Gene Page: AIRE
Summary ?
| GeneID | 326 |
| Symbol | AIRE |
| Synonyms | AIRE1|APECED|APS1|APSI|PGA1 |
| Description | autoimmune regulator |
| Reference | MIM:607358|HGNC:HGNC:360|Ensembl:ENSG00000160224|HPRD:06301|Vega:OTTHUMG00000086921 |
| Gene type | protein-coding |
| Map location | 21q22.3 |
| Pascal p-value | 0.646 |
| Sherlock p-value | 0.191 |
| Fetal beta | -1.46 |
| eGene | Frontal Cortex BA9 |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:GWASdb | Genome-wide Association Studies | GWASdb records for schizophrenia | |
| CV:PGCnp | Genome-wide Association Study | GWAS |
Section I. Genetics and epigenetics annotation
Section II. Transcriptome annotation
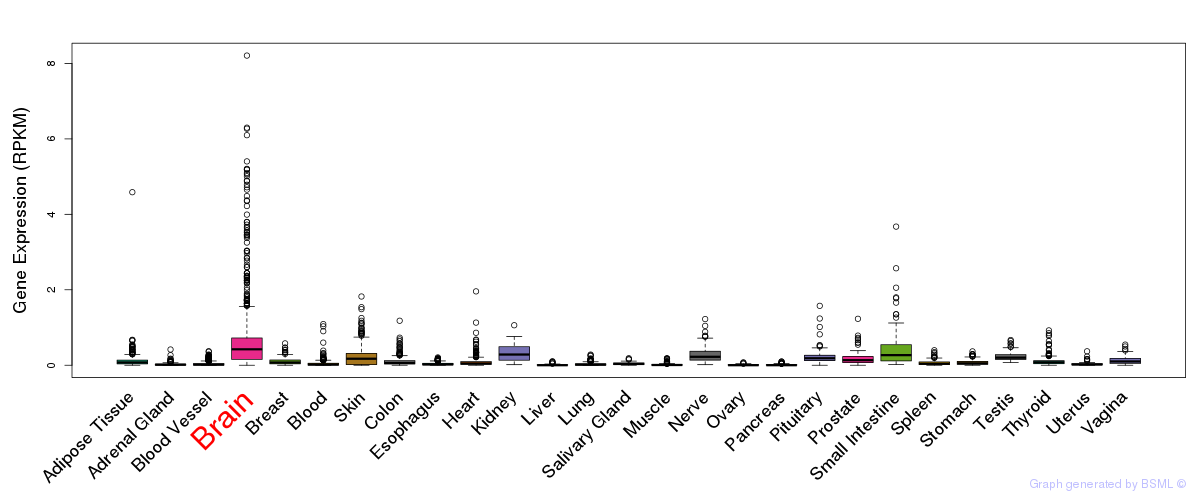
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| XPO1 | 0.93 | 0.95 |
| COPB1 | 0.92 | 0.95 |
| GPBP1 | 0.91 | 0.94 |
| ZNF639 | 0.91 | 0.94 |
| CEBPZ | 0.91 | 0.93 |
| RAD21 | 0.91 | 0.92 |
| LRRC40 | 0.91 | 0.92 |
| SMARCA1 | 0.90 | 0.94 |
| C14orf135 | 0.90 | 0.93 |
| NAP1L4 | 0.90 | 0.93 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| MT-CO2 | -0.74 | -0.86 |
| AF347015.33 | -0.73 | -0.84 |
| FXYD1 | -0.72 | -0.85 |
| MT-CYB | -0.72 | -0.83 |
| AF347015.27 | -0.71 | -0.82 |
| AF347015.31 | -0.71 | -0.83 |
| HLA-F | -0.71 | -0.78 |
| AF347015.8 | -0.71 | -0.84 |
| AF347015.15 | -0.69 | -0.82 |
| IFI27 | -0.68 | -0.82 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| KEGG UBIQUITIN MEDIATED PROTEOLYSIS | 138 | 98 | All SZGR 2.0 genes in this pathway |
| KEGG PRIMARY IMMUNODEFICIENCY | 35 | 28 | All SZGR 2.0 genes in this pathway |
| RIZ ERYTHROID DIFFERENTIATION APOBEC2 | 27 | 19 | All SZGR 2.0 genes in this pathway |
| WANG CISPLATIN RESPONSE AND XPC DN | 228 | 146 | All SZGR 2.0 genes in this pathway |
| HELLER SILENCED BY METHYLATION UP | 282 | 183 | All SZGR 2.0 genes in this pathway |
| MATZUK CENTRAL FOR FEMALE FERTILITY | 29 | 25 | All SZGR 2.0 genes in this pathway |
| LABBE TGFB1 TARGETS UP | 102 | 64 | All SZGR 2.0 genes in this pathway |
| MIKKELSEN MCV6 HCP WITH H3K27ME3 | 435 | 318 | All SZGR 2.0 genes in this pathway |
| MARTENS TRETINOIN RESPONSE UP | 857 | 456 | All SZGR 2.0 genes in this pathway |
| PASINI SUZ12 TARGETS UP | 112 | 65 | All SZGR 2.0 genes in this pathway |