| KEGG PORPHYRIN AND CHLOROPHYLL METABOLISM
| 41 | 23 | All SZGR 2.0 genes in this pathway |
| PID HIF1 TFPATHWAY
| 66 | 52 | All SZGR 2.0 genes in this pathway |
| REACTOME TRANSMEMBRANE TRANSPORT OF SMALL MOLECULES
| 413 | 270 | All SZGR 2.0 genes in this pathway |
| REACTOME SLC MEDIATED TRANSMEMBRANE TRANSPORT
| 241 | 157 | All SZGR 2.0 genes in this pathway |
| REACTOME TRANSPORT OF GLUCOSE AND OTHER SUGARS BILE SALTS AND ORGANIC ACIDS METAL IONS AND AMINE COMPOUNDS
| 89 | 54 | All SZGR 2.0 genes in this pathway |
| REACTOME METAL ION SLC TRANSPORTERS
| 22 | 13 | All SZGR 2.0 genes in this pathway |
| REACTOME IRON UPTAKE AND TRANSPORT
| 36 | 18 | All SZGR 2.0 genes in this pathway |
| PARENT MTOR SIGNALING UP
| 567 | 375 | All SZGR 2.0 genes in this pathway |
| NAKAMURA TUMOR ZONE PERIPHERAL VS CENTRAL DN
| 634 | 384 | All SZGR 2.0 genes in this pathway |
| PICCALUGA ANGIOIMMUNOBLASTIC LYMPHOMA UP
| 205 | 140 | All SZGR 2.0 genes in this pathway |
| SENGUPTA NASOPHARYNGEAL CARCINOMA WITH LMP1 UP
| 408 | 247 | All SZGR 2.0 genes in this pathway |
| TURASHVILI BREAST DUCTAL CARCINOMA VS DUCTAL NORMAL DN
| 198 | 110 | All SZGR 2.0 genes in this pathway |
| TURASHVILI BREAST LOBULAR CARCINOMA VS DUCTAL NORMAL DN
| 91 | 53 | All SZGR 2.0 genes in this pathway |
| THUM SYSTOLIC HEART FAILURE UP
| 423 | 283 | All SZGR 2.0 genes in this pathway |
| ODONNELL TFRC TARGETS UP
| 456 | 228 | All SZGR 2.0 genes in this pathway |
| SABATES COLORECTAL ADENOMA DN
| 291 | 176 | All SZGR 2.0 genes in this pathway |
| KIM WT1 TARGETS 8HR DN
| 129 | 84 | All SZGR 2.0 genes in this pathway |
| DODD NASOPHARYNGEAL CARCINOMA UP
| 1821 | 933 | All SZGR 2.0 genes in this pathway |
| GOZGIT ESR1 TARGETS DN
| 781 | 465 | All SZGR 2.0 genes in this pathway |
| BERENJENO ROCK SIGNALING NOT VIA RHOA UP
| 29 | 21 | All SZGR 2.0 genes in this pathway |
| MARKEY RB1 CHRONIC LOF DN
| 118 | 78 | All SZGR 2.0 genes in this pathway |
| GRAESSMANN APOPTOSIS BY SERUM DEPRIVATION UP
| 552 | 347 | All SZGR 2.0 genes in this pathway |
| GRAESSMANN RESPONSE TO MC AND SERUM DEPRIVATION UP
| 211 | 136 | All SZGR 2.0 genes in this pathway |
| GAUSSMANN MLL AF4 FUSION TARGETS E UP
| 97 | 60 | All SZGR 2.0 genes in this pathway |
| MCBRYAN PUBERTAL BREAST 4 5WK DN
| 196 | 131 | All SZGR 2.0 genes in this pathway |
| MCBRYAN PUBERTAL BREAST 6 7WK UP
| 197 | 135 | All SZGR 2.0 genes in this pathway |
| MCBRYAN PUBERTAL TGFB1 TARGETS UP
| 169 | 127 | All SZGR 2.0 genes in this pathway |
| DARWICHE SKIN TUMOR PROMOTER DN
| 185 | 115 | All SZGR 2.0 genes in this pathway |
| DARWICHE PAPILLOMA RISK LOW DN
| 165 | 107 | All SZGR 2.0 genes in this pathway |
| DARWICHE PAPILLOMA RISK HIGH DN
| 180 | 110 | All SZGR 2.0 genes in this pathway |
| DARWICHE SQUAMOUS CELL CARCINOMA DN
| 181 | 107 | All SZGR 2.0 genes in this pathway |
| KHETCHOUMIAN TRIM24 TARGETS UP
| 47 | 38 | All SZGR 2.0 genes in this pathway |
| INGRAM SHH TARGETS UP
| 127 | 79 | All SZGR 2.0 genes in this pathway |
| MANTOVANI NFKB TARGETS UP
| 43 | 33 | All SZGR 2.0 genes in this pathway |
| MANTOVANI VIRAL GPCR SIGNALING UP
| 86 | 54 | All SZGR 2.0 genes in this pathway |
| RICKMAN HEAD AND NECK CANCER E
| 89 | 44 | All SZGR 2.0 genes in this pathway |
| HOUSTIS ROS
| 36 | 29 | All SZGR 2.0 genes in this pathway |
| VANTVEER BREAST CANCER METASTASIS DN
| 121 | 65 | All SZGR 2.0 genes in this pathway |
| BYSTRYKH HEMATOPOIESIS STEM CELL QTL TRANS
| 882 | 572 | All SZGR 2.0 genes in this pathway |
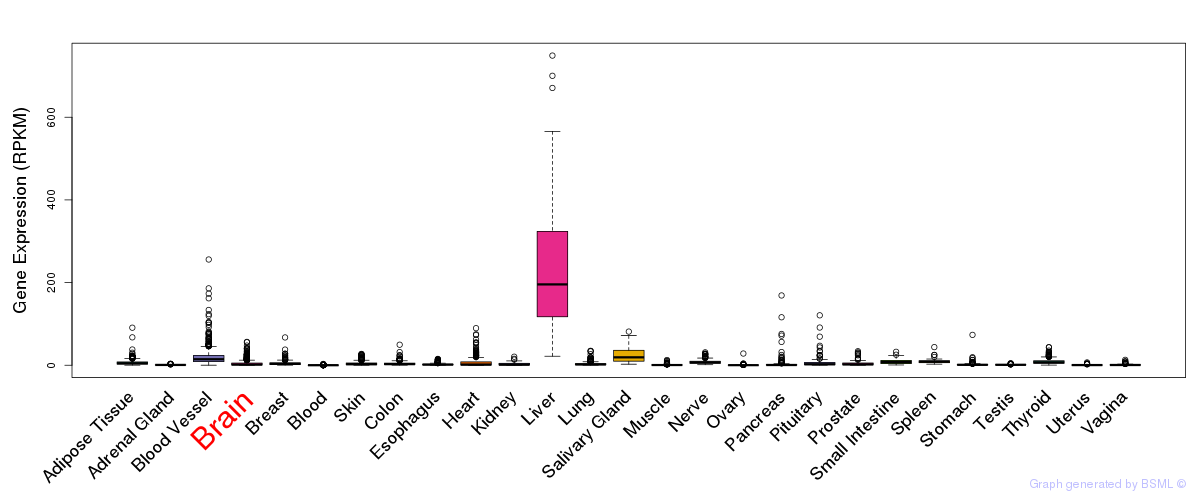
| HSIAO LIVER SPECIFIC GENES
| 244 | 154 | All SZGR 2.0 genes in this pathway |
| LEE AGING NEOCORTEX DN
| 80 | 49 | All SZGR 2.0 genes in this pathway |
| RUAN RESPONSE TO TNF UP
| 12 | 10 | All SZGR 2.0 genes in this pathway |
| HARRIS HYPOXIA
| 81 | 64 | All SZGR 2.0 genes in this pathway |
| BLALOCK ALZHEIMERS DISEASE UP
| 1691 | 1088 | All SZGR 2.0 genes in this pathway |
| RUAN RESPONSE TO TNF TROGLITAZONE UP
| 17 | 11 | All SZGR 2.0 genes in this pathway |
| CHANG IMMORTALIZED BY HPV31 UP
| 84 | 55 | All SZGR 2.0 genes in this pathway |
| SEMENZA HIF1 TARGETS
| 36 | 32 | All SZGR 2.0 genes in this pathway |
| CHENG RESPONSE TO NICKEL ACETATE
| 45 | 29 | All SZGR 2.0 genes in this pathway |
| ZAMORA NOS2 TARGETS DN
| 96 | 71 | All SZGR 2.0 genes in this pathway |
| HAN JNK SINGALING DN
| 39 | 27 | All SZGR 2.0 genes in this pathway |
| ZHONG SECRETOME OF LUNG CANCER AND MACROPHAGE
| 77 | 50 | All SZGR 2.0 genes in this pathway |
| ZHONG SECRETOME OF LUNG CANCER AND ENDOTHELIUM
| 66 | 47 | All SZGR 2.0 genes in this pathway |
| ZHONG SECRETOME OF LUNG CANCER AND FIBROBLAST
| 132 | 93 | All SZGR 2.0 genes in this pathway |
| MONNIER POSTRADIATION TUMOR ESCAPE DN
| 373 | 196 | All SZGR 2.0 genes in this pathway |
| CREIGHTON ENDOCRINE THERAPY RESISTANCE 3
| 720 | 440 | All SZGR 2.0 genes in this pathway |
| FOSTER TOLERANT MACROPHAGE UP
| 156 | 92 | All SZGR 2.0 genes in this pathway |
| FOSTER TOLERANT MACROPHAGE DN
| 409 | 268 | All SZGR 2.0 genes in this pathway |
| KONDO EZH2 TARGETS
| 245 | 148 | All SZGR 2.0 genes in this pathway |
| WONG MITOCHONDRIA GENE MODULE
| 217 | 122 | All SZGR 2.0 genes in this pathway |
| MARTINEZ RB1 TARGETS UP
| 673 | 430 | All SZGR 2.0 genes in this pathway |
| MARTINEZ TP53 TARGETS UP
| 602 | 364 | All SZGR 2.0 genes in this pathway |
| MARTINEZ RB1 AND TP53 TARGETS UP
| 601 | 369 | All SZGR 2.0 genes in this pathway |
| IWANAGA CARCINOGENESIS BY KRAS PTEN UP
| 181 | 112 | All SZGR 2.0 genes in this pathway |
| SMID BREAST CANCER RELAPSE IN BRAIN UP
| 39 | 20 | All SZGR 2.0 genes in this pathway |
| SMID BREAST CANCER RELAPSE IN BONE DN
| 315 | 197 | All SZGR 2.0 genes in this pathway |
| SMID BREAST CANCER RELAPSE IN PLEURA DN
| 27 | 15 | All SZGR 2.0 genes in this pathway |
| SMID BREAST CANCER BASAL UP
| 648 | 398 | All SZGR 2.0 genes in this pathway |
| BONOME OVARIAN CANCER SURVIVAL OPTIMAL DEBULKING
| 246 | 152 | All SZGR 2.0 genes in this pathway |
| ZHENG GLIOBLASTOMA PLASTICITY UP
| 250 | 168 | All SZGR 2.0 genes in this pathway |
| LABBE TGFB1 TARGETS DN
| 108 | 64 | All SZGR 2.0 genes in this pathway |
| LABBE TARGETS OF TGFB1 AND WNT3A DN
| 108 | 68 | All SZGR 2.0 genes in this pathway |
| WINTER HYPOXIA METAGENE
| 242 | 168 | All SZGR 2.0 genes in this pathway |
| WANG TUMOR INVASIVENESS DN
| 210 | 128 | All SZGR 2.0 genes in this pathway |
| BOYLAN MULTIPLE MYELOMA C D DN
| 252 | 155 | All SZGR 2.0 genes in this pathway |
| LUDWICZEK TREATING IRON OVERLOAD
| 7 | 5 | All SZGR 2.0 genes in this pathway |
| SWEET LUNG CANCER KRAS DN
| 435 | 289 | All SZGR 2.0 genes in this pathway |
| ZHANG TLX TARGETS 60HR UP
| 293 | 203 | All SZGR 2.0 genes in this pathway |
| COLINA TARGETS OF 4EBP1 AND 4EBP2
| 356 | 214 | All SZGR 2.0 genes in this pathway |
| CHIANG LIVER CANCER SUBCLASS CTNNB1 DN
| 170 | 105 | All SZGR 2.0 genes in this pathway |
| CHIANG LIVER CANCER SUBCLASS UNANNOTATED UP
| 85 | 50 | All SZGR 2.0 genes in this pathway |
| MIKKELSEN MEF LCP WITH H3K4ME3
| 128 | 68 | All SZGR 2.0 genes in this pathway |
| BOYLAN MULTIPLE MYELOMA PCA1 UP
| 101 | 66 | All SZGR 2.0 genes in this pathway |
| HOSHIDA LIVER CANCER SUBCLASS S3
| 266 | 180 | All SZGR 2.0 genes in this pathway |
| CAIRO HEPATOBLASTOMA DN
| 267 | 160 | All SZGR 2.0 genes in this pathway |
| MIKKELSEN NPC LCP WITH H3K4ME3
| 58 | 34 | All SZGR 2.0 genes in this pathway |
| STAMBOLSKY RESPONSE TO VITAMIN D3 UP
| 84 | 48 | All SZGR 2.0 genes in this pathway |
| HOELZEL NF1 TARGETS DN
| 115 | 73 | All SZGR 2.0 genes in this pathway |
| STEGER ADIPOGENESIS UP
| 21 | 16 | All SZGR 2.0 genes in this pathway |
| VANDESLUIS COMMD1 TARGETS GROUP 3 DN
| 39 | 21 | All SZGR 2.0 genes in this pathway |
| LEE BMP2 TARGETS UP
| 745 | 475 | All SZGR 2.0 genes in this pathway |
| WACKER HYPOXIA TARGETS OF VHL
| 13 | 11 | All SZGR 2.0 genes in this pathway |
| YANG BCL3 TARGETS UP
| 364 | 236 | All SZGR 2.0 genes in this pathway |
| DELACROIX RARG BOUND MEF
| 367 | 231 | All SZGR 2.0 genes in this pathway |
| DELACROIX RAR TARGETS UP
| 48 | 33 | All SZGR 2.0 genes in this pathway |
| KRIEG HYPOXIA NOT VIA KDM3A
| 770 | 480 | All SZGR 2.0 genes in this pathway |
| ACEVEDO FGFR1 TARGETS IN PROSTATE CANCER MODEL UP
| 289 | 184 | All SZGR 2.0 genes in this pathway |
 eQTL annotation
eQTL annotation