Gene Page: MEF2A
Summary ?
| GeneID | 4205 |
| Symbol | MEF2A |
| Synonyms | ADCAD1|RSRFC4|RSRFC9|mef2 |
| Description | myocyte enhancer factor 2A |
| Reference | MIM:600660|HGNC:HGNC:6993|Ensembl:ENSG00000068305|HPRD:02807|Vega:OTTHUMG00000171937 |
| Gene type | protein-coding |
| Map location | 15q26 |
| Pascal p-value | 0.156 |
| Sherlock p-value | 0.898 |
| Fetal beta | -0.883 |
| eGene | Myers' cis & trans |
Gene in Data Sources
| Gene set name | Method of gene set | Description | Info |
|---|---|---|---|
| CV:PGCnp | Genome-wide Association Study | GWAS | |
| PMID:cooccur | High-throughput literature-search | Systematic search in PubMed for genes co-occurring with SCZ keywords. A total of 3027 genes were included. | |
| Literature | High-throughput literature-search | Co-occurance with Schizophrenia keywords: schizophrenia,schizophrenias | Click to show details |
Section I. Genetics and epigenetics annotation
 eQTL annotation
eQTL annotation
| SNP ID | Chromosome | Position | eGene | Gene Entrez ID | pvalue | qvalue | TSS distance | eQTL type |
|---|---|---|---|---|---|---|---|---|
| rs4951059 | chr1 | 204258018 | MEF2A | 4205 | 0.18 | trans | ||
| rs17014160 | chr1 | 206855799 | MEF2A | 4205 | 0.1 | trans | ||
| rs12093398 | chr1 | 246113979 | MEF2A | 4205 | 0.04 | trans | ||
| rs12997903 | chr2 | 1952083 | MEF2A | 4205 | 0.11 | trans | ||
| rs6733654 | chr2 | 1963270 | MEF2A | 4205 | 0 | trans | ||
| rs891877 | chr2 | 2043348 | MEF2A | 4205 | 0.13 | trans | ||
| rs6754787 | chr2 | 30141949 | MEF2A | 4205 | 0.01 | trans | ||
| rs841285 | chr2 | 129330728 | MEF2A | 4205 | 0.13 | trans | ||
| rs7595584 | chr2 | 175652187 | MEF2A | 4205 | 8.135E-4 | trans | ||
| rs10191055 | chr2 | 201316238 | MEF2A | 4205 | 0.18 | trans | ||
| rs1155107 | chr3 | 26843376 | MEF2A | 4205 | 0.14 | trans | ||
| rs1395163 | chr3 | 31028951 | MEF2A | 4205 | 0.14 | trans | ||
| rs6446109 | chr3 | 60076988 | MEF2A | 4205 | 0.06 | trans | ||
| rs2339917 | chr3 | 179669447 | MEF2A | 4205 | 0.16 | trans | ||
| rs6848885 | chr4 | 21582684 | MEF2A | 4205 | 0.05 | trans | ||
| snp_a-4196049 | 0 | MEF2A | 4205 | 0.01 | trans | |||
| snp_a-1853527 | 0 | MEF2A | 4205 | 0 | trans | |||
| rs7677329 | chr4 | 21586153 | MEF2A | 4205 | 0.05 | trans | ||
| rs6813178 | chr4 | 71575238 | MEF2A | 4205 | 4.106E-4 | trans | ||
| rs6834662 | chr4 | 71605100 | MEF2A | 4205 | 0.15 | trans | ||
| rs12639667 | chr4 | 140243821 | MEF2A | 4205 | 0.06 | trans | ||
| rs16870339 | chr5 | 2705169 | MEF2A | 4205 | 4.945E-5 | trans | ||
| rs17146240 | chr5 | 119522933 | MEF2A | 4205 | 0.15 | trans | ||
| rs17146261 | chr5 | 119534252 | MEF2A | 4205 | 0.15 | trans | ||
| rs6930572 | chr6 | 31428168 | MEF2A | 4205 | 0.18 | trans | ||
| rs320082 | chr7 | 29105404 | MEF2A | 4205 | 0.03 | trans | ||
| rs317720 | chr7 | 29110633 | MEF2A | 4205 | 0.13 | trans | ||
| rs317714 | chr7 | 29115553 | MEF2A | 4205 | 0.02 | trans | ||
| rs317724 | chr7 | 29120565 | MEF2A | 4205 | 0.13 | trans | ||
| rs1831517 | chr9 | 2513405 | MEF2A | 4205 | 3.203E-4 | trans | ||
| rs1020772 | chr9 | 76532511 | MEF2A | 4205 | 0.07 | trans | ||
| rs10992675 | chr9 | 95963315 | MEF2A | 4205 | 0.06 | trans | ||
| rs4607971 | chr10 | 19407890 | MEF2A | 4205 | 0.01 | trans | ||
| rs7078127 | chr10 | 74847633 | MEF2A | 4205 | 0.14 | trans | ||
| rs12569493 | chr10 | 74886042 | MEF2A | 4205 | 0.14 | trans | ||
| rs16930362 | chr10 | 74890320 | MEF2A | 4205 | 0.04 | trans | ||
| rs2271904 | chr10 | 74894374 | MEF2A | 4205 | 0.14 | trans | ||
| rs16915029 | chr10 | 75001994 | MEF2A | 4205 | 0.03 | trans | ||
| rs11000607 | chr10 | 75089777 | MEF2A | 4205 | 0.19 | trans | ||
| rs6480700 | chr10 | 75101689 | MEF2A | 4205 | 0.16 | trans | ||
| rs12571454 | chr10 | 75208954 | MEF2A | 4205 | 0.05 | trans | ||
| rs1700369 | chr12 | 127842232 | MEF2A | 4205 | 0.13 | trans | ||
| rs1817693 | chr12 | 128395998 | MEF2A | 4205 | 0.19 | trans | ||
| rs2699067 | chr12 | 128397197 | MEF2A | 4205 | 0.19 | trans | ||
| rs2699066 | chr12 | 128397415 | MEF2A | 4205 | 0.19 | trans | ||
| rs10145685 | chr14 | 22794892 | MEF2A | 4205 | 0.03 | trans | ||
| rs17793827 | chr14 | 22797560 | MEF2A | 4205 | 0.01 | trans | ||
| rs10136383 | chr14 | 22801078 | MEF2A | 4205 | 0.01 | trans | ||
| rs2455956 | chr14 | 43524686 | MEF2A | 4205 | 0 | trans | ||
| rs2254834 | chr14 | 43596184 | MEF2A | 4205 | 0 | trans | ||
| rs1954396 | chr14 | 43601835 | MEF2A | 4205 | 1.569E-4 | trans | ||
| rs11158850 | chr14 | 70757171 | MEF2A | 4205 | 0 | trans | ||
| rs2239335 | chr16 | 22897794 | MEF2A | 4205 | 0.1 | trans | ||
| rs2369690 | chr16 | 22920851 | MEF2A | 4205 | 0.03 | trans | ||
| rs10500572 | chr16 | 73559356 | MEF2A | 4205 | 0.03 | trans | ||
| rs9965053 | chr18 | 3166191 | MEF2A | 4205 | 0.07 | trans | ||
| rs8087321 | chr18 | 3168615 | MEF2A | 4205 | 0.17 | trans | ||
| rs7239484 | chr18 | 55443711 | MEF2A | 4205 | 0.04 | trans | ||
| rs1761450 | chr19 | 54818034 | MEF2A | 4205 | 0.09 | trans |
Section II. Transcriptome annotation
General gene expression (GTEx)
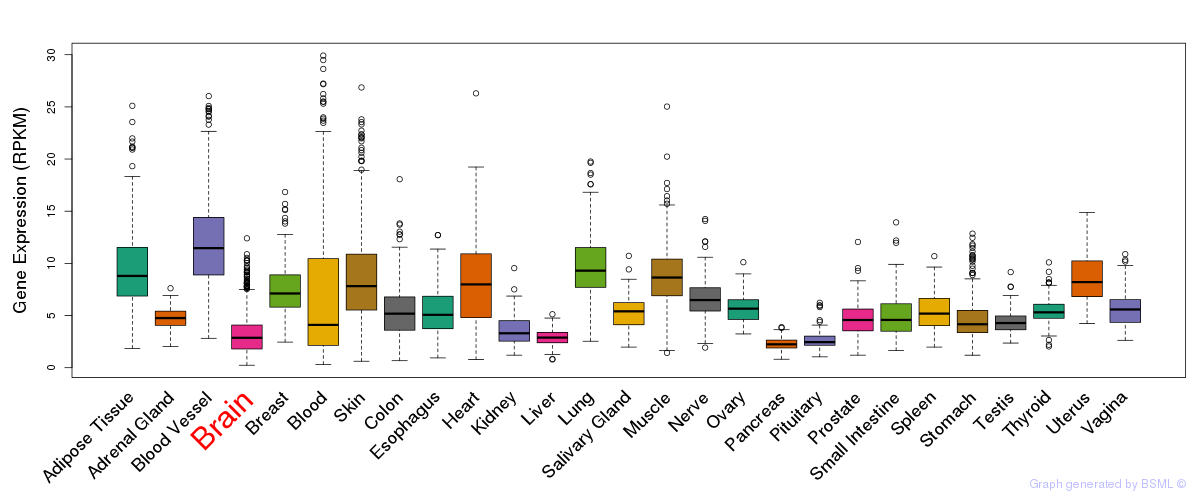
Gene expression during devlopment (BrainCloud)
Footnote:
A total of 269 time points ploted, with n=38 fetal samples (x=1:38). Each triangle represents one time point.
Gene expression of temporal and spatial changes (BrainSpan)
Footnote:
SC: sub-cortical regions; SM: sensory-motor regions; FC: frontal cortex; and TP: temporal-parietal cortex
ST1: fetal (13 - 26 postconception weeks), ST2: early infancy to late childhood (4 months to 11 years), and ST3: adolescence to adulthood (13 - 23 years)
The bar shown representes the lower 25% and upper 25% of the expression distribution.
No co-expressed genes in brain regions
Section IV. Protein-protein interaction annotation
| Interactors | Aliases B | Official full name B | Experimental | Source | PubMed ID |
|---|---|---|---|---|---|
| ASCL1 | ASH1 | HASH1 | MASH1 | bHLHa46 | achaete-scute complex homolog 1 (Drosophila) | - | HPRD,BioGRID | 8662987 |
| EP300 | KAT3B | p300 | E1A binding protein p300 | - | HPRD,BioGRID | 12371907 |
| HDAC4 | HA6116 | HD4 | HDAC-A | HDACA | KIAA0288 | histone deacetylase 4 | - | HPRD,BioGRID | 10487761 |
| HDAC5 | FLJ90614 | HD5 | NY-CO-9 | histone deacetylase 5 | - | HPRD,BioGRID | 10748098 |
| HDAC9 | DKFZp779K1053 | HD7 | HDAC | HDAC7 | HDAC7B | HDAC9B | HDAC9FL | HDRP | KIAA0744 | MITR | histone deacetylase 9 | Affinity Capture-Western Reconstituted Complex | BioGRID | 10487761 |10748098 |
| HDAC9 | DKFZp779K1053 | HD7 | HDAC | HDAC7 | HDAC7B | HDAC9B | HDAC9FL | HDRP | KIAA0744 | MITR | histone deacetylase 9 | - | HPRD | 10825153 |
| MAPK14 | CSBP1 | CSBP2 | CSPB1 | EXIP | Mxi2 | PRKM14 | PRKM15 | RK | SAPK2A | p38 | p38ALPHA | mitogen-activated protein kinase 14 | - | HPRD,BioGRID | 10330143 |
| MAPK7 | BMK1 | ERK4 | ERK5 | PRKM7 | mitogen-activated protein kinase 7 | - | HPRD,BioGRID | 9753748 |
| MEF2A | ADCAD1 | RSRFC4 | RSRFC9 | myocyte enhancer factor 2A | - | HPRD | 8575763 |
| MEF2D | DKFZp686I1536 | myocyte enhancer factor 2D | - | HPRD,BioGRID | 8798771 |9858528 |
| MYOD1 | MYF3 | MYOD | PUM | bHLHc1 | myogenic differentiation 1 | - | HPRD | 8548800|9418854 |
| MYOG | MYF4 | MYOGENIN | bHLHc3 | myogenin (myogenic factor 4) | - | HPRD,BioGRID | 1329097 |
| SMAD2 | JV18 | JV18-1 | MADH2 | MADR2 | MGC22139 | MGC34440 | hMAD-2 | hSMAD2 | SMAD family member 2 | - | HPRD,BioGRID | 11160896 |
| SMAD4 | DPC4 | JIP | MADH4 | SMAD family member 4 | - | HPRD | 11160896 |
| TEAD1 | AA | REF1 | TCF13 | TEF-1 | TEA domain family member 1 (SV40 transcriptional enhancer factor) | - | HPRD | 12061776 |
| THRA | AR7 | EAR7 | ERB-T-1 | ERBA | ERBA1 | MGC000261 | MGC43240 | NR1A1 | THRA1 | THRA2 | c-ERBA-1 | thyroid hormone receptor, alpha (erythroblastic leukemia viral (v-erb-a) oncogene homolog, avian) | - | HPRD,BioGRID | 12371907 |
| VGLL4 | KIAA0121 | VGL-4 | vestigial like 4 (Drosophila) | Two-hybrid | BioGRID | 15140898 |
Section V. Pathway annotation
| Pathway name | Pathway size | # SZGR 2.0 genes in pathway | Info |
|---|---|---|---|
| BIOCARTA AT1R PATHWAY | 36 | 28 | All SZGR 2.0 genes in this pathway |
| BIOCARTA HDAC PATHWAY | 32 | 25 | All SZGR 2.0 genes in this pathway |
| BIOCARTA MAPK PATHWAY | 87 | 68 | All SZGR 2.0 genes in this pathway |
| BIOCARTA P38MAPK PATHWAY | 40 | 31 | All SZGR 2.0 genes in this pathway |
| BIOCARTA PGC1A PATHWAY | 26 | 20 | All SZGR 2.0 genes in this pathway |
| BIOCARTA ERK5 PATHWAY | 18 | 15 | All SZGR 2.0 genes in this pathway |
| PID P38 ALPHA BETA DOWNSTREAM PATHWAY | 38 | 29 | All SZGR 2.0 genes in this pathway |
| REACTOME SIGNALLING BY NGF | 217 | 167 | All SZGR 2.0 genes in this pathway |
| REACTOME DEVELOPMENTAL BIOLOGY | 396 | 292 | All SZGR 2.0 genes in this pathway |
| REACTOME TRIF MEDIATED TLR3 SIGNALING | 74 | 54 | All SZGR 2.0 genes in this pathway |
| REACTOME NGF SIGNALLING VIA TRKA FROM THE PLASMA MEMBRANE | 137 | 105 | All SZGR 2.0 genes in this pathway |
| REACTOME NUCLEAR EVENTS KINASE AND TRANSCRIPTION FACTOR ACTIVATION | 24 | 20 | All SZGR 2.0 genes in this pathway |
| REACTOME ERK MAPK TARGETS | 21 | 17 | All SZGR 2.0 genes in this pathway |
| REACTOME MYOGENESIS | 28 | 20 | All SZGR 2.0 genes in this pathway |
| REACTOME MAP KINASE ACTIVATION IN TLR CASCADE | 50 | 37 | All SZGR 2.0 genes in this pathway |
| REACTOME MAPK TARGETS NUCLEAR EVENTS MEDIATED BY MAP KINASES | 30 | 24 | All SZGR 2.0 genes in this pathway |
| REACTOME TRAF6 MEDIATED INDUCTION OF NFKB AND MAP KINASES UPON TLR7 8 OR 9 ACTIVATION | 77 | 57 | All SZGR 2.0 genes in this pathway |
| REACTOME NFKB AND MAP KINASES ACTIVATION MEDIATED BY TLR4 SIGNALING REPERTOIRE | 72 | 53 | All SZGR 2.0 genes in this pathway |
| REACTOME MYD88 MAL CASCADE INITIATED ON PLASMA MEMBRANE | 83 | 63 | All SZGR 2.0 genes in this pathway |
| REACTOME INNATE IMMUNE SYSTEM | 279 | 178 | All SZGR 2.0 genes in this pathway |
| REACTOME ACTIVATED TLR4 SIGNALLING | 93 | 69 | All SZGR 2.0 genes in this pathway |
| REACTOME IMMUNE SYSTEM | 933 | 616 | All SZGR 2.0 genes in this pathway |
| REACTOME TOLL RECEPTOR CASCADES | 118 | 84 | All SZGR 2.0 genes in this pathway |
| GARY CD5 TARGETS UP | 473 | 314 | All SZGR 2.0 genes in this pathway |
| OSWALD HEMATOPOIETIC STEM CELL IN COLLAGEN GEL DN | 275 | 168 | All SZGR 2.0 genes in this pathway |
| HOEBEKE LYMPHOID STEM CELL UP | 95 | 64 | All SZGR 2.0 genes in this pathway |
| MULLIGHAN NPM1 MUTATED SIGNATURE 1 DN | 126 | 86 | All SZGR 2.0 genes in this pathway |
| MULLIGHAN NPM1 MUTATED SIGNATURE 2 DN | 77 | 46 | All SZGR 2.0 genes in this pathway |
| MULLIGHAN NPM1 SIGNATURE 3 DN | 162 | 116 | All SZGR 2.0 genes in this pathway |
| GRAESSMANN APOPTOSIS BY DOXORUBICIN DN | 1781 | 1082 | All SZGR 2.0 genes in this pathway |
| PEREZ TP53 TARGETS | 1174 | 695 | All SZGR 2.0 genes in this pathway |
| TURJANSKI MAPK7 TARGETS | 8 | 8 | All SZGR 2.0 genes in this pathway |
| TURJANSKI MAPK11 TARGETS | 5 | 5 | All SZGR 2.0 genes in this pathway |
| TURJANSKI MAPK14 TARGETS | 10 | 10 | All SZGR 2.0 genes in this pathway |
| SCHLOSSER SERUM RESPONSE DN | 712 | 443 | All SZGR 2.0 genes in this pathway |
| DACOSTA UV RESPONSE VIA ERCC3 DN | 855 | 609 | All SZGR 2.0 genes in this pathway |
| DACOSTA UV RESPONSE VIA ERCC3 COMMON DN | 483 | 336 | All SZGR 2.0 genes in this pathway |
| MARTORIATI MDM4 TARGETS FETAL LIVER DN | 514 | 319 | All SZGR 2.0 genes in this pathway |
| BUYTAERT PHOTODYNAMIC THERAPY STRESS UP | 811 | 508 | All SZGR 2.0 genes in this pathway |
| BALDWIN PRKCI TARGETS UP | 35 | 26 | All SZGR 2.0 genes in this pathway |
| BENPORATH NANOG TARGETS | 988 | 594 | All SZGR 2.0 genes in this pathway |
| NIKOLSKY BREAST CANCER 15Q26 AMPLICON | 22 | 19 | All SZGR 2.0 genes in this pathway |
| ROSS AML WITH CBFB MYH11 FUSION | 52 | 32 | All SZGR 2.0 genes in this pathway |
| MANALO HYPOXIA UP | 207 | 145 | All SZGR 2.0 genes in this pathway |
| HADDAD B LYMPHOCYTE PROGENITOR | 293 | 193 | All SZGR 2.0 genes in this pathway |
| NGUYEN NOTCH1 TARGETS DN | 86 | 67 | All SZGR 2.0 genes in this pathway |
| VERHAAK AML WITH NPM1 MUTATED DN | 246 | 180 | All SZGR 2.0 genes in this pathway |
| HOEGERKORP CD44 TARGETS TEMPORAL DN | 25 | 16 | All SZGR 2.0 genes in this pathway |
| BLALOCK ALZHEIMERS DISEASE DN | 1237 | 837 | All SZGR 2.0 genes in this pathway |
| WANG SMARCE1 TARGETS DN | 371 | 218 | All SZGR 2.0 genes in this pathway |
| LEE AGING CEREBELLUM DN | 86 | 66 | All SZGR 2.0 genes in this pathway |
| DELASERNA MYOD TARGETS UP | 89 | 51 | All SZGR 2.0 genes in this pathway |
| BLALOCK ALZHEIMERS DISEASE INCIPIENT DN | 165 | 106 | All SZGR 2.0 genes in this pathway |
| KEEN RESPONSE TO ROSIGLITAZONE DN | 106 | 68 | All SZGR 2.0 genes in this pathway |
| KRIGE RESPONSE TO TOSEDOSTAT 6HR UP | 953 | 554 | All SZGR 2.0 genes in this pathway |
| KRIGE RESPONSE TO TOSEDOSTAT 24HR UP | 783 | 442 | All SZGR 2.0 genes in this pathway |
| CREIGHTON ENDOCRINE THERAPY RESISTANCE 5 | 482 | 296 | All SZGR 2.0 genes in this pathway |
| MARSON BOUND BY FOXP3 UNSTIMULATED | 1229 | 713 | All SZGR 2.0 genes in this pathway |
| MARTINEZ RB1 TARGETS UP | 673 | 430 | All SZGR 2.0 genes in this pathway |
| MARTINEZ TP53 TARGETS UP | 602 | 364 | All SZGR 2.0 genes in this pathway |
| MARTINEZ RB1 AND TP53 TARGETS UP | 601 | 369 | All SZGR 2.0 genes in this pathway |
| MASSARWEH TAMOXIFEN RESISTANCE UP | 578 | 341 | All SZGR 2.0 genes in this pathway |
| WENDT COHESIN TARGETS UP | 33 | 19 | All SZGR 2.0 genes in this pathway |
| MILI PSEUDOPODIA HAPTOTAXIS UP | 518 | 299 | All SZGR 2.0 genes in this pathway |
| CHIARETTI T ALL RELAPSE PROGNOSIS | 19 | 15 | All SZGR 2.0 genes in this pathway |
| WONG ADULT TISSUE STEM MODULE | 721 | 492 | All SZGR 2.0 genes in this pathway |
| KOINUMA TARGETS OF SMAD2 OR SMAD3 | 824 | 528 | All SZGR 2.0 genes in this pathway |