6.853E-10 | 0 | 20853 | gtex_brain_putamen_basal |
| rs12901300 | 15 | 78892952 | CHRNA3 | ENSG00000080644.11 | 2.934E-8 | 0 | 20685 | gtex_brain_putamen_basal |
| rs3743077 | 15 | 78894896 | CHRNA3 | ENSG00000080644.11 | 1.735E-7 | 0 | 18741 | gtex_brain_putamen_basal |
| rs62010328 | 15 | 78894971 | CHRNA3 | ENSG00000080644.11 | 3.513E-10 | 0 | 18666 | gtex_brain_putamen_basal |
| rs75104798 | 15 | 78897865 | CHRNA3 | ENSG00000080644.11 | 7.203E-11 | 0 | 15772 | gtex_brain_putamen_basal |
| rs7182583 | 15 | 78899210 | CHRNA3 | ENSG00000080644.11 | 8.888E-12 | 0 | 14427 | gtex_brain_putamen_basal |
| rs111725959 | 15 | 78899346 | CHRNA3 | ENSG00000080644.11 | 6.724E-11 | 0 | 14291 | gtex_brain_putamen_basal |
| rs7495518 | 15 | 78900574 | CHRNA3 | ENSG00000080644.11 | 3.595E-10 | 0 | 13063 | gtex_brain_putamen_basal |
| rs139978901 | 15 | 78902026 | CHRNA3 | ENSG00000080644.11 | 9.923E-8 | 0 | 11611 | gtex_brain_putamen_basal |
| rs183822442 | 15 | 78902775 | CHRNA3 | ENSG00000080644.11 | 2.634E-8 | 0 | 10862 | gtex_brain_putamen_basal |
| rs190259891 | 15 | 78902906 | CHRNA3 | ENSG00000080644.11 | 1.287E-7 | 0 | 10731 | gtex_brain_putamen_basal |
| rs77403874 | 15 | 78903165 | CHRNA3 | ENSG00000080644.11 | 1.081E-7 | 0 | 10472 | gtex_brain_putamen_basal |
| rs9646187 | 15 | 78906128 | CHRNA3 | ENSG00000080644.11 | 1.882E-10 | 0 | 7509 | gtex_brain_putamen_basal |
| rs2869546 | 15 | 78907345 | CHRNA3 | ENSG00000080644.11 | 2.265E-11 | 0 | 6292 | gtex_brain_putamen_basal |
| rs3743075 | 15 | 78909452 | CHRNA3 | ENSG00000080644.11 | 8.892E-9 | 0 | 4185 | gtex_brain_putamen_basal |
| rs3743074 | 15 | 78909480 | CHRNA3 | ENSG00000080644.11 | 7.993E-10 | 0 | 4157 | gtex_brain_putamen_basal |
| rs3743073 | 15 | 78909539 | CHRNA3 | ENSG00000080644.11 | 1.716E-9 | 0 | 4098 | gtex_brain_putamen_basal |
| rs2067808 | 15 | 78911780 | CHRNA3 | ENSG00000080644.11 | 6.746E-11 | 0 | 1857 | gtex_brain_putamen_basal |
| rs1878399 | 15 | 78912003 | CHRNA3 | ENSG00000080644.11 | 2.291E-8 | 0 | 1634 | gtex_brain_putamen_basal |
| rs201576627 | 15 | 78912201 | CHRNA3 | ENSG00000080644.11 | 3.817E-7 | 0 | 1436 | gtex_brain_putamen_basal |
| rs4366683 | 15 | 78912203 | CHRNA3 | ENSG00000080644.11 | 3.078E-7 | 0 | 1434 | gtex_brain_putamen_basal |
| rs386419558 | 15 | 78912203 | CHRNA3 | ENSG00000080644.11 | 1.478E-7 | 0 | 1434 | gtex_brain_putamen_basal |
| rs4352017 | 15 | 78912204 | CHRNA3 | ENSG00000080644.11 | 1.343E-7 | 0 | 1433 | gtex_brain_putamen_basal |
| rs8025188 | 15 | 78912278 | CHRNA3 | ENSG00000080644.11 | 2.431E-7 | 0 | 1359 | gtex_brain_putamen_basal |
| rs8023462 | 15 | 78914734 | CHRNA3 | ENSG00000080644.11 | 2.037E-12 | 0 | -1097 | gtex_brain_putamen_basal |
| rs4887070 | 15 | 78916187 | CHRNA3 | ENSG00000080644.11 | 3.896E-11 | 0 | -2550 | gtex_brain_putamen_basal |
| rs2904130 | 15 | 78916622 | CHRNA3 | ENSG00000080644.11 | 1.453E-13 | 0 | -2985 | gtex_brain_putamen_basal |
| rs1948 | 15 | 78917399 | CHRNA3 | ENSG00000080644.11 | 3E-12 | 0 | -3762 | gtex_brain_putamen_basal |
| rs62010332 | 15 | 78918762 | CHRNA3 | ENSG00000080644.11 | 1.342E-13 | 0 | -5125 | gtex_brain_putamen_basal |
| rs7178270 | 15 | 78921077 | CHRNA3 | ENSG00000080644.11 | 7.694E-9 | 0 | -7440 | gtex_brain_putamen_basal |
| rs950776 | 15 | 78926018 | CHRNA3 | ENSG00000080644.11 | 8.286E-10 | 0 | -12381 | gtex_brain_putamen_basal |
| rs79609284 | 15 | 78928411 | CHRNA3 | ENSG00000080644.11 | 3.242E-7 | 0 | -14774 | gtex_brain_putamen_basal |
| rs57728226 | 15 | 78929350 | CHRNA3 | ENSG00000080644.11 | 1.726E-6 | 0 | -15713 | gtex_brain_putamen_basal |
| rs36045869 | 15 | 78929498 | CHRNA3 | ENSG00000080644.11 | 1.726E-6 | 0 | -15861 | gtex_brain_putamen_basal |
| rs11637890 | 15 | 78935419 | CHRNA3 | ENSG00000080644.11 | 1.797E-7 | 0 | -21782 | gtex_brain_putamen_basal |
| rs11633223 | 15 | 78935476 | CHRNA3 | ENSG00000080644.11 | 1.797E-7 | 0 | -21839 | gtex_brain_putamen_basal |
Section II. Transcriptome annotation
General gene expression (GTEx)
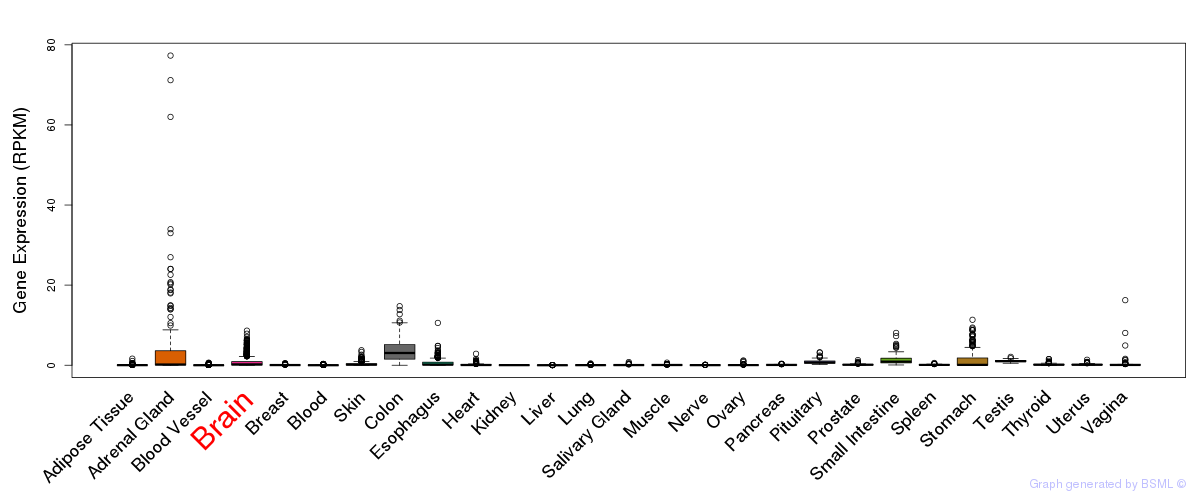
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
No co-expressed genes in brain regions
Section III. Gene Ontology annotation
| Molecular function | GO term | Evidence | Neuro keywords | PubMed ID |
|---|---|---|---|---|
| GO:0030594 | neurotransmitter receptor activity | IEA | Neurotransmitter (GO term level: 5) | - |
| GO:0015464 | acetylcholine receptor activity | IDA | Neurotransmitter (GO term level: 6) | 8906617 |
| GO:0004889 | nicotinic acetylcholine-activated cation-selective channel activity | IDA | 8906617 | |
| GO:0004889 | nicotinic acetylcholine-activated cation-selective channel activity | IEA | - | |
| GO:0005216 | ion channel activity | IEA | - | |
| GO:0005230 | extracellular ligand-gated ion channel activity | IEA | - | |
| Biological process | GO term | Evidence | Neuro keywords | PubMed ID |
| GO:0060084 | synaptic transmission involved in micturition | IEA | neuron, Synap (GO term level: 8) | - |
| GO:0060084 | synaptic transmission involved in micturition | IMP | neuron, Synap (GO term level: 8) | 11450844 |
| GO:0060079 | regulation of excitatory postsynaptic membrane potential | IEA | neuron, Synap (GO term level: 10) | - |
| GO:0060079 | regulation of excitatory postsynaptic membrane potential | ISS | neuron, Synap (GO term level: 10) | - |
| GO:0007271 | synaptic transmission, cholinergic | IEA | neuron, Cholinergic, Synap, Neurotransmitter (GO term level: 7) | - |
| GO:0007271 | synaptic transmission, cholinergic | ISS | neuron, Cholinergic, Synap, Neurotransmitter (GO term level: 7) | - |
| GO:0014056 | regulation of acetylcholine secretion | IEA | Neurotransmitter (GO term level: 10) | - |
| GO:0014056 | regulation of acetylcholine secretion | ISS | Neurotransmitter (GO term level: 10) | - |
| GO:0048814 | regulation of dendrite morphogenesis | IEA | dendrite (GO term level: 13) | - |
| GO:0048814 | regulation of dendrite morphogenesis | ISS | dendrite (GO term level: 13) | - |
| GO:0007399 | nervous system development | IMP | neurite (GO term level: 5) | 11450844 |
| GO:0007165 | signal transduction | IDA | 8906617 | |
| GO:0007626 | locomotory behavior | IEA | - | |
| GO:0007626 | locomotory behavior | ISS | - | |
| GO:0006811 | ion transport | IEA | - | |
| GO:0006811 | ion transport | NAS | 8906617 | |
| GO:0006940 | regulation of smooth muscle contraction | IEA | - | |
| GO:0006940 | regulation of smooth muscle contraction | ISS | - | |
| GO:0007171 | activation of transmembrane receptor protein tyrosine kinase activity | IEA | - | |
| GO:0007171 | activation of transmembrane receptor protein tyrosine kinase activity | ISS | - | |
| GO:0042391 | regulation of membrane potential | IEA | - | |
| GO:0042391 | regulation of membrane potential | ISS | - | |
| GO:0035095 | behavioral response to nicotine | IEA | - | |
| GO:0035095 | behavioral response to nicotine | IMP | 18227835 | |
| Cellular component | GO term | Evidence | Neuro keywords | PubMed ID |
| GO:0005892 | nicotinic acetylcholine-gated receptor-channel complex | IDA | Neurotransmitter (GO term level: 9) | 8906617 |
| GO:0005892 | nicotinic acetylcholine-gated receptor-channel complex | IEA | Neurotransmitter (GO term level: 9) | - |
| GO:0045211 | postsynaptic membrane | IEA | Synap, Neurotransmitter (GO term level: 5) | - |
| GO:0045202 | synapse | IEA | neuron, Synap, Neurotransmitter, Glial (GO term level: 2) | - |
| GO:0016021 | integral to membrane | IEA | - | |
| GO:0016021 | integral to membrane | NAS | 8906617 | |
| GO:0005886 | plasma membrane | IEA | - | |
| GO:0030054 | cell junction | IEA | - |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| KEGG NEUROACTIVE LIGAND RECEPTOR INTERACTION | 272 | 195 | All SZGR 2.0 genes in this pathway |
| REACTOME TRANSMISSION ACROSS CHEMICAL SYNAPSES | 186 | 155 | All SZGR 2.0 genes in this pathway |
| REACTOME NEURONAL SYSTEM | 279 | 221 | All SZGR 2.0 genes in this pathway |
| REACTOME NEUROTRANSMITTER RECEPTOR BINDING AND DOWNSTREAM TRANSMISSION IN THE POSTSYNAPTIC CELL | 137 | 110 | All SZGR 2.0 genes in this pathway |
| REACTOME ACETYLCHOLINE BINDING AND DOWNSTREAM EVENTS | 16 | 13 | All SZGR 2.0 genes in this pathway |
| REACTOME HIGHLY CALCIUM PERMEABLE POSTSYNAPTIC NICOTINIC ACETYLCHOLINE RECEPTORS | 13 | 12 | All SZGR 2.0 genes in this pathway |
| REACTOME PRESYNAPTIC NICOTINIC ACETYLCHOLINE RECEPTORS | 12 | 10 | All SZGR 2.0 genes in this pathway |
| DOANE RESPONSE TO ANDROGEN UP | 184 | 125 | All SZGR 2.0 genes in this pathway |
| GRABARCZYK BCL11B TARGETS DN | 57 | 35 | All SZGR 2.0 genes in this pathway |
| KIM WT1 TARGETS 12HR UP | 162 | 116 | All SZGR 2.0 genes in this pathway |
| DODD NASOPHARYNGEAL CARCINOMA UP | 1821 | 933 | All SZGR 2.0 genes in this pathway |
| CONCANNON APOPTOSIS BY EPOXOMICIN DN | 172 | 112 | All SZGR 2.0 genes in this pathway |
| KANG GIST WITH PDGFRA UP | 50 | 27 | All SZGR 2.0 genes in this pathway |
| BENPORATH SUZ12 TARGETS | 1038 | 678 | All SZGR 2.0 genes in this pathway |
| SHEN SMARCA2 TARGETS DN | 357 | 212 | All SZGR 2.0 genes in this pathway |
| SHIPP DLBCL CURED VS FATAL UP | 39 | 22 | All SZGR 2.0 genes in this pathway |
| KANG IMMORTALIZED BY TERT UP | 89 | 61 | All SZGR 2.0 genes in this pathway |
| RODWELL AGING KIDNEY NO BLOOD UP | 222 | 139 | All SZGR 2.0 genes in this pathway |
| RODWELL AGING KIDNEY UP | 487 | 303 | All SZGR 2.0 genes in this pathway |
| MEISSNER NPC HCP WITH H3 UNMETHYLATED | 536 | 296 | All SZGR 2.0 genes in this pathway |
| MEISSNER BRAIN HCP WITH H3K27ME3 | 269 | 159 | All SZGR 2.0 genes in this pathway |
| MIKKELSEN MCV6 HCP WITH H3K27ME3 | 435 | 318 | All SZGR 2.0 genes in this pathway |
| MIKKELSEN MEF HCP WITH H3K27ME3 | 590 | 403 | All SZGR 2.0 genes in this pathway |