Gene Page: TRAPPC1
Summary ?
| GeneID | 58485 |
| Symbol | TRAPPC1 |
| Synonyms | BET5|MUM2 |
| Description | trafficking protein particle complex 1 |
| Reference | MIM:610969|HGNC:HGNC:19894|Ensembl:ENSG00000170043|HPRD:18219|Vega:OTTHUMG00000108171 |
| Gene type | protein-coding |
| Map location | 17p13.1 |
| Pascal p-value | 0.013 |
| Sherlock p-value | 0.005 |
| Fetal beta | -0.567 |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS |
Section I. Genetics and epigenetics annotation
Section II. Transcriptome annotation
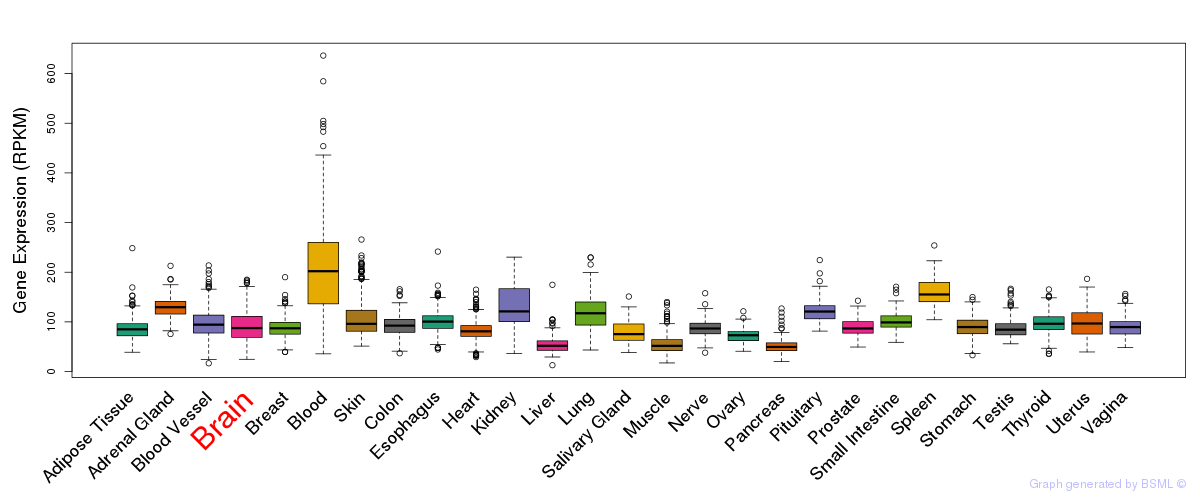
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
Top co-expressed genes in brain regions
| Top 10 positively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| PRDX6 | 0.85 | 0.77 |
| RLBP1 | 0.84 | 0.70 |
| LGALS3 | 0.83 | 0.72 |
| NQO1 | 0.83 | 0.79 |
| BAALC | 0.81 | 0.72 |
| GJB6 | 0.80 | 0.69 |
| PSMB8 | 0.80 | 0.71 |
| SAT1 | 0.80 | 0.70 |
| TLCD1 | 0.80 | 0.76 |
| NDRG2 | 0.80 | 0.74 |
| Top 10 negatively co-expressed genes | ||
| Gene | Pearson's Correlation | Spearman's Correlation |
| UPF2 | -0.64 | -0.68 |
| ZNF407 | -0.63 | -0.71 |
| RTF1 | -0.63 | -0.71 |
| SUPT6H | -0.63 | -0.70 |
| BRD4 | -0.63 | -0.72 |
| NCOR1 | -0.62 | -0.69 |
| GIGYF2 | -0.62 | -0.69 |
| ANKRD17 | -0.62 | -0.67 |
| TCOF1 | -0.62 | -0.67 |
| C14orf102 | -0.62 | -0.68 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| CHANDRAN METASTASIS TOP50 DN | 45 | 26 | All SZGR 2.0 genes in this pathway |
| LASTOWSKA NEUROBLASTOMA COPY NUMBER UP | 181 | 108 | All SZGR 2.0 genes in this pathway |
| RICKMAN METASTASIS DN | 261 | 155 | All SZGR 2.0 genes in this pathway |
| NOUZOVA TRETINOIN AND H4 ACETYLATION | 143 | 85 | All SZGR 2.0 genes in this pathway |
| JIANG HYPOXIA CANCER | 83 | 52 | All SZGR 2.0 genes in this pathway |
| PELLICCIOTTA HDAC IN ANTIGEN PRESENTATION UP | 64 | 40 | All SZGR 2.0 genes in this pathway |
| MARSON BOUND BY FOXP3 UNSTIMULATED | 1229 | 713 | All SZGR 2.0 genes in this pathway |
| GOLDRATH ANTIGEN RESPONSE | 346 | 192 | All SZGR 2.0 genes in this pathway |
| CHANDRAN METASTASIS DN | 306 | 191 | All SZGR 2.0 genes in this pathway |
| CHICAS RB1 TARGETS CONFLUENT | 567 | 365 | All SZGR 2.0 genes in this pathway |
| FIGUEROA AML METHYLATION CLUSTER 3 UP | 170 | 97 | All SZGR 2.0 genes in this pathway |
| FIGUEROA AML METHYLATION CLUSTER 4 UP | 112 | 64 | All SZGR 2.0 genes in this pathway |
| FIGUEROA AML METHYLATION CLUSTER 6 UP | 140 | 81 | All SZGR 2.0 genes in this pathway |
| FIGUEROA AML METHYLATION CLUSTER 7 UP | 118 | 68 | All SZGR 2.0 genes in this pathway |
| JOHNSTONE PARVB TARGETS 2 UP | 140 | 94 | All SZGR 2.0 genes in this pathway |