Gene Page: TBC1D2
Summary ?
| GeneID | 55357 |
| Symbol | TBC1D2 |
| Synonyms | PARIS-1|PARIS1|TBC1D2A |
| Description | TBC1 domain family member 2 |
| Reference | MIM:609871|HGNC:HGNC:18026|Ensembl:ENSG00000095383|Vega:OTTHUMG00000020343 |
| Gene type | protein-coding |
| Map location | 9q22.33 |
| Pascal p-value | 0.004 |
| Fetal beta | -0.138 |
| eGene | Caudate basal ganglia Cerebellar Hemisphere Cerebellum Cortex Hippocampus Hypothalamus Nucleus accumbens basal ganglia Putamen basal ganglia Meta |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| PMID:cooccur | High-throughput literature-search | Systematic search in PubMed for genes co-occurring with SCZ keywords. A total of 3027 genes were included. | |
| Literature | High-throughput literature-search | Co-occurance with Schizophrenia keywords: schizophrenia,schizophrenias | Click to show details |
Section I. Genetics and epigenetics annotation
 eQTL annotation
eQTL annotation
| SNP ID | Chromosome | Position | eGene | Gene Entrez ID | pvalue | qvalue | TSS distance | eQTL type |
|---|---|---|---|---|---|---|---|---|
| rs10985366 | 9 | 100954077 | TBC1D2 | ENSG00000095383.15 | 1.559E-6 | 0.01 | 63838 | gtex_brain_putamen_basal |
| rs3780450 | 9 | 100965008 | TBC1D2 | ENSG00000095383.15 | 1.14E-6 | 0.01 | 52907 | gtex_brain_putamen_basal |
| rs3780449 | 9 | 100965307 | TBC1D2 | ENSG00000095383.15 | 1.316E-6 | 0.01 | 52608 | gtex_brain_putamen_basal |
| rs3739667 | 9 | 100965472 | TBC1D2 | ENSG00000095383.15 | 2.42E-6 | 0.01 | 52443 | gtex_brain_putamen_basal |
| rs10985441 | 9 | 100970881 | TBC1D2 | ENSG00000095383.15 | 1.134E-6 | 0.01 | 47034 | gtex_brain_putamen_basal |
| rs3739666 | 9 | 100971348 | TBC1D2 | ENSG00000095383.15 | 1.134E-6 | 0.01 | 46567 | gtex_brain_putamen_basal |
| rs2023361 | 9 | 100972162 | TBC1D2 | ENSG00000095383.15 | 1.237E-6 | 0.01 | 45753 | gtex_brain_putamen_basal |
| rs733040 | 9 | 100972388 | TBC1D2 | ENSG00000095383.15 | 4.989E-7 | 0.01 | 45527 | gtex_brain_putamen_basal |
| rs942166 | 9 | 100972699 | TBC1D2 | ENSG00000095383.15 | 1.628E-6 | 0.01 | 45216 | gtex_brain_putamen_basal |
| rs4743186 | 9 | 100976086 | TBC1D2 | ENSG00000095383.15 | 1.628E-6 | 0.01 | 41829 | gtex_brain_putamen_basal |
| rs2417893 | 9 | 100977392 | TBC1D2 | ENSG00000095383.15 | 1.628E-6 | 0.01 | 40523 | gtex_brain_putamen_basal |
| rs2900463 | 9 | 100977457 | TBC1D2 | ENSG00000095383.15 | 1.628E-6 | 0.01 | 40458 | gtex_brain_putamen_basal |
| rs7021760 | 9 | 100978294 | TBC1D2 | ENSG00000095383.15 | 1.628E-6 | 0.01 | 39621 | gtex_brain_putamen_basal |
| rs3739665 | 9 | 100979343 | TBC1D2 | ENSG00000095383.15 | 1.628E-6 | 0.01 | 38572 | gtex_brain_putamen_basal |
| rs3739664 | 9 | 100979345 | TBC1D2 | ENSG00000095383.15 | 1.628E-6 | 0.01 | 38570 | gtex_brain_putamen_basal |
| rs7855855 | 9 | 100983566 | TBC1D2 | ENSG00000095383.15 | 6.517E-7 | 0.01 | 34349 | gtex_brain_putamen_basal |
Section II. Transcriptome annotation
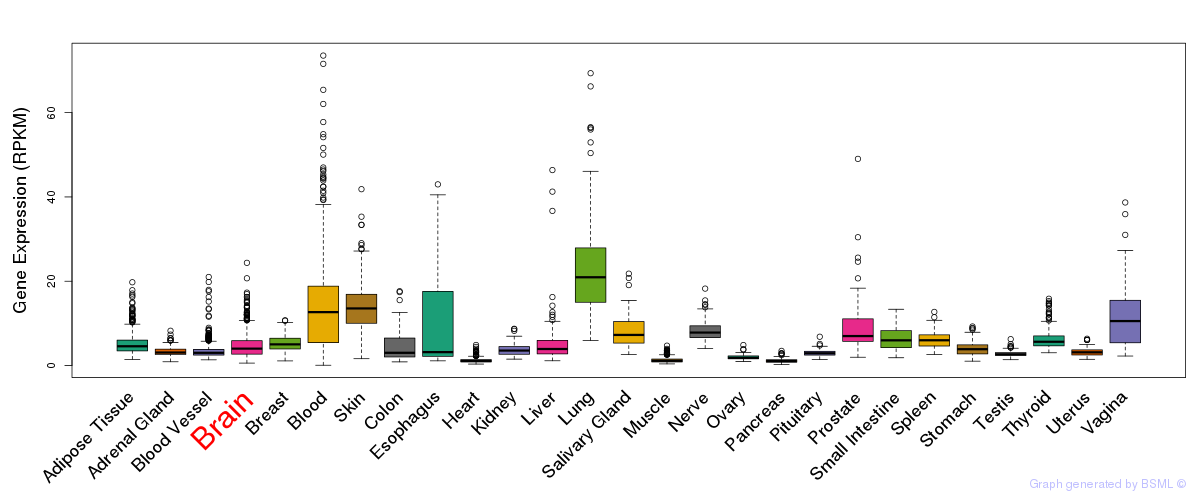
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
No co-expressed genes in brain regions
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| FULCHER INFLAMMATORY RESPONSE LECTIN VS LPS DN | 463 | 290 | All SZGR 2.0 genes in this pathway |
| ZHOU INFLAMMATORY RESPONSE LIVE DN | 384 | 220 | All SZGR 2.0 genes in this pathway |
| KIM WT1 TARGETS 8HR UP | 164 | 122 | All SZGR 2.0 genes in this pathway |
| KIM WT1 TARGETS 12HR UP | 162 | 116 | All SZGR 2.0 genes in this pathway |
| GOZGIT ESR1 TARGETS DN | 781 | 465 | All SZGR 2.0 genes in this pathway |
| BUYTAERT PHOTODYNAMIC THERAPY STRESS DN | 637 | 377 | All SZGR 2.0 genes in this pathway |
| RICKMAN METASTASIS DN | 261 | 155 | All SZGR 2.0 genes in this pathway |
| KOYAMA SEMA3B TARGETS UP | 292 | 168 | All SZGR 2.0 genes in this pathway |
| AMIT EGF RESPONSE 480 HELA | 164 | 118 | All SZGR 2.0 genes in this pathway |
| BLALOCK ALZHEIMERS DISEASE UP | 1691 | 1088 | All SZGR 2.0 genes in this pathway |
| CREIGHTON ENDOCRINE THERAPY RESISTANCE 3 | 720 | 440 | All SZGR 2.0 genes in this pathway |
| ACEVEDO LIVER CANCER UP | 973 | 570 | All SZGR 2.0 genes in this pathway |
| ACEVEDO LIVER TUMOR VS NORMAL ADJACENT TISSUE UP | 863 | 514 | All SZGR 2.0 genes in this pathway |
| BERNARD PPAPDC1B TARGETS DN | 58 | 39 | All SZGR 2.0 genes in this pathway |
| TAYLOR METHYLATED IN ACUTE LYMPHOBLASTIC LEUKEMIA | 77 | 52 | All SZGR 2.0 genes in this pathway |
| KOINUMA TARGETS OF SMAD2 OR SMAD3 | 824 | 528 | All SZGR 2.0 genes in this pathway |
| FOSTER KDM1A TARGETS DN | 211 | 119 | All SZGR 2.0 genes in this pathway |
| ZWANG CLASS 1 TRANSIENTLY INDUCED BY EGF | 516 | 308 | All SZGR 2.0 genes in this pathway |