Gene Page: ASF1A
Summary ?
| GeneID | 25842 |
| Symbol | ASF1A |
| Synonyms | CGI-98|CIA|HSPC146 |
| Description | anti-silencing function 1A histone chaperone |
| Reference | MIM:609189|HGNC:HGNC:20995|Ensembl:ENSG00000111875|HPRD:16459|Vega:OTTHUMG00000015470 |
| Gene type | protein-coding |
| Map location | 6q22.31 |
| Pascal p-value | 0.001 |
| Sherlock p-value | 0.005 |
| Support | Chromatin Remodeling Genes |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS |
Section I. Genetics and epigenetics annotation
Section II. Transcriptome annotation
General gene expression (GTEx)
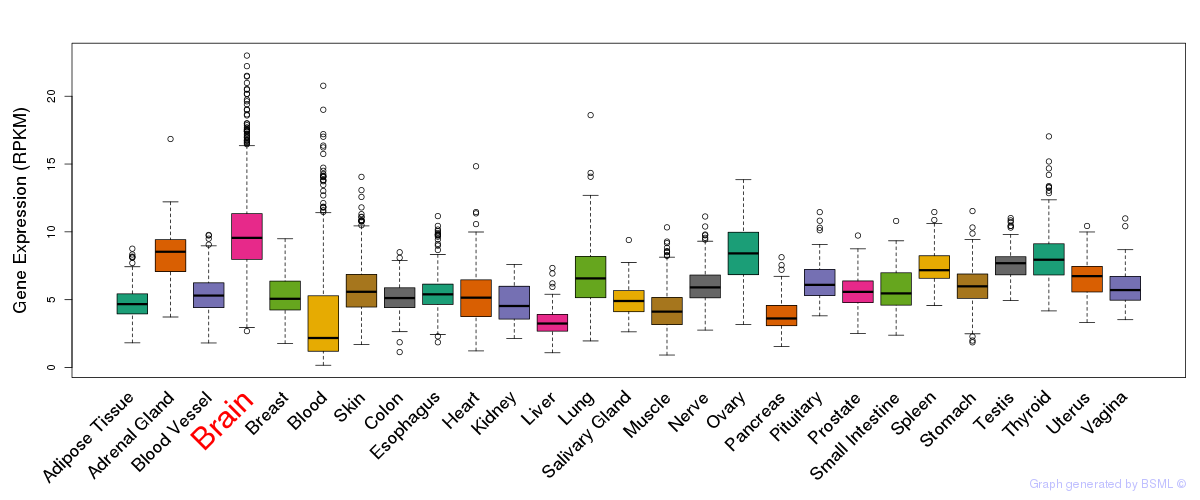
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| SART1 | 0.75 | 0.71 |
| MIB2 | 0.73 | 0.74 |
| CDC37 | 0.72 | 0.70 |
| OGFR | 0.72 | 0.70 |
| HIRIP3 | 0.72 | 0.68 |
| PPAN | 0.71 | 0.69 |
| C9orf86 | 0.70 | 0.69 |
| KRI1 | 0.70 | 0.69 |
| MBD3 | 0.70 | 0.66 |
| PELP1 | 0.69 | 0.66 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| C1orf54 | -0.56 | -0.51 |
| CLEC2B | -0.56 | -0.50 |
| LY96 | -0.54 | -0.46 |
| LPAR6 | -0.53 | -0.52 |
| B2M | -0.52 | -0.47 |
| SCRG1 | -0.52 | -0.39 |
| HBD | -0.52 | -0.48 |
| SYCP3 | -0.51 | -0.45 |
| HBB | -0.51 | -0.48 |
| PXMP3 | -0.51 | -0.50 |
Section IV. Protein-protein interaction annotation
| Interactors | Aliases B | Official full name B | Experimental | Source | PubMed ID |
|---|---|---|---|---|---|
| ASF1B | CIA-II | FLJ10604 | ASF1 anti-silencing function 1 homolog B (S. cerevisiae) | - | HPRD | 12842904 |
| CHAF1A | CAF-1 | CAF1 | CAF1B | CAF1P150 | MGC71229 | P150 | chromatin assembly factor 1, subunit A (p150) | Affinity Capture-Western Reconstituted Complex | BioGRID | 11897662 |
| CHAF1B | CAF-1 | CAF-IP60 | CAF1 | CAF1A | CAF1P60 | MPHOSPH7 | MPP7 | chromatin assembly factor 1, subunit B (p60) | Affinity Capture-Western Reconstituted Complex | BioGRID | 11897662 |
| H3F3A | H3.3A | H3F3 | MGC87782 | MGC87783 | H3 histone, family 3A | H3.3 interacts with Asf1a. This interaction was modeled on a demonstrated interaction between H3.3 from an unspecified species and human Asf1a. | BIND | 15664198 |
| HIRA | DGCR1 | TUP1 | TUPLE1 | HIR histone cell cycle regulation defective homolog A (S. cerevisiae) | HIRA interacts with ASF1a. | BIND | 15621527 |
| HIST1H3C | H3.1 | H3/c | H3FC | histone cluster 1, H3c | - | HPRD | 10759893 |
| HIST1H3E | H3.1 | H3/d | H3FD | histone cluster 1, H3e | H3.1 interacts with Asf1a. This interaction was modeled on a demonstrated interaction between H3.1 from an unspecified species and human Asf1a. | BIND | 15664198 |
| HIST1H4F | H4 | H4/c | H4FC | histone cluster 1, H4f | - | HPRD | 10759893 |
| HIST3H3 | H3.4 | H3/g | H3FT | H3t | MGC126886 | MGC126888 | histone cluster 3, H3 | H3 interacts with Asf1a. | BIND | 15664198 |
| HIST4H4 | H4/p | MGC24116 | histone cluster 4, H4 | Asf1a interacts with H4AcK12. This interaction was modeled on a demonstrated interaction between Asf1a from an unspecified species and human H4AcK12. | BIND | 15664198 |
| NASP | DKFZp547F162 | FLB7527 | FLJ31599 | FLJ35510 | MGC19722 | MGC20372 | MGC2297 | PRO1999 | nuclear autoantigenic sperm protein (histone-binding) | Asf1a interacts with NASP. This interaction was modeled on a demonstrated interaction between Asf1a from an unspecified species and human NASP. | BIND | 15664198 |
| ST13 | AAG2 | FAM10A1 | FAM10A4 | FLJ27260 | HIP | HOP | HSPABP | HSPABP1 | MGC129952 | P48 | PRO0786 | SNC6 | suppression of tumorigenicity 13 (colon carcinoma) (Hsp70 interacting protein) | Asf1a interacts with p48. This interaction was modeled on a demonstrated interaction between Asf1a from an unspecified species and human p48. | BIND | 15664198 |
| TAF1 | BA2R | CCG1 | CCGS | DYT3 | KAT4 | N-TAF1 | NSCL2 | OF | P250 | TAF2A | TAFII250 | TAF1 RNA polymerase II, TATA box binding protein (TBP)-associated factor, 250kDa | - | HPRD | 12093919 |
| TAF1 | BA2R | CCG1 | CCGS | DYT3 | KAT4 | N-TAF1 | NSCL2 | OF | P250 | TAF2A | TAFII250 | TAF1 RNA polymerase II, TATA box binding protein (TBP)-associated factor, 250kDa | CIA interacts with TAFII250. | BIND | 12093919 |
| TLK1 | KIAA0137 | PKU-beta | tousled-like kinase 1 | - | HPRD,BioGRID | 11470414 |
| TLK2 | MGC44450 | PKU-ALPHA | tousled-like kinase 2 | - | HPRD,BioGRID | 11470414 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| ONKEN UVEAL MELANOMA UP | 783 | 507 | All SZGR 2.0 genes in this pathway |
| GAZDA DIAMOND BLACKFAN ANEMIA ERYTHROID DN | 493 | 298 | All SZGR 2.0 genes in this pathway |
| HORIUCHI WTAP TARGETS DN | 310 | 188 | All SZGR 2.0 genes in this pathway |
| WANG CLIM2 TARGETS DN | 186 | 114 | All SZGR 2.0 genes in this pathway |
| KINSEY TARGETS OF EWSR1 FLII FUSION UP | 1278 | 748 | All SZGR 2.0 genes in this pathway |
| SCIBETTA KDM5B TARGETS DN | 81 | 55 | All SZGR 2.0 genes in this pathway |
| KIM WT1 TARGETS DN | 459 | 276 | All SZGR 2.0 genes in this pathway |
| GRAHAM CML QUIESCENT VS NORMAL QUIESCENT UP | 87 | 45 | All SZGR 2.0 genes in this pathway |
| MATTIOLI MGUS VS PCL | 116 | 62 | All SZGR 2.0 genes in this pathway |
| FARMER BREAST CANCER BASAL VS LULMINAL | 330 | 215 | All SZGR 2.0 genes in this pathway |
| PUJANA XPRSS INT NETWORK | 168 | 103 | All SZGR 2.0 genes in this pathway |
| PUJANA BRCA1 PCC NETWORK | 1652 | 1023 | All SZGR 2.0 genes in this pathway |
| PUJANA BRCA2 PCC NETWORK | 423 | 265 | All SZGR 2.0 genes in this pathway |
| PUJANA ATM PCC NETWORK | 1442 | 892 | All SZGR 2.0 genes in this pathway |
| PUJANA CHEK2 PCC NETWORK | 779 | 480 | All SZGR 2.0 genes in this pathway |
| PUJANA BRCA CENTERED NETWORK | 117 | 72 | All SZGR 2.0 genes in this pathway |
| BENPORATH PROLIFERATION | 147 | 80 | All SZGR 2.0 genes in this pathway |
| PENG GLUCOSE DEPRIVATION DN | 169 | 112 | All SZGR 2.0 genes in this pathway |
| FLECHNER BIOPSY KIDNEY TRANSPLANT OK VS DONOR UP | 555 | 346 | All SZGR 2.0 genes in this pathway |
| MOREAUX B LYMPHOCYTE MATURATION BY TACI DN | 73 | 45 | All SZGR 2.0 genes in this pathway |
| MOREAUX MULTIPLE MYELOMA BY TACI DN | 172 | 107 | All SZGR 2.0 genes in this pathway |
| BLALOCK ALZHEIMERS DISEASE DN | 1237 | 837 | All SZGR 2.0 genes in this pathway |
| MARSON BOUND BY FOXP3 STIMULATED | 1022 | 619 | All SZGR 2.0 genes in this pathway |
| MARSON BOUND BY FOXP3 UNSTIMULATED | 1229 | 713 | All SZGR 2.0 genes in this pathway |
| GAZIN EPIGENETIC SILENCING BY KRAS | 26 | 16 | All SZGR 2.0 genes in this pathway |
| ACEVEDO LIVER CANCER UP | 973 | 570 | All SZGR 2.0 genes in this pathway |
| PODAR RESPONSE TO ADAPHOSTIN UP | 147 | 98 | All SZGR 2.0 genes in this pathway |
| AGUIRRE PANCREATIC CANCER COPY NUMBER DN | 238 | 145 | All SZGR 2.0 genes in this pathway |
| YAGI AML WITH T 8 21 TRANSLOCATION | 368 | 247 | All SZGR 2.0 genes in this pathway |
| PUJANA BREAST CANCER WITH BRCA1 MUTATED UP | 56 | 27 | All SZGR 2.0 genes in this pathway |
| HIRSCH CELLULAR TRANSFORMATION SIGNATURE DN | 103 | 67 | All SZGR 2.0 genes in this pathway |
| RAO BOUND BY SALL4 ISOFORM B | 517 | 302 | All SZGR 2.0 genes in this pathway |