Gene Page: FIGNL1
Summary ?
| GeneID | 63979 |
| Symbol | FIGNL1 |
| Synonyms | - |
| Description | fidgetin like 1 |
| Reference | MIM:615383|HGNC:HGNC:13286|Ensembl:ENSG00000132436|HPRD:10958|Vega:OTTHUMG00000022866 |
| Gene type | protein-coding |
| Map location | 7p12.1 |
| Pascal p-value | 0.054 |
| Fetal beta | 1.427 |
| DMG | 1 (# studies) |
| eGene | Anterior cingulate cortex BA24 Caudate basal ganglia Cerebellar Hemisphere Cerebellum Cortex Frontal Cortex BA9 Hippocampus Hypothalamus Nucleus accumbens basal ganglia Putamen basal ganglia Myers' cis & trans Meta |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| DMG:Wockner_2014 | Genome-wide DNA methylation analysis | This dataset includes 4641 differentially methylated probes corresponding to 2929 unique genes between schizophrenia patients (n=24) and controls (n=24). | 1 |
Section I. Genetics and epigenetics annotation
 Differentially methylated gene
Differentially methylated gene
| Probe | Chromosome | Position | Nearest gene | P (dis) | Beta (dis) | FDR (dis) | Study |
|---|---|---|---|---|---|---|---|
| cg05072008 | 7 | 50518647 | FIGNL1 | 2.472E-4 | 0.522 | 0.037 | DMG:Wockner_2014 |
 eQTL annotation
eQTL annotation
| SNP ID | Chromosome | Position | eGene | Gene Entrez ID | pvalue | qvalue | TSS distance | eQTL type |
|---|---|---|---|---|---|---|---|---|
| rs2244353 | chr7 | 50739454 | FIGNL1 | 63979 | 0.14 | cis | ||
| rs2244351 | chr7 | 50739525 | FIGNL1 | 63979 | 0.14 | cis | ||
| rs2244347 | chr7 | 50739587 | FIGNL1 | 63979 | 0.11 | cis | ||
| rs11763774 | 7 | 50462138 | FIGNL1 | ENSG00000132436.7 | 5.577E-7 | 0 | 55950 | gtex_brain_ba24 |
| rs397717279 | 7 | 50477015 | FIGNL1 | ENSG00000132436.7 | 4.018E-7 | 0 | 41073 | gtex_brain_ba24 |
| rs12719030 | 7 | 50485073 | FIGNL1 | ENSG00000132436.7 | 5.331E-7 | 0 | 33015 | gtex_brain_ba24 |
| rs4947641 | 7 | 50489038 | FIGNL1 | ENSG00000132436.7 | 1.257E-7 | 0 | 29050 | gtex_brain_ba24 |
| rs11764792 | 7 | 50496841 | FIGNL1 | ENSG00000132436.7 | 2.483E-7 | 0 | 21247 | gtex_brain_ba24 |
| rs6945253 | 7 | 50501440 | FIGNL1 | ENSG00000132436.7 | 3.938E-8 | 0 | 16648 | gtex_brain_ba24 |
| rs4947827 | 7 | 50507863 | FIGNL1 | ENSG00000132436.7 | 7.432E-8 | 0 | 10225 | gtex_brain_ba24 |
| rs12719056 | 7 | 50510076 | FIGNL1 | ENSG00000132436.7 | 7.432E-8 | 0 | 8012 | gtex_brain_ba24 |
| rs12154303 | 7 | 50515627 | FIGNL1 | ENSG00000132436.7 | 5.903E-8 | 0 | 2461 | gtex_brain_ba24 |
| rs11768267 | 7 | 50533539 | FIGNL1 | ENSG00000132436.7 | 7.877E-8 | 0 | -15451 | gtex_brain_ba24 |
| rs11575534 | 7 | 50533956 | FIGNL1 | ENSG00000132436.7 | 8.858E-8 | 0 | -15868 | gtex_brain_ba24 |
| rs730092 | 7 | 50535754 | FIGNL1 | ENSG00000132436.7 | 4.629E-8 | 0 | -17666 | gtex_brain_ba24 |
| rs71018472 | 7 | 50536827 | FIGNL1 | ENSG00000132436.7 | 1.292E-7 | 0 | -18739 | gtex_brain_ba24 |
| rs7803247 | 7 | 50540743 | FIGNL1 | ENSG00000132436.7 | 7.887E-8 | 0 | -22655 | gtex_brain_ba24 |
| rs732215 | 7 | 50544063 | FIGNL1 | ENSG00000132436.7 | 5.342E-7 | 0 | -25975 | gtex_brain_ba24 |
| rs2329365 | 7 | 50544945 | FIGNL1 | ENSG00000132436.7 | 9.966E-7 | 0 | -26857 | gtex_brain_ba24 |
| rs4989851 | 7 | 50544954 | FIGNL1 | ENSG00000132436.7 | 9.966E-7 | 0 | -26866 | gtex_brain_ba24 |
| rs2876827 | 7 | 50544975 | FIGNL1 | ENSG00000132436.7 | 9.966E-7 | 0 | -26887 | gtex_brain_ba24 |
| rs3887825 | 7 | 50545093 | FIGNL1 | ENSG00000132436.7 | 7.635E-7 | 0 | -27005 | gtex_brain_ba24 |
| rs4992502 | 7 | 50545191 | FIGNL1 | ENSG00000132436.7 | 9.95E-7 | 0 | -27103 | gtex_brain_ba24 |
| rs12718527 | 7 | 50545549 | FIGNL1 | ENSG00000132436.7 | 9.95E-7 | 0 | -27461 | gtex_brain_ba24 |
| rs12718528 | 7 | 50545571 | FIGNL1 | ENSG00000132436.7 | 9.95E-7 | 0 | -27483 | gtex_brain_ba24 |
| rs12718529 | 7 | 50545593 | FIGNL1 | ENSG00000132436.7 | 9.95E-7 | 0 | -27505 | gtex_brain_ba24 |
| rs11763807 | 7 | 50545618 | FIGNL1 | ENSG00000132436.7 | 9.95E-7 | 0 | -27530 | gtex_brain_ba24 |
| rs11763809 | 7 | 50545621 | FIGNL1 | ENSG00000132436.7 | 9.943E-7 | 0 | -27533 | gtex_brain_ba24 |
| rs11238138 | 7 | 50545992 | FIGNL1 | ENSG00000132436.7 | 1.117E-6 | 0 | -27904 | gtex_brain_ba24 |
| rs4329234 | 7 | 50546006 | FIGNL1 | ENSG00000132436.7 | 5.572E-7 | 0 | -27918 | gtex_brain_ba24 |
| rs4433100 | 7 | 50546037 | FIGNL1 | ENSG00000132436.7 | 1.117E-6 | 0 | -27949 | gtex_brain_ba24 |
| rs4579483 | 7 | 50546100 | FIGNL1 | ENSG00000132436.7 | 1.131E-6 | 0 | -28012 | gtex_brain_ba24 |
| rs4580999 | 7 | 50546261 | FIGNL1 | ENSG00000132436.7 | 1.117E-6 | 0 | -28173 | gtex_brain_ba24 |
| rs4436083 | 7 | 50546279 | FIGNL1 | ENSG00000132436.7 | 1.117E-6 | 0 | -28191 | gtex_brain_ba24 |
| rs1349492 | 7 | 50546459 | FIGNL1 | ENSG00000132436.7 | 1.115E-6 | 0 | -28371 | gtex_brain_ba24 |
| rs10899734 | 7 | 50546764 | FIGNL1 | ENSG00000132436.7 | 2.625E-6 | 0 | -28676 | gtex_brain_ba24 |
| rs10899735 | 7 | 50546891 | FIGNL1 | ENSG00000132436.7 | 2.625E-6 | 0 | -28803 | gtex_brain_ba24 |
| rs10899736 | 7 | 50546925 | FIGNL1 | ENSG00000132436.7 | 7.002E-7 | 0 | -28837 | gtex_brain_ba24 |
| rs11575462 | 7 | 50547182 | FIGNL1 | ENSG00000132436.7 | 2.58E-6 | 0 | -29094 | gtex_brain_ba24 |
| rs11575461 | 7 | 50547183 | FIGNL1 | ENSG00000132436.7 | 1.41E-6 | 0 | -29095 | gtex_brain_ba24 |
| rs11575457 | 7 | 50547584 | FIGNL1 | ENSG00000132436.7 | 2.625E-6 | 0 | -29496 | gtex_brain_ba24 |
| rs59827423 | 7 | 50547661 | FIGNL1 | ENSG00000132436.7 | 2.625E-6 | 0 | -29573 | gtex_brain_ba24 |
| rs4947577 | 7 | 50548609 | FIGNL1 | ENSG00000132436.7 | 2.63E-6 | 0 | -30521 | gtex_brain_ba24 |
| rs4948196 | 7 | 50548651 | FIGNL1 | ENSG00000132436.7 | 2.625E-6 | 0 | -30563 | gtex_brain_ba24 |
| rs13311361 | 7 | 50549008 | FIGNL1 | ENSG00000132436.7 | 2.625E-6 | 0 | -30920 | gtex_brain_ba24 |
| rs11980368 | 7 | 50549939 | FIGNL1 | ENSG00000132436.7 | 1.335E-7 | 0 | -31851 | gtex_brain_ba24 |
| rs12718541 | 7 | 50550144 | FIGNL1 | ENSG00000132436.7 | 2.625E-6 | 0 | -32056 | gtex_brain_ba24 |
| rs2167367 | 7 | 50551730 | FIGNL1 | ENSG00000132436.7 | 2.625E-6 | 0 | -33642 | gtex_brain_ba24 |
| rs2122822 | 7 | 50552152 | FIGNL1 | ENSG00000132436.7 | 2.63E-6 | 0 | -34064 | gtex_brain_ba24 |
| rs1451371 | 7 | 50553051 | FIGNL1 | ENSG00000132436.7 | 7.002E-7 | 0 | -34963 | gtex_brain_ba24 |
| rs1451372 | 7 | 50553871 | FIGNL1 | ENSG00000132436.7 | 2.625E-6 | 0 | -35783 | gtex_brain_ba24 |
| rs1597639 | 7 | 50555841 | FIGNL1 | ENSG00000132436.7 | 2.523E-6 | 0 | -37753 | gtex_brain_ba24 |
| rs1901313 | 7 | 50556798 | FIGNL1 | ENSG00000132436.7 | 2.473E-6 | 0 | -38710 | gtex_brain_ba24 |
| rs6953675 | 7 | 50557506 | FIGNL1 | ENSG00000132436.7 | 2.439E-6 | 0 | -39418 | gtex_brain_ba24 |
| rs7802199 | 7 | 50558441 | FIGNL1 | ENSG00000132436.7 | 2.39E-6 | 0 | -40353 | gtex_brain_ba24 |
| rs3470 | 7 | 50558966 | FIGNL1 | ENSG00000132436.7 | 2.366E-6 | 0 | -40878 | gtex_brain_ba24 |
| rs398004707 | 7 | 50559785 | FIGNL1 | ENSG00000132436.7 | 2.759E-6 | 0 | -41697 | gtex_brain_ba24 |
| rs6592952 | 7 | 50560255 | FIGNL1 | ENSG00000132436.7 | 2.304E-6 | 0 | -42167 | gtex_brain_ba24 |
| rs13234331 | 7 | 50560314 | FIGNL1 | ENSG00000132436.7 | 6.119E-7 | 0 | -42226 | gtex_brain_ba24 |
| . | 7 | 50560381 | FIGNL1 | ENSG00000132436.7 | 1.028E-6 | 0 | -42293 | gtex_brain_ba24 |
| rs199887122 | 7 | 50560392 | FIGNL1 | ENSG00000132436.7 | 3.553E-8 | 0 | -42304 | gtex_brain_ba24 |
| rs12718558 | 7 | 50561104 | FIGNL1 | ENSG00000132436.7 | 2.625E-6 | 0 | -43016 | gtex_brain_ba24 |
| rs11575423 | 7 | 50562549 | FIGNL1 | ENSG00000132436.7 | 1.109E-7 | 0 | -44461 | gtex_brain_ba24 |
| rs3807566 | 7 | 50564204 | FIGNL1 | ENSG00000132436.7 | 2.517E-6 | 0 | -46116 | gtex_brain_ba24 |
| rs3807563 | 7 | 50566390 | FIGNL1 | ENSG00000132436.7 | 2.456E-6 | 0 | -48302 | gtex_brain_ba24 |
| rs3779083 | 7 | 50568823 | FIGNL1 | ENSG00000132436.7 | 2.52E-6 | 0 | -50735 | gtex_brain_ba24 |
| rs3779082 | 7 | 50568840 | FIGNL1 | ENSG00000132436.7 | 2.625E-6 | 0 | -50752 | gtex_brain_ba24 |
| rs3823674 | 7 | 50571996 | FIGNL1 | ENSG00000132436.7 | 2.517E-6 | 0 | -53908 | gtex_brain_ba24 |
| rs12718574 | 7 | 50573848 | FIGNL1 | ENSG00000132436.7 | 2.667E-6 | 0 | -55760 | gtex_brain_ba24 |
| rs6945253 | 7 | 50501440 | FIGNL1 | ENSG00000132436.7 | 8.047E-7 | 0.01 | 16648 | gtex_brain_putamen_basal |
| rs4947827 | 7 | 50507863 | FIGNL1 | ENSG00000132436.7 | 1.548E-6 | 0.01 | 10225 | gtex_brain_putamen_basal |
| rs12719056 | 7 | 50510076 | FIGNL1 | ENSG00000132436.7 | 1.548E-6 | 0.01 | 8012 | gtex_brain_putamen_basal |
| rs10232974 | 7 | 50510092 | FIGNL1 | ENSG00000132436.7 | 1.863E-6 | 0.01 | 7996 | gtex_brain_putamen_basal |
| rs10243511 | 7 | 50510269 | FIGNL1 | ENSG00000132436.7 | 1.863E-6 | 0.01 | 7819 | gtex_brain_putamen_basal |
| rs2305358 | 7 | 50512284 | FIGNL1 | ENSG00000132436.7 | 1.863E-6 | 0.01 | 5804 | gtex_brain_putamen_basal |
| rs10230307 | 7 | 50512857 | FIGNL1 | ENSG00000132436.7 | 3.773E-6 | 0.01 | 5231 | gtex_brain_putamen_basal |
| rs10230343 | 7 | 50512929 | FIGNL1 | ENSG00000132436.7 | 1.863E-6 | 0.01 | 5159 | gtex_brain_putamen_basal |
| rs10239496 | 7 | 50515557 | FIGNL1 | ENSG00000132436.7 | 1.863E-6 | 0.01 | 2531 | gtex_brain_putamen_basal |
| rs935795 | 7 | 50518392 | FIGNL1 | ENSG00000132436.7 | 2.145E-6 | 0.01 | -304 | gtex_brain_putamen_basal |
| rs397961541 | 7 | 50518435 | FIGNL1 | ENSG00000132436.7 | 3.419E-6 | 0.01 | -347 | gtex_brain_putamen_basal |
| rs4947954 | 7 | 50520688 | FIGNL1 | ENSG00000132436.7 | 2.168E-6 | 0.01 | -2600 | gtex_brain_putamen_basal |
| rs1451376 | 7 | 50522325 | FIGNL1 | ENSG00000132436.7 | 3.773E-6 | 0.01 | -4237 | gtex_brain_putamen_basal |
| rs4947980 | 7 | 50522421 | FIGNL1 | ENSG00000132436.7 | 1.882E-6 | 0.01 | -4333 | gtex_brain_putamen_basal |
| rs7790230 | 7 | 50522857 | FIGNL1 | ENSG00000132436.7 | 3.773E-6 | 0.01 | -4769 | gtex_brain_putamen_basal |
| rs4947510 | 7 | 50525420 | FIGNL1 | ENSG00000132436.7 | 3.773E-6 | 0.01 | -7332 | gtex_brain_putamen_basal |
| rs7787209 | 7 | 50528153 | FIGNL1 | ENSG00000132436.7 | 7.062E-7 | 0.01 | -10065 | gtex_brain_putamen_basal |
| rs4515495 | 7 | 50528764 | FIGNL1 | ENSG00000132436.7 | 2.882E-6 | 0.01 | -10676 | gtex_brain_putamen_basal |
Section II. Transcriptome annotation
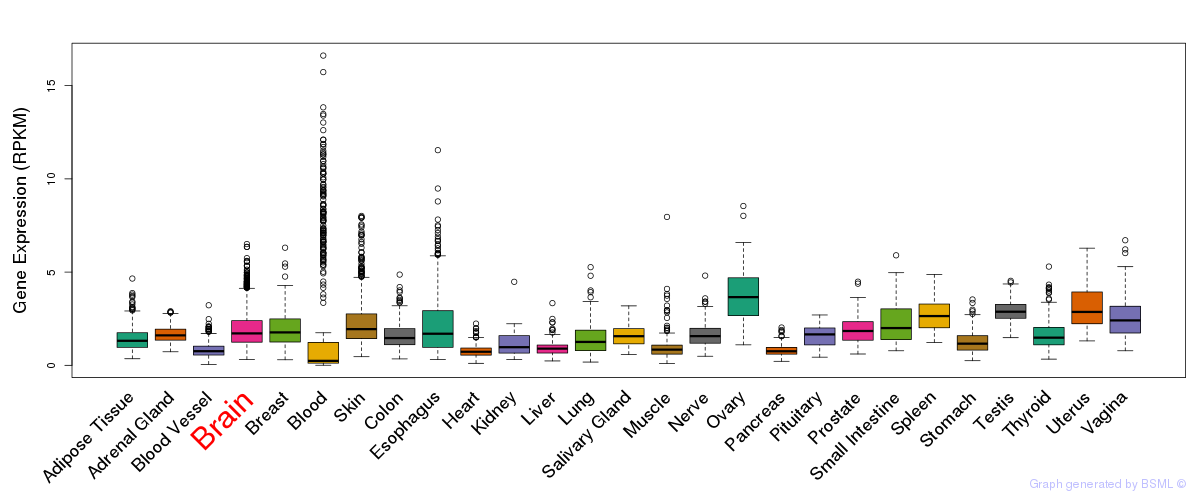
General gene expression (GTEx)
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
No co-expressed genes in brain regions
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| SENGUPTA NASOPHARYNGEAL CARCINOMA UP | 294 | 178 | All SZGR 2.0 genes in this pathway |
| GARY CD5 TARGETS DN | 431 | 263 | All SZGR 2.0 genes in this pathway |
| VECCHI GASTRIC CANCER EARLY UP | 430 | 232 | All SZGR 2.0 genes in this pathway |
| KINSEY TARGETS OF EWSR1 FLII FUSION UP | 1278 | 748 | All SZGR 2.0 genes in this pathway |
| DODD NASOPHARYNGEAL CARCINOMA DN | 1375 | 806 | All SZGR 2.0 genes in this pathway |
| RODRIGUES THYROID CARCINOMA POORLY DIFFERENTIATED UP | 633 | 376 | All SZGR 2.0 genes in this pathway |
| RODRIGUES THYROID CARCINOMA ANAPLASTIC UP | 722 | 443 | All SZGR 2.0 genes in this pathway |
| CHIARADONNA NEOPLASTIC TRANSFORMATION KRAS UP | 126 | 72 | All SZGR 2.0 genes in this pathway |
| LINDGREN BLADDER CANCER CLUSTER 3 UP | 329 | 196 | All SZGR 2.0 genes in this pathway |
| MARKEY RB1 ACUTE LOF DN | 228 | 137 | All SZGR 2.0 genes in this pathway |
| GRAESSMANN APOPTOSIS BY DOXORUBICIN DN | 1781 | 1082 | All SZGR 2.0 genes in this pathway |
| GRAESSMANN RESPONSE TO MC AND DOXORUBICIN DN | 770 | 415 | All SZGR 2.0 genes in this pathway |
| BERENJENO TRANSFORMED BY RHOA UP | 536 | 340 | All SZGR 2.0 genes in this pathway |
| KONG E2F3 TARGETS | 97 | 58 | All SZGR 2.0 genes in this pathway |
| RASHI RESPONSE TO IONIZING RADIATION 3 | 48 | 31 | All SZGR 2.0 genes in this pathway |
| IWANAGA E2F1 TARGETS INDUCED BY SERUM | 31 | 19 | All SZGR 2.0 genes in this pathway |
| LOCKWOOD AMPLIFIED IN LUNG CANCER | 214 | 139 | All SZGR 2.0 genes in this pathway |
| LE EGR2 TARGETS UP | 108 | 75 | All SZGR 2.0 genes in this pathway |
| BYSTRYKH HEMATOPOIESIS STEM CELL QTL TRANS | 882 | 572 | All SZGR 2.0 genes in this pathway |
| IVANOVA HEMATOPOIESIS LATE PROGENITOR | 544 | 307 | All SZGR 2.0 genes in this pathway |
| PAL PRMT5 TARGETS UP | 203 | 135 | All SZGR 2.0 genes in this pathway |
| BURTON ADIPOGENESIS 3 | 101 | 64 | All SZGR 2.0 genes in this pathway |
| WANG CISPLATIN RESPONSE AND XPC UP | 202 | 115 | All SZGR 2.0 genes in this pathway |
| GAVIN FOXP3 TARGETS CLUSTER P6 | 91 | 44 | All SZGR 2.0 genes in this pathway |
| MARSON BOUND BY E2F4 UNSTIMULATED | 728 | 415 | All SZGR 2.0 genes in this pathway |
| FUJII YBX1 TARGETS DN | 202 | 132 | All SZGR 2.0 genes in this pathway |
| TOYOTA TARGETS OF MIR34B AND MIR34C | 463 | 262 | All SZGR 2.0 genes in this pathway |
| ZHANG BREAST CANCER PROGENITORS UP | 425 | 253 | All SZGR 2.0 genes in this pathway |
| ZHENG GLIOBLASTOMA PLASTICITY UP | 250 | 168 | All SZGR 2.0 genes in this pathway |
| BOCHKIS FOXA2 TARGETS | 425 | 261 | All SZGR 2.0 genes in this pathway |
| GOLDRATH ANTIGEN RESPONSE | 346 | 192 | All SZGR 2.0 genes in this pathway |
| ZHANG TLX TARGETS 36HR DN | 185 | 116 | All SZGR 2.0 genes in this pathway |
| ZHANG TLX TARGETS 60HR DN | 277 | 166 | All SZGR 2.0 genes in this pathway |
| ZHANG TLX TARGETS DN | 89 | 50 | All SZGR 2.0 genes in this pathway |
| CHICAS RB1 TARGETS GROWING | 243 | 155 | All SZGR 2.0 genes in this pathway |
| DUTERTRE ESTRADIOL RESPONSE 24HR UP | 324 | 193 | All SZGR 2.0 genes in this pathway |
| PILON KLF1 TARGETS DN | 1972 | 1213 | All SZGR 2.0 genes in this pathway |
| JOHNSTONE PARVB TARGETS 3 DN | 918 | 550 | All SZGR 2.0 genes in this pathway |
| LEE BMP2 TARGETS DN | 882 | 538 | All SZGR 2.0 genes in this pathway |
| FEVR CTNNB1 TARGETS DN | 553 | 343 | All SZGR 2.0 genes in this pathway |
| GOBERT OLIGODENDROCYTE DIFFERENTIATION UP | 570 | 339 | All SZGR 2.0 genes in this pathway |
| JUBAN TARGETS OF SPI1 AND FLI1 UP | 115 | 73 | All SZGR 2.0 genes in this pathway |
| PEDERSEN METASTASIS BY ERBB2 ISOFORM 7 | 403 | 240 | All SZGR 2.0 genes in this pathway |
| PLASARI TGFB1 TARGETS 10HR DN | 244 | 157 | All SZGR 2.0 genes in this pathway |
| DELACROIX RAR BOUND ES | 462 | 273 | All SZGR 2.0 genes in this pathway |
| FORTSCHEGGER PHF8 TARGETS DN | 784 | 464 | All SZGR 2.0 genes in this pathway |
| RAO BOUND BY SALL4 ISOFORM B | 517 | 302 | All SZGR 2.0 genes in this pathway |